SCD1.3 (Potri.015G049500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | SCD1.3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G049500.2 pacid=42775963 polypeptide=Potri.015G049500.2.p locus=Potri.015G049500 ID=Potri.015G049500.2.v4.1 annot-version=v4.1
ATGGCAGGGATCTTCGAGTATTTTGTAGTGTGTGGGTTGGGTCCGGAGATGAGAACTATGGATGGAAACAAAGGATATCATGGAATGCGAGTGTTGTATT
TGCCTTCTCTACTCGATCAATATCCTCCAGATAATCACTGTCTTTATCCTCCGCCTCCTCCTCAACTCCCAACTTGTGTGCTTCCTGCTGGTGTGGAGTT
TTACCCGTCAGGACTTGATGCAAATGATTCCTCCACATTTCCTAAAAGTTACCCCATTGTTTTGACTGAGGGTGATGGATCCAAAATTTATGTTAGTTGC
ATTGCCTTTCGAGATCCAGTTAGCGAGGATATTGCTGAAGCTTACCGTATACCTCCAAATTCATTTGCTGACAAATGTATCTGCCTTGTTTCCCGCTCTC
CTAGTTTTGGTGTTCTTAGAACTGCACTGGAGGAGTTATTTGCACTTTGCTTTTCCCCTGCTGGAAGTAGCAAGCCATTATGGGATGTTATATCATACAT
GGTCTCCAATGTACCTTTGCCTACTCCTGGAAAGGATCGGGTCTTGTTTGCCATTGAGAATTGCCTGCTTTCTGTGGAGGCTCCACCCAAAGATGGACTC
CCCCATGTTGAGATATCATTTCAGCCATTGGTACAGTGTTTGGATGTGGACAACTTGCTCAAACTTTTTACTGCCGTGTTGCTTGAAAGGAGGATTTTAC
TTCGTTCCAACAAGTATTCACTTTTAACCCTAGCATCAGAGGCCATATGTCATTTGATTTATCCCCTTAGGTGGCAGCATGTCTACATTCCATTACTATT
TTTCAGTGGAGTTGACTATATTGATGCTCCTACACCTTATATGATGGGTCTCCATTCAGCTGTTGATACATCTTACCTTGCAATGGATGGTGTAGTGGTT
GTGGACCTTGAGTATAACCGAATTTGTACCTCAGAGGAAATACCCCCTATTCCAGAGCCAGAATTAAGTACCTTGCGGGGTGAGATACTGAAGCTATTGT
ACCCCAATGTCATGGGAATAGATCAAATGAAAGCTGGTCTGGTTAGCTCATCTGAGCAATATTTTAAAGGCTGCAACAAGCCTTGGGGAGAAGATCATGA
CCTTCAACTTAGGTTAATATTCCTAAAGTTCTTTGCATCCATTTTAGGTGGATACCGGAATTTCATTGAAAACACTGCAACTCATGCCTTCAACACTCAA
GCATTTTTGAGGAAGCGCTCTCGGTCAACCAACCAACCACCAGATGCCATGATAACACAGTTCTTAGACTCCCATGGTTTCTTGGATTATTTGGAGAGAG
TTATTGATTCTGATGAAAACAATTATAATCTTCTTGACAAGTTACAAGACGCAATTGGAAGGGGTCAGAATCCCATTTCAGTTCTCCCCTCTTCGTGGGT
AGAGCCTGAAATTATAACCATATCAGATCCAGATGTTGGAATATTGGGATCAGGCGCTAAATTTACTTATGATAGGTTTCCCGCAAACATCAGGTCAGAA
GAACATGAGGAGAAGAGGAAGCAAATTCTTGCAGCAGCAAGTGGGGCTTTTGATTACATAAAGCATACTCCTAGTTCACCGTCTGTGCAAGTGGGCAAGG
