Potri.015G040100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G040100.2 pacid=42776499 polypeptide=Potri.015G040100.2.p locus=Potri.015G040100 ID=Potri.015G040100.2.v4.1 annot-version=v4.1
ATGAAGAACTTTCTCAAGAAGCTTCATATCATGCCTAATCAATCAGAGGATGCAGAGGGGTCAAATTCATCAAGAGGCCATAAGTCCACTAATGGATCAT
CCCCTGATAATAAGTCTTTGCATTCTAGGTCCCAAGAGAATAAACCCTTTTCTGGCCTTTCAAATTGGTTAAGTTCAGTTGCTAATAGAAAAAGTCCTAG
TCCACCATCATCTTCAAATGTGACTAGAGGAGAGAAGGTGGAGCAACCGGAGTCAATTAGTAGTAGTGGTTTTGATGTTGTTTCAGAAGCTGCAAGGCGT
GATTCAGGGTCTACCACCTCCAGGGATCCTGATATTGAAGAGGAGTATCAGATACAGCTGGCTCTGGAGTTGAGTGCAAGTGAGGATCCCGAGGCAGTTC
AGATTGAGGCTGTTAAGCAAATAAGCTTGGGCTCTTGTGCACCCGAGAATACTCCTGCTGAAGTCATTGCTTATCGATATTGGAATTACAATGCTCTTAG
CTATGATGACAAGGTCTTGGATGGTTTTTATGACCTGTATGGAATCATGACCGAGTCCACAACAGACAGAATGCCTCCTCTCGTTGATTTGCAAGGAACA
CCAGTCTCAGATGGTGTCACCTGGGAAGCAGTGCTGGTCAATAGGGCTGCTGATGCTAGTTTGTTGAAACTTGAACAGAAGGCACTGGAGATGACTGTCA
AGTCAAGGTCAGAATGTCAAATTTTCATTGGCAGTGCTTTGGTGGGAAGGCTTGCTGTTTTAGTTTCTGATTACATGGGTGGATCAGTTGGGGATCCCAG
CAATTTGTCAAGAGCATGGCGCAGTCTTAGCTATAGTTTGAAAGCAACCCTTGGTAGCATGGTTTTGCCACTTGGTTCTCTTACAATTGGCTTGCCTCGT
CATCGTGCATTAATGTTCAAGGTTTTGGCTGATAGTGTTGGCATACCATGCCGGTTGGTAAAAGGGCATCTGTACACTGGTTCTGATGATGTAGCTATGA
ACTTTGTAAAGCTCGATGATGGAAGGGAATACATTGTTGATCTAACAGCAGACCCTGGCACCCTTATTCCTTCTGATGCAGCTGGATCACATATAGAATA
CGATGAGACATTCTTTTCTTCCAGTCCTTTATCTAGAGACATTGACTCTTCTCATATAGCTTCTTCTAGTAGTGGACACACAAGTTCATTTGAAGAACAT
TCAGAACTTGGGACATTGGAAAAGCAATCCAGATTGAGAAATATTGCTGCTGTAGGGAACCAATCTGATGGAAGAAGTGAATCTCATGAGGGGGCAAGTT
TAACTAGGCCGAGTAAAAGTGGAGAAGAATCTACGATGTCTTCAGATGATTTTGGAAAAACTTCCAATGCTGAGAAGGTGCCAGTGCGGGAGCTTCCTGG
