Potri.015G042500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G042500.1 pacid=42775295 polypeptide=Potri.015G042500.1.p locus=Potri.015G042500 ID=Potri.015G042500.1.v4.1 annot-version=v4.1
ATGCCTTGCTGTATCAGAATGAACACAATCATGTCTCTAAGACCCACTTCCACCAAAACAGCAAAATCTCTCCATTTCCATTACAAAGCTAAAACCACCC
TTAAACCCTCTTTATTTTCTTCATCATCTTTTGTTTCCGTTAGAAACCACCAAACCCTATCTTTTACTAACCAGCCATTGCCCCGCCCTCTTCGATTCAG
GCATCATGGGATTTTTTCGTTCAAGTGTTTTGGTGTTGCAGGTTTTCATTCCAAGAAACTTGCTTTGCCATATACCAGGTCTATGCAGAGCTTTAATTAT
GGCAGGTTTGCTTACCGGGATGTTTCTAGTGATGAATCCGATTATGAATTAGGCTCCTCCCAGAAGGAAATGACTGGCTCGACCCTTGACAACGTTGATG
ATTGGAAATGGAAGTTAACCATGCTTCTTCAAAGTAAAGATCAGCAAGAAGTTGTATCCAGGGAGAAAAAGGACAGACGTGATTTTGGGCATCTATCAGC
TATGGCAACCCGAATGGGTTTACACAGCCGTCAATATTCAAGAATTGTTGTGTTTAGTAAAGTCCCCTTGCCAAATTACAGACATGATCTGGATGATAAG
CGTCCACAGAGAGAGGTGATCTTACCTTTTGGGTTACAAAGAGAAGTAGATGCTCATTTCAAAGCTTACATTTCCAAAAAGCCCACGAGCAGAGGATTAT
TTCCTCCCAATTCCTTGTCAAGATCAAACGGTGGTAGGAGTATGGACACCGATGAAAGAATCTATGAGCGGCCAGAACTGTCAGTACAAAATAGTGTTGC
CATGGAGAGAATTCTTAGTCGGAAGAGTTTGCAGCTTCGTAATCAACAAGAAAAGTGGCAGGAATCTCCTGAGGGCCAAAAGATGATTGAATTTCGTAGA
AGCCTTCCTGCATATAAAGAAAAGGATGTGCTGTTAAAAGCCATATCTGAAAATCAGGTTATTGTTGTCTCTGGTGAAACTGGTTGTGGTAAAACCACAC
AACTTCCTCAATACATACTAGAATCTGAGATTGAAGCTGCTCGCGGAGCTGCATGTAGTATTATATGTACACAGCCTAGAAGAATATCGGCTATGGCTGT
TTCTGAAAGAGTTGCTGCAGAACGAGGGGAGAAACTGGGAGAATCAGTTGGTTATAAAGTTCGGCTCGAGGGTATGAGAGGAAGGGACACTCGACTTCTA
TTCTGTACGACTGGCATATTGTTGAGGAGATTACTTCTGGACAGAAATTTAAAAGGTGTAACTCATGTCATCGTTGATGAGATTCATGAACGCGGAATGA
ATGAAGATTTCCTTCTTATTGTACTGAGAGATCTGCTTCCTCGTCGACCGGAGTTGAGATTGATTTTGATGAGTGCAACCTTAAATGCGGAGCTTTTCTC
TTCCTACTTTGGCGATGCTCCTGCAATTCATATTCCTGGTTTTACATATCCTGTCCGTGCACATTTTTTAGAGAATATTCTTGAAATAACCGGGTATCGG
CTGACTCCTTACAATCAAATCGACGATTATGGTCAAGAAAAGACGTGGAAAATGCAAAAACAAGCTCAAGCTTTCAAAAAGAGGAAGAGCCAGATTGCTT