ACTCTCTTAGTCCCATGGAGAGGGCTGCAGAAAGGGAGCGCATGGTTTTGGACATTAAAGTTAAGCTGCAGGGTTTGTGGCTGCGTCTTCTTAAACTAAG
AGCCACTGATGATCCACTTTCATCTTTTGAATATGGCACTATCCTTGCACTGATTGAGTCTGATGCGGAGGGGATTGGTGGTAGTGGATTTGTTGAATGT
ATAAGGGAGCATATACACTCAGGATGGCACTGTCAATTGACTGATGAGCAGTTTATTGCTGTGAAAGAACTGCTTAAAACAGCAATTAGTCGCGCAACCT
CCAGGAATGATGTGTCAACCATTAGAGATGCCCTTGAAGTATCTGCTGAGATGTATAAAAGAGATGCTAATAATGTTTCTGACTATGTCCAGCGTCATCT
TATTTCTCTGTCTATATGGGAGGAGCTACGGTTCTGGGAGGGGTACTTTGAGTATCTAATGGAGCATCCTTCAAGCAAGTCAGCCAATTATTCAGCTTTA
GTAACAACACAACTAATTCTTGTGGCATTACACATGGCTGGATTGGGACTTCTTGATACTGATGCTTGGCACATGATTGAAACAATAGCAGAGAAGAATA
ATATTGGATACAAGCAGTTTATAAAACTCAGAGGATTCTTGTCACACATCCAGCAAGTTCGTATCAGTTACTGGGGAATCTCTTCTGTAAAAGCCCAATC
CATGCGATCTCCTGGATTGTCTTCCCCACGCCCAAAAGATTCTATGGATGAGAATGAGCAACCTGCAGAAGCTTCTGTTATTGGCAGGAGCTGGGTCCAA
AGCATGTTTAGCAGAGATCCATCCAGAGCAAACTCTTTTGGTCGTGTTCGTAAAGGGGCTTCCGATGGTACTTCAGCAGTAAATGAAAATGGAACCCCTA
GAAAACAAGATTCATCTGCTGCTGGGCAGAAGAAACTTCAAACCAATGTTCGCATACTTAGAGGCCATAGTGGTGCCGTTACTGCCTTACATTGTGTTAC
CAGAAGAGAGGTCTGGGATCTAGTTGGTGACCGTGAAGATGCTGGTTTCTTTATCAGTGGAAGCACAGACTGCATGGTTAAAATCTGGGATCCTAGCATT
CGTGGTTCCGAACTTCGTGCAACTCTGAAAGGGCATACCAGAACTGTTCGAGCTATAAGTTCTGATCGAGGAAAAGTGGTATCAGGATCTGATGATCAGT
CTGTTATTGTATGGGATAAGCAAACATCCCAGCTTCTTGAAGAGCTGAAGGGCCATGATGCACAGGTTAGCTGTGTCCGTATGCTGTCTGGTGAACGTGT
CCTGACTGCTGCACATGATGGAACTGTTAAGATGTGGGATGTCCGAACTGATACTTGTGTGGCAACGGTTGGCCGTTGCTCGAGTGCAGTCCTATGCATG
GAGTATGATGATTCCACTGGAATCCTTGCTGCTGCTGGTAGAGATGCGGTTGCGAATATATGGGACATCCGTGCTGGTAGGCAGATGCACAAGCTTTTAG
GGCACACAAAATGGATTCGGTCAATTAGAATGGTTGGAGACACACTGATTACTGGTAGTGATGATTGGACTGCAAGAGTGTGGTCCGTTTCACGAGGAAC
ATGTGATGCTGTTCTAGCCTGCCATGCTGGTCCAATACTGTGTGTTGAGTATTCCATGTCAGACAGAGGAATAATAACAGGTTCAACTGATGGTCTTCTC
CGATTTTGGGAAAATGAGGAAGAAGGCATAAGATGCGTGAAGAATGTTACTATTCATACCGCTCCTATATTATCCATCAATGCTGGGGAACACTGGCTGG