CAGACCTATTTACCCTTATGCACATGCTCGCTCTCCTTCTTGGACTGAAGGTGTTAGTTCTCCTGCTGCCCGTAGAATGAAAGTAAAAGATGTTTCACAA
TATATGATTGATGCTGCAAAAGAGAATCCACAGTTAGCCCAGAAACTTCACGATGTGTTGCTTGAAAGTGGTGTTGTTGCCCCCCCTAACCTGTTTACTG
AAATCTATGCTGAACAGTTGGATTTGTCTACTGCTGAGACCAAGTCCCCAACAGTAGATAAGGTTGATCACAAACAGAGGACTGAAATCCGAAGTGTGAA
GGATCAAGATGATCTTGTTCCTGCTCGATTTTTGCCTCCTCTGCCTCCTCATAGGCTGCCATATAAAGCCAGTTCACCTGGTAATCCACCTGATCAATCT
AAACCTGTGGAGGGTTCAGGGGTCAACCATCCTTTTGATACAAGGGAGATCACTGGACTACCCATACCATTGCAGTCTGAAGTTACTCCTGTGAAATATG
TAAAAAAAGTTCCTGTAGCTGCTGCAGCTGCTGCCGCTGCAGCTGTTGTTGCATCTTCTATGGTTGTTGCTGCAGCAAAGTCAGGTACTGACTCAAACCT
TGAACTACCAGTTGCAGCTGCTGCCACTGCCACAGCTGCAGCAGTGGTGGCAACCACAGCAGCTGTCAACAAGCAATATGAGCAGGGTGCACGGAGTGAT
GGAGATGCAGATAGTGCTGGTTATGAGCCACGAGGTAGTGGGGACAAGGGTAGTGGAGGACGCAGCAGTGAAGGCCATGGCAGTGGTGGCCAAGAATGTG
ATGCTTTAGGTGCCAATTCAGAAGGCGAGAGAATATCTGACAGATCAGTTGGAAATGATAGCTCAAAGTCTGATGCTGCGATGGATGATGTTGCAGAGTG
TGAGATTCCATGGGATGAAATTTCCTTGGGTGAACGTATTGGACTTGGATCATATGGAGAGGTATATCGTGGAGACTGGCATGGAACAGAAGTTGCTGTG
AAGAGGTTCCTAGACCAAGATATTACGGGTGAATCTCTTGCAGAATTCAGAAGTGAGGTTCGAATCATGAAGAGAGTCAGGCATCCCAATGTGGTTCTCT
TCATGGGAGCTGTAACTCGTGCCCCAAATCTTTCAATTGTTACAGAGTTTCTTCCTAGAGGTAGCTTGTATAGATTACTTCACCGGCCTAACAATCAATT
GGATGAGCGGAGGCGTTTGAGGATGGCCTTTGATGCTGCTCGGGGAATGAATTATTTGCACAACTGCACCCCAATGATAGTACACCGAGATTTGAAGTCC
CCCAATCTTCTGGTTGATAAAAATTGGGTAGTTAAGGTATGTGATTTTGGATTATCGAGGATGAAGCACAGCACATTTCTCTCTTCAAGATCAACTGCAG
GAACGGCTGAGTGGATGGCCCCAGAAGTGCTAAGAAATGAACCTTCAGACGAAAAGTGTGATGTTTATAGCTTCGGGGTTATATTGTGGGAGCTCTCTAC
ATTGCAACAACCATGGGGTGGAATGAACCCAATGCAAGTTGTTGGTGCGGTTGGATTTCAACATCGCCGTCTCGACATTCCAAATGATATGGATCCTACA
