Potri.015G037000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G037000.6 pacid=42776584 polypeptide=Potri.015G037000.6.p locus=Potri.015G037000 ID=Potri.015G037000.6.v4.1 annot-version=v4.1
ATGGCGACTGACGGTGCTGCGGTTGTTGGATTCGTCGGATTGGATGATTTGAGCTTAGACATGGCCGCTGCGCTTCTTCGTGCTGGCTACAAAGTTCAAG
CTTTCGAGATAGATGAAACTTTGGTTGATAAATTTTTGAATCTAGGGGGAACGAGAAGTGCCAGCCTTATTGAGGCTGGGAAAGAAGTTGCAGCTTTGAT
TGTTCTAATATCTCACGTGGATCAAATCAATGATGTATTTTTTGGCCAACAGGGTGTTTTGAAAGGATTGCAGAAAGGGGCACTTATCATCCTACGTTCA
ACCATTCTGCCATCATACATCCAGAATCTAGAGAAACGTCTCAGAGATGAAGATTCGATGGCTCATCTAATAGAAGCATATGTCTCCAGGGGATTTTCTG
AAGTCTTAAAGGGAAGAACCATGATTACATCCTCGGGAAGGTCTGAAGCTAATGCGAAGGCGCAGCCTATTCTTTCTGCAATGTCCGAAAAGCTTTTCAC
CTTCGAAGGTGAAGTAGGCACTGGAAGTAAAATTAAAATGGTGAATGAGCTGCTTGAGGGCATTCATCTTGTTGCTGCACTGGAGGCGATTTCTCTTTGC
ACTCAAGCTGGAATTCATCCATGGATAGTCTATGATATTATTTCCAATGCTGCTGGAAACTCATGGGTTTTCAAAAATCATATTCCCCAGTTTTTGAGAG
GTGATACCAAGGTGCATTCCTACAGGACCGTGGTCCAAAATTTGGGTATTGTTTTGGATACGGCCAAATCGCTTATATTTCCTCTTCCACTTTTGTCTGT
TGCTCATCAACAACTTATTCTTGGATCTTCATATGGTCAGGGTGATGATAGTGATGTCACATTTGTCAAGGTCTGGGGAAAGTTGCTTGGAGCAAATATT
CAAGATGCAGCCAGTGCAGAACTCTACGAACCTGAGCAACTGGCTAGACAAATTGTTGCCAAATCTGTTGTCGTAAAGAGAATTGGTTTCATTGGTCTTG
GAGCAATGGGATTTGGCATGGCAACTCACCTGTTGAAATCAAATTTCTGTGTTGTTGGTTATGATGTGTATAAACCAACTCTCACTCGATTTGCAAATGC
TGGTGGTTTGATTGGCAACTCTCCAGCAGAAACATCTAAAGATGTTGATGTACTAGTAGTTATGGTCACAAATGAAACTCAAGCGGAGAGTGTTTTGTAT
GGAGATCTAGGAGCAGTTGCAGCCTTACCATCAGGAGCATCTATCATCCTATCTTCTACAGTATCCCCTGCATTTGTGAGCCAACTGGAGCGACGCTTAC
AAGGTGAAGGCAAGGGCTTGAAATTGGTGGATGCTCCTGTTTCTGGTGGTGTCAAGAGGGCCTCAGAAGGAACACTTACGATAATGGCTTCTGGTACTGA
TGAAGCTCTTACATGTACCGGCTCAGTTCTCTCAGCATTAAGTGAGAAGCTTTATGTGATTAGAGGAGGTTGTGGGGCTGGAAGTGGTGTAAAGATGATT
AATCAGTTGCTCGCTGGAGTTCATATAGCATCAGGGGCTGAGGCAATGGCGCTTGGAGCTCGATTAGGTCTCAATACAAGGATGCTGTTTGATTTCGTCA
AGAACAGTGGAGGAACATCCTGGATGTTTGAGAATCGTGTTCCGCACATGTTGGACAACGATTATACACCGTATTCTGCACTCGATATCTTTGTGAAGGA
TCTGGGAATTGTTTGCCGTGAATCGTCATCCCTTAAAGTTCCACTTCATATTGCAACTGTTGCACACCAACTATTCCTGGCAGGCTCTGCTGCTGGCTGG
