Potri.015G025300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G025300.5 pacid=42776166 polypeptide=Potri.015G025300.5.p locus=Potri.015G025300 ID=Potri.015G025300.5.v4.1 annot-version=v4.1
ATGCTGCTGATAAAGCACGACAGGTCGTTTCATCATCTTTATGTGCTTATTGTGCTTCTGTTGCTCATGAAACTTGCTCCTGGGTTCATTTCTGGAGTGA
AAGGGGCAACTTTCGGGTGCATAGAGAGGGAGAGACAAGCCCTTCTCAAGTTTAAAGAAGACCTTATTGATGACTTTGGTCTTCTTTCAACTTGGGGAAG
TGAAGAAGAGAAGAGGGATTGCTGCAAATGGAGAGGAGTTCGATGCAACAATCGAACTGGTCATGTTACTCATCTTGATCTTCACCAGGAGAATTACATT
AACGGATATTTAACTGGTAAGATCAGTAATTCACTACTTGAGTTACAGCATTTGAGTTACCTGAATCTAAATAGAAACTCTTTTGAAGGGAGTTCTTTTC
CATATTTCATTGGTTCTCTAAAAAAATTAAGGTATCTTGATCTCTCTTCTATTGGTATTGTTGGAACTCTTTCAAATCAGTTTTGGAACCTTTCAAGACT
ACAGTATCTTGATCTTAGTGGTAATTATTATGTTAATTTTACAAGTCTTGACTTTCTCTCCAACCTTTTTTCTCTTGAATATCTTGATTTGAGTGGAAAC
AATCTCAGCCAAGTCATTGACTGGATACAAACTGTTAAGAAGTTTCCTTTCTTAAAAATCTTGCTGTTCAGGAATTGTGATCTTTCAAATAACAGTCCTC
CCTCTCTTTCTTCTACAAATTCTTCTAAATCCCTAGCTGTCATTGATCTCTCTCACAACTATCTTGCTTCTTCCACATTCAATTGGTTGTCTAATTTTAG
TAATAACCTTGTTGATCTTGATCTCAGTTATAATGATGGTGTTACTTTTAAAAGTCTTGACTTCCTTTCTAACCTTTTCTTTCTTGAGCATCTTCAGCTC
TCATATATCCAATTACAAGGCTTAATTCCAGAAGCTTTCGCAAACATGATTTCTCTTAGGACTCTTGATCTATCCTTTAACGAATTACAAGGTTTAATTC
CAGATGCTTTCACAAACATGACTTCTCTTAGGACTCTTGATCTATCCTGTAACCAATTACAAGGTTCAATTCCAGATGCTTTCACAAACATGACTTCTCT
TAGGACTCTTTATCTATCCTTTAACCATTTACAAGGTTCAATTCCAGATGCTTTCACAAACATGACATCTTTTAGGACTCTTGATCTATCCTTTAACCAA
TTACAAGGCGATTTGAGTACATTTGGGCGGATGTGCAGTTTAAAAGTACTGCATATGAGTGGAAACAATCTCACTGGAGAGCTGTCTCAGTTGTTTCAAG
ACTCGCATGGATGCGTGGAGAGCTCACTAGAGATTCTGCAACTAGACGGAAATCAGCTTCATGGTTCCGTGCCTGACATCACAAGATTCACTTCCATGAC
AGAGCTGGATCTCTCTCGGAATCAACTGAATGGATCTTTACCGAAACGATTTAGTCAACGTTCTGAAATTGTTATCTTGTACTTGAATGACAACCAACTC