ATTGCAGATATTATTAGGAACTGCTGGAAGACAGATCCAAAATTAAGGCCAACATTTGCAGAAATCATGGCTGCGCTGAAGCCATTGCAGAAGCCTATAA
CTGGTCCACAAGTGCCCAGACCAAATGCATCATTGAGAAGTGGGCGTGAGAAGGTTCAGCTGTTTCAAGAAGCTGAAGACCAGGCAGGCTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G040100.2 pacid=42776499 polypeptide=Potri.015G040100.2.p locus=Potri.015G040100 ID=Potri.015G040100.2.v4.1 annot-version=v4.1
MKNFLKKLHIMPNQSEDAEGSNSSRGHKSTNGSSPDNKSLHSRSQENKPFSGLSNWLSSVANRKSPSPPSSSNVTRGEKVEQPESISSSGFDVVSEAARR
DSGSTTSRDPDIEEEYQIQLALELSASEDPEAVQIEAVKQISLGSCAPENTPAEVIAYRYWNYNALSYDDKVLDGFYDLYGIMTESTTDRMPPLVDLQGT
PVSDGVTWEAVLVNRAADASLLKLEQKALEMTVKSRSECQIFIGSALVGRLAVLVSDYMGGSVGDPSNLSRAWRSLSYSLKATLGSMVLPLGSLTIGLPR
HRALMFKVLADSVGIPCRLVKGHLYTGSDDVAMNFVKLDDGREYIVDLTADPGTLIPSDAAGSHIEYDETFFSSSPLSRDIDSSHIASSSSGHTSSFEEH
SELGTLEKQSRLRNIAAVGNQSDGRSESHEGASLTRPSKSGEESTMSSDDFGKTSNAEKVPVRELPGRPIYPYAHARSPSWTEGVSSPAARRMKVKDVSQ
YMIDAAKENPQLAQKLHDVLLESGVVAPPNLFTEIYAEQLDLSTAETKSPTVDKVDHKQRTEIRSVKDQDDLVPARFLPPLPPHRLPYKASSPGNPPDQS
KPVEGSGVNHPFDTREITGLPIPLQSEVTPVKYVKKVPVAAAAAAAAAVVASSMVVAAAKSGTDSNLELPVAAAATATAAAVVATTAAVNKQYEQGARSD
GDADSAGYEPRGSGDKGSGGRSSEGHGSGGQECDALGANSEGERISDRSVGNDSSKSDAAMDDVAECEIPWDEISLGERIGLGSYGEVYRGDWHGTEVAV
KRFLDQDITGESLAEFRSEVRIMKRVRHPNVVLFMGAVTRAPNLSIVTEFLPRGSLYRLLHRPNNQLDERRRLRMAFDAARGMNYLHNCTPMIVHRDLKS
PNLLVDKNWVVKVCDFGLSRMKHSTFLSSRSTAGTAEWMAPEVLRNEPSDEKCDVYSFGVILWELSTLQQPWGGMNPMQVVGAVGFQHRRLDIPNDMDPT
IADIIRNCWKTDPKLRPTFAEIMAALKPLQKPITGPQVPRPNASLRSGREKVQLFQEAEDQAG
|
DESeq2's median of ratios [POPLAR]
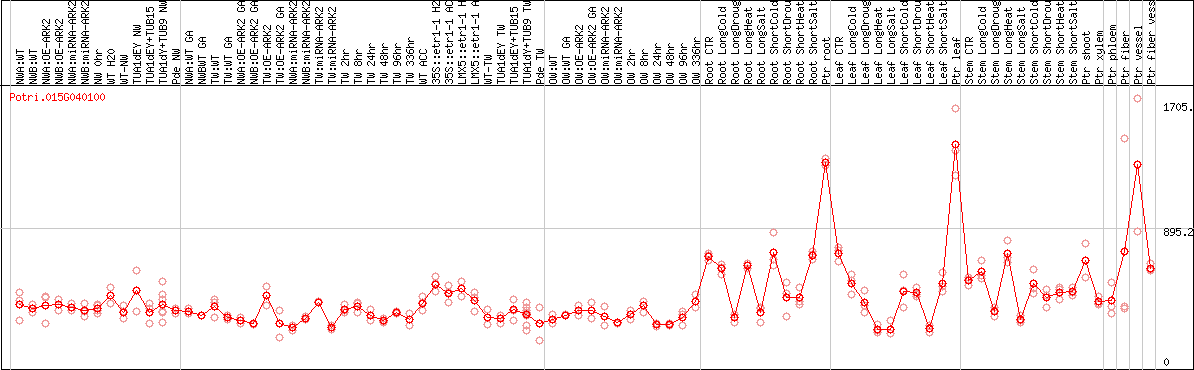
Coexpressed genes
Potri.015G040100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
