Potri.015G036300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G036300.4 pacid=42776023 polypeptide=Potri.015G036300.4.p locus=Potri.015G036300 ID=Potri.015G036300.4.v4.1 annot-version=v4.1
ATGGCGATCGTAACAGGAGATCGTTATCTAGAAAAACTAGTCAAATTCGTTGAAGAACAAGCCGGTTCGTTAATTGACGGGACCCTTGTCTTAAAGCTTT
ACCCAGGCGGTCTCCGCTACGTCGATTCACGACTCGAATCACTACACGAGTTAGAGAATTTATTATCCGGAGCTCCAGTTGATTATCTTCGCGCTTACAT
TTCTGACCTCGGTGACCACCGTGCTCTCGAGCAGTTACGGCGGATTCTCCGTTTGTTGACGGAGTTAAAGGTTGTTTCGGTTTTGCCTCCGACGACGCGT
GATCCGACGCCGGTTTGTTTGGTTCCGTTTGGGAGATTGAGGGTTTTGGAACTTAGAGGTTGTGATTTGTCTACGTCAGCTGCCAAAGGGTTGCTTGAAT
TGAGACATACTCTGGAGAAGATTGTTTGTCACAATTCTACTGATGCATTACGGCATGTATTTGCTAGTAGGATTGTGGAGATAAAGGACTCACCGCAATG
GAATCGGTTGTCATTTGTATCGTGTGCGTGTAATCGACTGATTTTGATGGATGAGTCTTTGCAACTTCTTCCTGTTGTGGAAACTCTTGATTTGAGTCGA
AATAAGTTTGCAAAGGTGGATAATTTGCGGAAGTGTACCAAATTGAAGCACTTGGATCTTGGGTTCAATCATCTGAGATCAATTGCACCCTTTTATGAGA
TCTCCTGTCACATTGTTAAACTTGTTTTGAGGAACAATGCCCTAACAACATTACATGGGCTTGAGAATTTGAAGTCACTTGAAGCCCTTGATGTGTCCTA
TAATATTATTTCCAATTTCTCAGAGCTGGAGTTCCTTACTGGCCTCCCATGTTTGCGAAACTTGTGGTTGGAAGGAAATCCTTTGTGCGGTGCCCGATGG
TATCGGGCTCAAGTATTCAGCTATGTTGTTCATCCAGAAGCAGTGAAGTTAGACGACCAAGAAATCAGCGCAAGAGAGTTTTGGAAGAGGCAAATCATAA
TTGCCAGGAGGCAGAAACGACCTACCAGCTTTGGATTTTATTCTCCTGCTATAGGAGATGATGAAGGAGATGGAAATATCAACAGGAAAAGGAGTAAAGT
ATCTCGTCTTGCTTCAATTTCGAATAAAGAAGAGACCATTTATTTTTCCTCTGACCAAGAGTCTCCATCTTTTGATAATGAGATTCAAAGCAAAGAGGAG
AATGACGTATCTGATGATGAAGCTGAAATAGTTGATCTGATTAACAGAGTTGAGCTCATGAAAAAGGAGCGTTCTACTCTTTGGTTGCGGGAATTTAAGG
ACTGGATGGATCATGAATCGGAGAATATTGTGGATTGCAGTACATATTGTGGGGTTACATTGCATCATGCAAAGGAAAATCATCCAACAAACAAGTCAAC