GGTCGGCAAGATGATGCAGGTGTCGTGAAGGTTTATGAGACCCTCACAGGTGTTAAAGTTGAAGGAACACTTCCTGTACTTAAGAAAGAAGTTGTCCTGC
AGTCTCTTCCACCTGAGTGGCCCCTGGACCCCATTGATGATATTCATAGACTGAACCAGAGCAATTCAAAAACGTTGGTGGTTCTAGATGATGATCCAAC
TGGAACTCAAACTGTTCATGATATTGAAGTATTAACAGAATGGAGTGTTGGGTCAATAGTTGAGCAATTTAGGAAAAAGCCTAAATGCTTTTTTATCTTG
ACCAACTCCAGGTCATTGAGTTCTGAAAAGGCGAGTGCACTGATTAAAGATATCTGTGGAAACCTAAGCATAGCCGCTAAGTCAGTTGAGAATATTGATT
ATACAGTAGTTTTGAGAGGCGATTCAACACTACGTGGTCATTTTCCAGAGGAAGCAGATGCAGCAGTTTCACTACTTGGTGAGATGGACGCATGGATCAT
TTGCCCATTTTTTCTTCAAGGAGGGCGTTATACAATCAAAGATATACACTATGTTGCAGATTCTGATTGGCTTGTCCCTGCAGGGGACACAGAGTTCGCC
AGAGATGCTTCTTTTGGCTACAAATCTTCAAACCTCCGTGAGTGGGTTGAGGAGAAAACTAGAGGCCGTATTCCTGCTAGCAGTGTTTCATCTATATCTA
TCAATCTTTTGAGGAAAGGTGGGCCAGATGCTGTTTGTGATACTCTTTGCAACTTGCAGAAGGGTTCAACTTGCATAGTTAATGCAGCAAGTGATCGAGA
CATGGCTGTATTTTCAGCTGGAATGATTCAGGCAGAATTAAGGGGGAAATCCTTTTTATGCCGCACTGCTGCTAGTTTTGTTTCTACCCGAATTGGAATT
ATTCCAAAAGCTCCAATACTGCCAAAGGATCTAGGAATAACCAAAGAAAGAAAGGGCGGGCTTATAGTTGTGGGATCGTATGTTCCCAAAACAACAAAAC
AGGTTGAAGAACTTAAATTACAATGTGGCCAGTTTTTAAAGAAACTAGAGGTTTCTGTTGATAAAATTGCCATGAAGTCATTAGAAGAGAGGGAGGAGGA
AATCAATAGAGTGGCTGAGATGGCAAATTTGCTCCTTGGGGCTTGTAAAGACACTTTAATCATGACCAGTCGAGAACTCATTACTGGAAAAACTGCTTCT
GAGAGCTTGGAAATCAACTTTAAGGTGAGCTCTGCTTTGGTGGAAATTGTGAGGCGGATATCAACAAGACCACGATACATTCTTGCAAAGGGTGGAATCA
CGTCATCCGACCTAGCTACAAAAGCTCTAGAAGCAAAATGTGCCAAAGTAGTTGGACAAGCCCTAGCTGGCATTCCATTGTGGCAGCTTGGTCCAGAAAG
TAGGCATCCAGGAGTTCCATACATTGTTTTCCCAGGTAATGTTGGTGACAGTAAAGCACTAGCTGATGTTGTGAAATCTTGGGCGCTTCCTTCCCGACTT
TCATCAACAAAAGAACTTCTTCTTAACGCAGAGAGAGGTGGATATGCTGTTGGAGCATTTAATGTCTATAATATGGAAGGGGCTGAGGCTGTTGTAGCTG
CAGCAGAGGAAGAAAATAGTCCTGCGATTCTACAGATTCATCCTAGTGCACTGAAGCAAGGAGGAATTCCATTGGTTGCTTGTTGTGTCTCTGCTGCTGA
ACAGGCCAATGTTCCAATCACTGTTCATTTTGATCATGGAACTTCGAAGCAAGAGCTAGTTGAAGCTCTTGATTTGGGATTTGATTCATTAATGGTAGAC
GGTTCACATCTTTCTCTCAAGGATAATATAGCATACACGAAGTATATATCACTTTTGGCCCACTCCAAAAACATGCTGGTGGAGGCTGAACTAGGGAGAT
TGTCAGGGACAGAGGATGATTTGACTGTTGAAGATTACGAAGCAAGGCTGACTGATGTTAATCAGGCTGAAGAGTTCATTGATGAGACTGGTATAGATGC