CTTCTGTTGAGGATGCCCTTGAAGTTGCTGACTTTAAGGGGTGTAGTTCACGCACTCGGGAGTCTTTATCTTGTTGGAATCCTGACTCAATTGGTTTTAA
CCTCATTGAGCATGTTCTTTGTCACATAGTTAAGAAAGAGAGACCTGGTGCTGTGTTGGTTTTTATGACTGGTTGGGATGACATTAACTCTTTAAAAGAT
CAGCTACAAGCTCATCCTATCTTGGGTGATCCTTGCAGAGTATTGTTACTGGCCTGCCATGGTTCCATGGCCAGCTCTGAGCAGAGGTTAATATTTGATA
AACCTGAAGATGGAGTGAGGAAAATTGTCTTAGCTACGAATATGGCTGAGACTAGTATTACTATCAATGATGTGGTATTTGTGGTTGACTGTGGAAAGGC
AAAGGAGACATCGTATGATGCTTTAAATAACACTCCTTGTTTGCTTCCATCCTGGATCTCAAAGGCTGCTGCCCGGCAAAGAAAGGGAAGAGCTGGTCGT
GTTCAACCTGGCGAGTGTTATCATCTTTATCCCAGATGTGTTTATGATGCATTTGCTGACTATCAATTACCAGAACTTTTGAGGACACCTTTACAGTCTT
TGAGTTTGCAAATCAAAAGTCTCCAACTTGGAAGTATATCAGAGTTCCTATCAAGGGCCCTGCAGCCACCAGAGCCTCTCTCGGTTCAAAATGCTGTTGA
GTATTTGAAGTTGATTGGGGCTTTAGATGAGCATGAAAATCTGACAGTGCTAGGACGTCATTTGTCAGTGCTTCCAGTTGAGCCTAAGCTCGGGAAAATG
CTCATATTGGGGACCATTTTCAACTGCCTAGATCCAATAATGACTGTCGTTGCTGGTCTCAGTGTTAGAGATCCATTCTTAATACCATTTGACAAGAAGG
ATCTTGCAGAGTCTGCAAAAGCACAATTCGCTGGTCGTGATTGCAGTGACCATCTTGCACTTGTCCGGGCCTATAATGGTTGGAAAGATGCAGAAAGACA
ACAATATGGTCATGAGTACTGTTGGAAGAATTTTCTTTCTGCACAAACTCTAAAAGCCATTGATTCCTTAAGGAAACAATTCTTTTATTTGCTCAAGGAT
ACTGGACTGGTTGATAAGCAGATAGAAAATTGCAATTCAAGGAGCATTGATGAGCATCTTATGCGAGCAGTCATCTGTGCTGGCCTGTTCCCTGGATTAT
GCTCTGTTGTGAACAAAGAGAAGTCGATAACACTGAAAACAATGGAGGATGGCCAGGTGCTTCTGTACTCGAATTCTGTGAATGCTGGGGTGCCCAAAAT
TCCATACCCTTGGTTAGTTTTCAATGAAAAAGTGAAAGTGAATTCAGTTTTTCTCCGTGATTCAACTGGTGTTTCTGATTCCGTGCTGCTTTTATTTGGA
GGAAACATTGAGAAGGGTGGACTAGATGGGCACCTAAAAATGTTGGGAGGATACTTGGAATTTTTTATGAAACCCACATTAGGAGATATGTATTTAAGCT
TAAAACGTGAGCTTGAGGAACTGATTCAGAATAAACTTCTAGATCCCAAGTTGGATATACAGTCCCACAATGAGCTTCTGATGGCAATAAGATTGCTGGT
TTCAGAGGACCAATGTGAGGGTAGATTTGTATTTGGTCGACAGCTGCCAGCACCCTCAAAGAAGGCAGAAAAGGCCAAAAATGTTGCTGGAGATGGGGGT
GACAATTCTAAGAATGAGCTTCAAACTTTACTTGCCAGGGCAGGGCATGAATCACCCGCTTATAAGACAAAACAACTGAAAAACAACCAGTTCCGTTCTA