ACAGAAGGACCATTGTGATAGTTCTAGAGATTCTATGGATGATCTTCAAGCTTCTGGAGATGAGACCAGTACAAATCTCCTTGAGTCTAATAGTTCATTT
GTTGATACTGGATCATATGGTGGGGTTGCTTTACCAGGGATGGGGAACATGAATCTAAGACAGAAGCATCAAAAGTCTTATTTACATGAAGGGTCTGGTA
GTATGTCTATGCAAAGTAGAAGTTCCCACACAGGTAGCTCTACTGTCCAAGAAGTCCATACAATAGTTGGAAATGGAAGCATATCACTGTTGACCACTCA
TTCATCTCCTGCATATCCTAGGTCTCCTCCTCATTATGAGGAGGACATTTTGCAACGACGCAACAACTTGGTGGAAGAAATTTTGCAGCTATCTGCAGAA
TCCTATTCAGTGGCATCATCTGACAGTAATACTAGCTCTAGTGATGATGATTTATACGAATTTGGAGATTCATCATATGAAGCTGCTAAATCACAAAATG
AAGAGTATTTGAATCCCAAGGCCGGAGGGCAATTATCTTCTAATCCTTTGAAGGATCAAGGACATGGAATCCATCATGTGATGGAAAACGACAGTTATCT
AAATGATTCACAAACCTCAATTTCCACAAAATTTTTGAGTTCGAACTCCAATGATTTCTCTGCTGGCTCTCATGATGGTGAAAATGCTCATTTTGCCAAC
CCAGAGGCTGATTTGCTGGAGAAAGGAAAAAACAAAAGGAAACCAAGAAGAATAGTCATTTCTTTGCTAGAGAATATGGTTGGCAGGATAGGGAGACCAG
AAAAATTAAATGGAAATGGGGATACTTGTGGAGCTGGTTTGGTGGATGAGCAAGGGGAACAAATTGTTTGTGAGAGTGATTTTCATGTTACTGATAAGAA
ACAGCTGCACACAAATTCTTTTACAACCCTGGATGCTGTAAATGTCAATGGATTCTCTGATGATTTTATTGAGAATTATTTTAATGAGAAAGTAGCAGAT
TCCAGAATTAATGAAAGCTGTAGGAATTATATGCGATGTGATTGCATACTAGAGCCAGAATCCATGTACAGAGAAAGAGAAGTAGTTTTGTTGCTGAGCA
GTGAAGACAAGCTGTATGTACTGCTCATTGATGTAGCATTTGATGGATCGGGAAGCATTTTAAGTTTGCTGGGTTGGCACAGGGTTGAAGATGTTAGAGA
AGTTTTAGTTGGTATAGGACTTCAGGTCGTGAGGGTATACATTGAAAGGGGAGCAACCTACCTGTTTCTAACGCGGAGCATTGAGAAGTCAAGGCAGGTA
CTTGATATCTTGCAAGTTTCTGGACCATGCACAACAAATAATAAGTGTTTGCTGAAAAGTTTGGAGCAGGTTCAGGCTGAGTTGTTTTGGCAGAAAATAT
GTCGAGGTTTGAAACTGAGCATATTCCAATATTCTATGGTGCTTTTTAGGCACAGGAAGAACGAAGAGGACTCGTGGCTTCCCAGATCACTGTTTGTAAG
TGGGGGGCATGTGCTTTTGTGTGTTGAAGATCTTAAGCAGTTCAGGAGTTCCTCAGTGGATGCATCTTCGCCTCCATATTTCTTGCTTGATTCTTGTTGC
TCCATTAGTGATGTGTCTGAGCTGGTTATTGAGGCCAGGGAGAGCTGGTTTATAACCTTAGCCCTTCAACATGCCCCAAAAAGCCTATCATCCAAATCTC