ACCGGATCACTTGCGGATGTTACGATGCTTTCATCATTGAGAGAGTTTGTGATTGCTAACAATCGCTTAGATGGGAATGTTTCTGAAAGTATTGGAAGCC
TATATCAGCTTGAGCAATTGGATGTTGGCAGGAATTCCCTACAAGGTGTGATGTCTGAAGCTCACTTTTCAAATCTCTCCAAATTAACGGTATTGGATTT
AACCGATAACTCCCTAGCTTTGAAATTTGAGTCTAACTGGGCTCCCACTTTTCAATTGGATCGCATATTTCTTTCGTCTTGTAATTTGGGTCCTCATTTC
CCTCAGTGGCTTCGAAATCAAAACAATTTCATGGAGCTAGATATCTCTGGTTCTAGAATTTCAGATACAGTCCCTAATTGGTTTTGGAACTTGTCCAATT
CCAAGTTGCAACTCTTAAACCTTTCTCACAACAAGATGAGTGGAATTTTGCCAGATTTCTCATCAAAATATTCCATTCTCCGAAACATGGATTTGAGTTT
CAACCAGTTTGAGGGTCCATTACCACTTTTTTCATCAGATACCATATCAACCTTGTTTCTCTCGAACAATAAGTTTTCAGGTTCAGCTTCGTTTCTATGT
AATATTGGAAGAAATATCAGTGTCCTTGACCTCTCAAATAACCTGTTGACGGGATGGATTCCTGATTGTTCAATGAATTTCACTCGTCTAAATATTCTGA
ATTTCGCCAGCAACAACTTTTCAGGTAAAATTCCCAGCTCCATTGGATCAATGTTCCATCTTCAAACATTGAGTTTGCATAATAATAGCTTTGTTGGAGA
ATTACCTTCGTCTTTGAGGAAGTGTACTTCTCTAGTATTTTTAGACCTGAGCTCAAATATGTTGCGAGGAGAAATACCAGGCTGGATAGGAGAAAGCATG
CCCTCTCTGGAAGTCCTTTCCCTACAATCTAATGGATTCAATGGAAGCATACCGCAAAATCTTTGTCACCTATCAAATATTCTGATCTTGGACCTATCTC
TAAACAATATCTCAGGAATCATTCCAAAGTGCCTTAATAACCTCACTTTCATGGTTCGTAAAACAGCGTCAGAATATCTGAATAATGCTGTTTCTAGTCT
GTATTCCAGTACACCAGATGTTCTTAGTGCATATCAGAATAAAATCACGGTTGGATGGAAAGGAAGAGAGGATGATTATGGAAGCACACTAGGACTTTTG
AGGATCATTAATTTTGCAAGAAATAAATTAATAGGTGAAATTCCAGAAGAAATAACAGGACTCTTACTGTTGCTTGCACTGAATTTATCCGGAAATAATT
TGACTGGAGAAATCCCTCAAAAGATTTGGCAGTTGAAACAGTTGGAATCACTTGATTTGTCAGGAAACCAACTATCAGGAGTGATTCCTATCACAATGGC
TGATCTAAATTTTCTGGCCTTCTTGAACCTCTCCAACAATCACTTGTCGGGGAGGATTCCGTCAAGCACTCAGCTGCAAGGCTTTAATGCTTCACAATTT
ACCGGTAACCTTGCATTGTGCGGGAAGCCACTCTTACAGAGATGCCCTGGAGATGAGACGAATCAAAGTCCACCTGCTAATGATGACAACCGAGGCAAAG