TTTGGCAGTGTGTATTGGGAATGTACATGGAAAGTACCCTGCAAGTGGTCCAAATCTCAGACTTGATTTGCTTAAGGATTTGCATGCCTTGAGTTCTAAG
AAAGGTGTATTTCTGGTGCTGCATGGAGCATCTGGCTTGTCCGAGGAACTTATCAAGGCTAGTATTCAGCGGGGTGTGACGAAGTTCAATGTTAACACTG
AGGTGCGCAACGCTTACATGAACTCACTAAGCAACCCCAAGAAAGATCTGGTCCATGTGATGGCTTCAGCAAAAGAAGCCATGAAAGCTGTGGTAGCAGA
GAAGATGCGCTTGTTTGGTTCATCTGGAAAAGCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G037000.6 pacid=42776584 polypeptide=Potri.015G037000.6.p locus=Potri.015G037000 ID=Potri.015G037000.6.v4.1 annot-version=v4.1
MATDGAAVVGFVGLDDLSLDMAAALLRAGYKVQAFEIDETLVDKFLNLGGTRSASLIEAGKEVAALIVLISHVDQINDVFFGQQGVLKGLQKGALIILRS
TILPSYIQNLEKRLRDEDSMAHLIEAYVSRGFSEVLKGRTMITSSGRSEANAKAQPILSAMSEKLFTFEGEVGTGSKIKMVNELLEGIHLVAALEAISLC
TQAGIHPWIVYDIISNAAGNSWVFKNHIPQFLRGDTKVHSYRTVVQNLGIVLDTAKSLIFPLPLLSVAHQQLILGSSYGQGDDSDVTFVKVWGKLLGANI
QDAASAELYEPEQLARQIVAKSVVVKRIGFIGLGAMGFGMATHLLKSNFCVVGYDVYKPTLTRFANAGGLIGNSPAETSKDVDVLVVMVTNETQAESVLY
GDLGAVAALPSGASIILSSTVSPAFVSQLERRLQGEGKGLKLVDAPVSGGVKRASEGTLTIMASGTDEALTCTGSVLSALSEKLYVIRGGCGAGSGVKMI
NQLLAGVHIASGAEAMALGARLGLNTRMLFDFVKNSGGTSWMFENRVPHMLDNDYTPYSALDIFVKDLGIVCRESSSLKVPLHIATVAHQLFLAGSAAGW
GRQDDAGVVKVYETLTGVKVEGTLPVLKKEVVLQSLPPEWPLDPIDDIHRLNQSNSKTLVVLDDDPTGTQTVHDIEVLTEWSVGSIVEQFRKKPKCFFIL
TNSRSLSSEKASALIKDICGNLSIAAKSVENIDYTVVLRGDSTLRGHFPEEADAAVSLLGEMDAWIICPFFLQGGRYTIKDIHYVADSDWLVPAGDTEFA
RDASFGYKSSNLREWVEEKTRGRIPASSVSSISINLLRKGGPDAVCDTLCNLQKGSTCIVNAASDRDMAVFSAGMIQAELRGKSFLCRTAASFVSTRIGI
IPKAPILPKDLGITKERKGGLIVVGSYVPKTTKQVEELKLQCGQFLKKLEVSVDKIAMKSLEEREEEINRVAEMANLLLGACKDTLIMTSRELITGKTAS
ESLEINFKVSSALVEIVRRISTRPRYILAKGGITSSDLATKALEAKCAKVVGQALAGIPLWQLGPESRHPGVPYIVFPGNVGDSKALADVVKSWALPSRL
SSTKELLLNAERGGYAVGAFNVYNMEGAEAVVAAAEEENSPAILQIHPSALKQGGIPLVACCVSAAEQANVPITVHFDHGTSKQELVEALDLGFDSLMVD
GSHLSLKDNIAYTKYISLLAHSKNMLVEAELGRLSGTEDDLTVEDYEARLTDVNQAEEFIDETGIDALAVCIGNVHGKYPASGPNLRLDLLKDLHALSSK
KGVFLVLHGASGLSEELIKASIQRGVTKFNVNTEVRNAYMNSLSNPKKDLVHVMASAKEAMKAVVAEKMRLFGSSGKA
|
DESeq2's median of ratios [POPLAR]
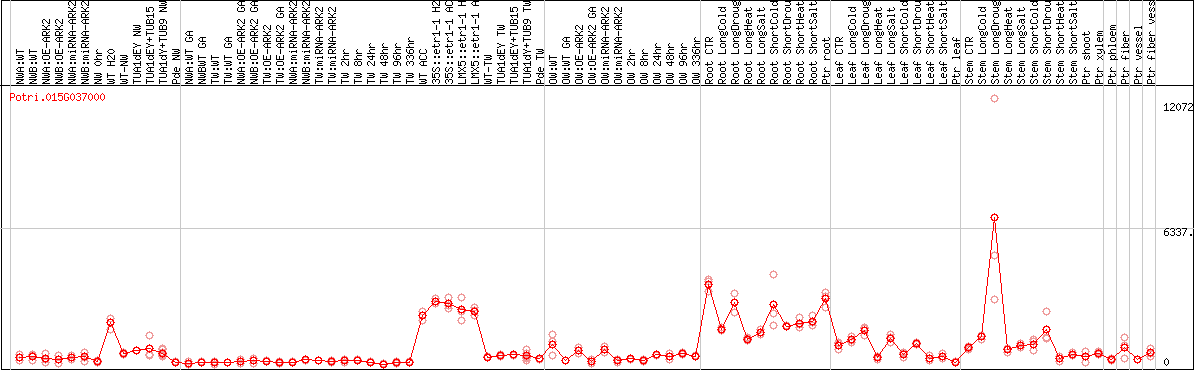
Coexpressed genes
Potri.015G037000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