GAATTGGAGCAGCTGACAATTCTATGTCCCTCTTTCACCAGCCTCAAGAGAGACTTGGTGGCTTCTCTAGCACGGGATCAAAAATGTCTGGGTGGCAACT
TTACAGAACACCTCAAAGGACAGTTGCTATGGTTAGATGTGTGGCTTCTGACCTTGAAAGGAAAAGGATATGCAGCGGTGGTCGTAATGGAGTTCTTAGG
TTATGGGAAGCGACCATCAATATTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G049500.2 pacid=42775963 polypeptide=Potri.015G049500.2.p locus=Potri.015G049500 ID=Potri.015G049500.2.v4.1 annot-version=v4.1
MAGIFEYFVVCGLGPEMRTMDGNKGYHGMRVLYLPSLLDQYPPDNHCLYPPPPPQLPTCVLPAGVEFYPSGLDANDSSTFPKSYPIVLTEGDGSKIYVSC
IAFRDPVSEDIAEAYRIPPNSFADKCICLVSRSPSFGVLRTALEELFALCFSPAGSSKPLWDVISYMVSNVPLPTPGKDRVLFAIENCLLSVEAPPKDGL
PHVEISFQPLVQCLDVDNLLKLFTAVLLERRILLRSNKYSLLTLASEAICHLIYPLRWQHVYIPLLFFSGVDYIDAPTPYMMGLHSAVDTSYLAMDGVVV
VDLEYNRICTSEEIPPIPEPELSTLRGEILKLLYPNVMGIDQMKAGLVSSSEQYFKGCNKPWGEDHDLQLRLIFLKFFASILGGYRNFIENTATHAFNTQ
AFLRKRSRSTNQPPDAMITQFLDSHGFLDYLERVIDSDENNYNLLDKLQDAIGRGQNPISVLPSSWVEPEIITISDPDVGILGSGAKFTYDRFPANIRSE
EHEEKRKQILAAASGAFDYIKHTPSSPSVQVGKDSLSPMERAAERERMVLDIKVKLQGLWLRLLKLRATDDPLSSFEYGTILALIESDAEGIGGSGFVEC
IREHIHSGWHCQLTDEQFIAVKELLKTAISRATSRNDVSTIRDALEVSAEMYKRDANNVSDYVQRHLISLSIWEELRFWEGYFEYLMEHPSSKSANYSAL
VTTQLILVALHMAGLGLLDTDAWHMIETIAEKNNIGYKQFIKLRGFLSHIQQVRISYWGISSVKAQSMRSPGLSSPRPKDSMDENEQPAEASVIGRSWVQ
SMFSRDPSRANSFGRVRKGASDGTSAVNENGTPRKQDSSAAGQKKLQTNVRILRGHSGAVTALHCVTRREVWDLVGDREDAGFFISGSTDCMVKIWDPSI
RGSELRATLKGHTRTVRAISSDRGKVVSGSDDQSVIVWDKQTSQLLEELKGHDAQVSCVRMLSGERVLTAAHDGTVKMWDVRTDTCVATVGRCSSAVLCM
EYDDSTGILAAAGRDAVANIWDIRAGRQMHKLLGHTKWIRSIRMVGDTLITGSDDWTARVWSVSRGTCDAVLACHAGPILCVEYSMSDRGIITGSTDGLL
RFWENEEEGIRCVKNVTIHTAPILSINAGEHWLGIGAADNSMSLFHQPQERLGGFSSTGSKMSGWQLYRTPQRTVAMVRCVASDLERKRICSGGRNGVLR
LWEATINI
|
DESeq2's median of ratios [POPLAR]
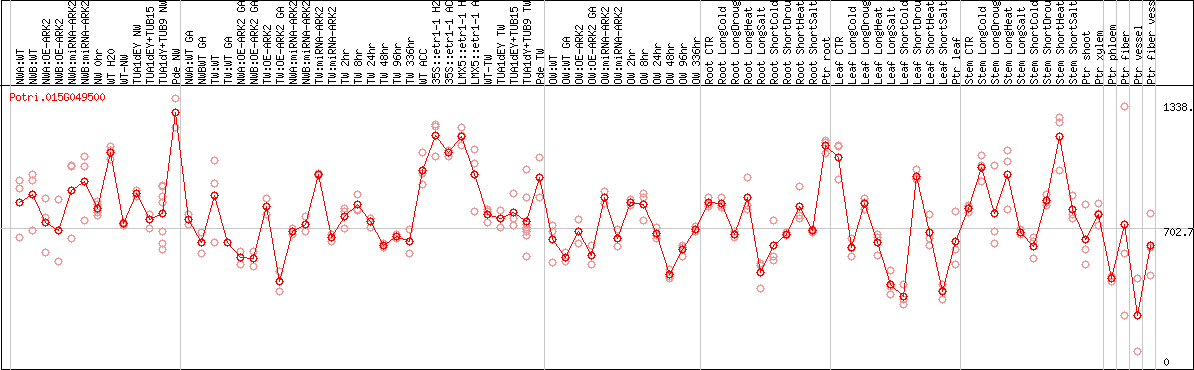
Coexpressed genes
Potri.015G049500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