CTGTTTTCTTTAACGGGCTGGACTTTGCTGGGCAGCCTTGCAGCAGTAAGAAACTTGCAGAAAAGGATGCAGCTGCTGCAGCTCTACTGTGGTTGAAGGG
GGAGACACATTCGTATTCCAGAAATACTGACCACTTCTCGGTGCTTCTGAAGAAGAGCAAGACCACAAATCAAAACAGAATCCCAGTTCGTGGTGGTAAA
TGGAATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G042500.1 pacid=42775295 polypeptide=Potri.015G042500.1.p locus=Potri.015G042500 ID=Potri.015G042500.1.v4.1 annot-version=v4.1
MPCCIRMNTIMSLRPTSTKTAKSLHFHYKAKTTLKPSLFSSSSFVSVRNHQTLSFTNQPLPRPLRFRHHGIFSFKCFGVAGFHSKKLALPYTRSMQSFNY
GRFAYRDVSSDESDYELGSSQKEMTGSTLDNVDDWKWKLTMLLQSKDQQEVVSREKKDRRDFGHLSAMATRMGLHSRQYSRIVVFSKVPLPNYRHDLDDK
RPQREVILPFGLQREVDAHFKAYISKKPTSRGLFPPNSLSRSNGGRSMDTDERIYERPELSVQNSVAMERILSRKSLQLRNQQEKWQESPEGQKMIEFRR
SLPAYKEKDVLLKAISENQVIVVSGETGCGKTTQLPQYILESEIEAARGAACSIICTQPRRISAMAVSERVAAERGEKLGESVGYKVRLEGMRGRDTRLL
FCTTGILLRRLLLDRNLKGVTHVIVDEIHERGMNEDFLLIVLRDLLPRRPELRLILMSATLNAELFSSYFGDAPAIHIPGFTYPVRAHFLENILEITGYR
LTPYNQIDDYGQEKTWKMQKQAQAFKKRKSQIASSVEDALEVADFKGCSSRTRESLSCWNPDSIGFNLIEHVLCHIVKKERPGAVLVFMTGWDDINSLKD
QLQAHPILGDPCRVLLLACHGSMASSEQRLIFDKPEDGVRKIVLATNMAETSITINDVVFVVDCGKAKETSYDALNNTPCLLPSWISKAAARQRKGRAGR
VQPGECYHLYPRCVYDAFADYQLPELLRTPLQSLSLQIKSLQLGSISEFLSRALQPPEPLSVQNAVEYLKLIGALDEHENLTVLGRHLSVLPVEPKLGKM
LILGTIFNCLDPIMTVVAGLSVRDPFLIPFDKKDLAESAKAQFAGRDCSDHLALVRAYNGWKDAERQQYGHEYCWKNFLSAQTLKAIDSLRKQFFYLLKD
TGLVDKQIENCNSRSIDEHLMRAVICAGLFPGLCSVVNKEKSITLKTMEDGQVLLYSNSVNAGVPKIPYPWLVFNEKVKVNSVFLRDSTGVSDSVLLLFG
GNIEKGGLDGHLKMLGGYLEFFMKPTLGDMYLSLKRELEELIQNKLLDPKLDIQSHNELLMAIRLLVSEDQCEGRFVFGRQLPAPSKKAEKAKNVAGDGG
DNSKNELQTLLARAGHESPAYKTKQLKNNQFRSTVFFNGLDFAGQPCSSKKLAEKDAAAAALLWLKGETHSYSRNTDHFSVLLKKSKTTNQNRIPVRGGK
WN
|
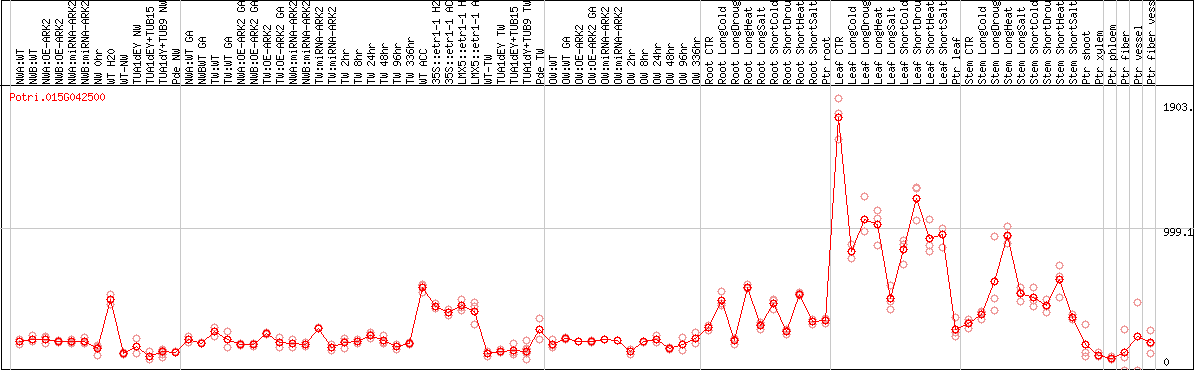
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G042500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