AAAAGGATATCAAAACAAATGACAAGGATAACGAAGGTTCAGGTTCTCTGACATGGAAGCTTAAGTGGTTCTCAAAAGAGAACCTATTCAATTTTGTGGC
TTTATTAAGGGCAATTCATGCAGGGGTAGCCGCGCTACCTATGCTAGTAACATATACTCCGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G036300.4 pacid=42776023 polypeptide=Potri.015G036300.4.p locus=Potri.015G036300 ID=Potri.015G036300.4.v4.1 annot-version=v4.1
MAIVTGDRYLEKLVKFVEEQAGSLIDGTLVLKLYPGGLRYVDSRLESLHELENLLSGAPVDYLRAYISDLGDHRALEQLRRILRLLTELKVVSVLPPTTR
DPTPVCLVPFGRLRVLELRGCDLSTSAAKGLLELRHTLEKIVCHNSTDALRHVFASRIVEIKDSPQWNRLSFVSCACNRLILMDESLQLLPVVETLDLSR
NKFAKVDNLRKCTKLKHLDLGFNHLRSIAPFYEISCHIVKLVLRNNALTTLHGLENLKSLEALDVSYNIISNFSELEFLTGLPCLRNLWLEGNPLCGARW
YRAQVFSYVVHPEAVKLDDQEISAREFWKRQIIIARRQKRPTSFGFYSPAIGDDEGDGNINRKRSKVSRLASISNKEETIYFSSDQESPSFDNEIQSKEE
NDVSDDEAEIVDLINRVELMKKERSTLWLREFKDWMDHESENIVDCSTYCGVTLHHAKENHPTNKSTQKDHCDSSRDSMDDLQASGDETSTNLLESNSSF
VDTGSYGGVALPGMGNMNLRQKHQKSYLHEGSGSMSMQSRSSHTGSSTVQEVHTIVGNGSISLLTTHSSPAYPRSPPHYEEDILQRRNNLVEEILQLSAE
SYSVASSDSNTSSSDDDLYEFGDSSYEAAKSQNEEYLNPKAGGQLSSNPLKDQGHGIHHVMENDSYLNDSQTSISTKFLSSNSNDFSAGSHDGENAHFAN
PEADLLEKGKNKRKPRRIVISLLENMVGRIGRPEKLNGNGDTCGAGLVDEQGEQIVCESDFHVTDKKQLHTNSFTTLDAVNVNGFSDDFIENYFNEKVAD
SRINESCRNYMRCDCILEPESMYREREVVLLLSSEDKLYVLLIDVAFDGSGSILSLLGWHRVEDVREVLVGIGLQVVRVYIERGATYLFLTRSIEKSRQV
LDILQVSGPCTTNNKCLLKSLEQVQAELFWQKICRGLKLSIFQYSMVLFRHRKNEEDSWLPRSLFVSGGHVLLCVEDLKQFRSSSVDASSPPYFLLDSCC
SISDVSELVIEARESWFITLALQHAPKSLSSKSQKDIKTNDKDNEGSGSLTWKLKWFSKENLFNFVALLRAIHAGVAALPMLVTYTP
|
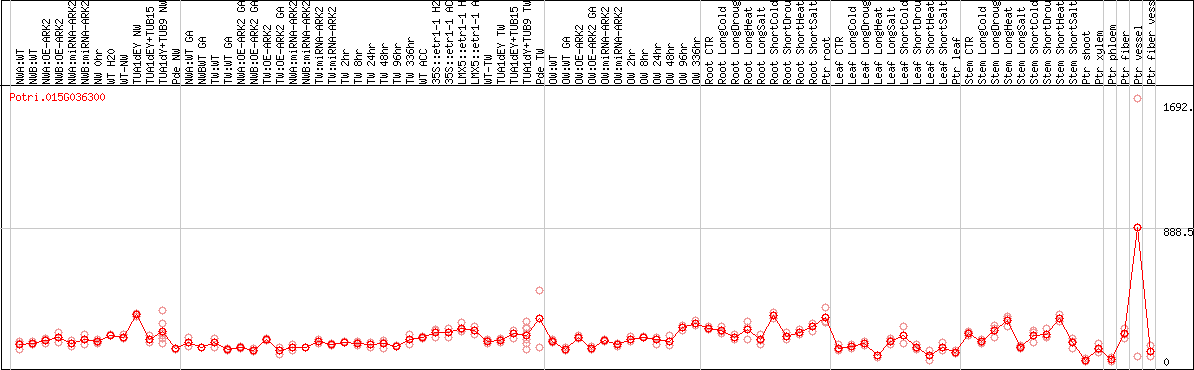
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G036300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