AAGTCGTTGCTGATGAATTCATGAAATGGTTTTGTACTGCTATGGGAATTGGTTTTAGTGTATTCTTTTGGGGAGTTTCAGGTGCTTTACTGCTAAAACG
TTCCTGGAGACATGCCTATTTCCGGTTCTTGGATGAATCATGGGACTGGCTCTATGTGAAGGTAGCAGTGCGCAAGGCACGACTTCAACGAGCCTTTCAG
TGGCTGCACGAGCACGGTCTTGCTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G025300.5 pacid=42776166 polypeptide=Potri.015G025300.5.p locus=Potri.015G025300 ID=Potri.015G025300.5.v4.1 annot-version=v4.1
MLLIKHDRSFHHLYVLIVLLLLMKLAPGFISGVKGATFGCIERERQALLKFKEDLIDDFGLLSTWGSEEEKRDCCKWRGVRCNNRTGHVTHLDLHQENYI
NGYLTGKISNSLLELQHLSYLNLNRNSFEGSSFPYFIGSLKKLRYLDLSSIGIVGTLSNQFWNLSRLQYLDLSGNYYVNFTSLDFLSNLFSLEYLDLSGN
NLSQVIDWIQTVKKFPFLKILLFRNCDLSNNSPPSLSSTNSSKSLAVIDLSHNYLASSTFNWLSNFSNNLVDLDLSYNDGVTFKSLDFLSNLFFLEHLQL
SYIQLQGLIPEAFANMISLRTLDLSFNELQGLIPDAFTNMTSLRTLDLSCNQLQGSIPDAFTNMTSLRTLYLSFNHLQGSIPDAFTNMTSFRTLDLSFNQ
LQGDLSTFGRMCSLKVLHMSGNNLTGELSQLFQDSHGCVESSLEILQLDGNQLHGSVPDITRFTSMTELDLSRNQLNGSLPKRFSQRSEIVILYLNDNQL
TGSLADVTMLSSLREFVIANNRLDGNVSESIGSLYQLEQLDVGRNSLQGVMSEAHFSNLSKLTVLDLTDNSLALKFESNWAPTFQLDRIFLSSCNLGPHF
PQWLRNQNNFMELDISGSRISDTVPNWFWNLSNSKLQLLNLSHNKMSGILPDFSSKYSILRNMDLSFNQFEGPLPLFSSDTISTLFLSNNKFSGSASFLC
NIGRNISVLDLSNNLLTGWIPDCSMNFTRLNILNFASNNFSGKIPSSIGSMFHLQTLSLHNNSFVGELPSSLRKCTSLVFLDLSSNMLRGEIPGWIGESM
PSLEVLSLQSNGFNGSIPQNLCHLSNILILDLSLNNISGIIPKCLNNLTFMVRKTASEYLNNAVSSLYSSTPDVLSAYQNKITVGWKGREDDYGSTLGLL
RIINFARNKLIGEIPEEITGLLLLLALNLSGNNLTGEIPQKIWQLKQLESLDLSGNQLSGVIPITMADLNFLAFLNLSNNHLSGRIPSSTQLQGFNASQF
TGNLALCGKPLLQRCPGDETNQSPPANDDNRGKEVVADEFMKWFCTAMGIGFSVFFWGVSGALLLKRSWRHAYFRFLDESWDWLYVKVAVRKARLQRAFQ
WLHEHGLA
|
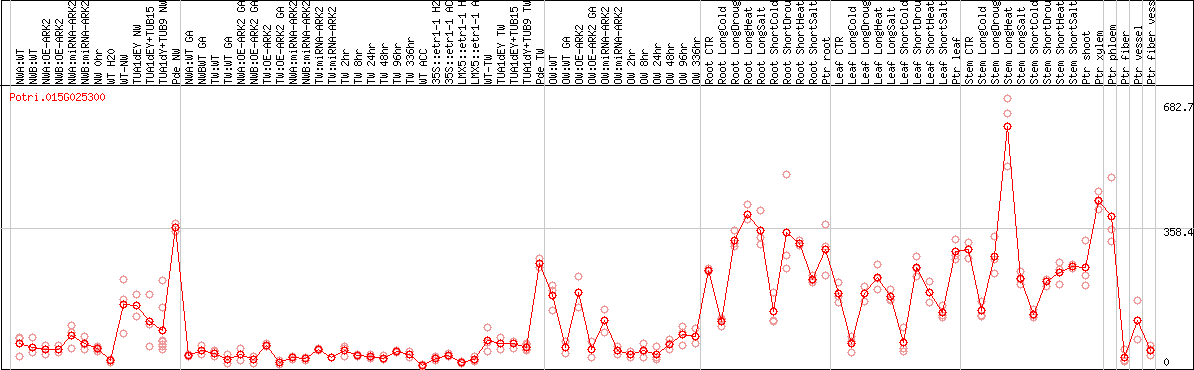
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G025300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
