External link
Symbol
CAB4.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G47470 442 / 2e-159
CAB4, LHCA4
light-harvesting chlorophyll-protein complex I subunit A4 (.1)
AT3G61470 225 / 8e-74
LHCA2
photosystem I light harvesting complex gene 2 (.1)
AT1G45474 212 / 1e-68
LHCA5
photosystem I light harvesting complex gene 5 (.1.2)
AT1G19150 200 / 1e-63
LHCA2*1, LHCA2*1, LHCA2*1, LHCA2*1, LHCA2*1, LHCA2*1, LHCA2*1, LHCA6, LHCA2*1, LHCA2*1, LHCA2*1, LH
photosystem I light harvesting complex gene 6 (.1)
AT1G61520 161 / 2e-48
LHCA3*1, LHCA3*1, LHCA3*1
photosystem I light harvesting complex gene 3 (.1.2.3)
AT3G54890 148 / 8e-44
LHCA1
photosystem I light harvesting complex gene 1 (.1.2.3.4)
AT2G05070 116 / 2e-31
LHCB2.2
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.2 (.1)
AT2G05100 116 / 2e-31
LHCB2.3, LHCB2.1
LIGHT-HARVESTING CHLOROPHYLL B-BINDING 2, photosystem II light harvesting complex gene 2.1 (.1)
AT1G29930 116 / 3e-31
LHCB1.3, CAB140, CAB1
LIGHT-HARVESTING CHLOROPHYLL A/B-PROTEIN 1.3, CHLOROPHYLL A/B PROTEIN 140, chlorophyll A/B binding protein 1 (.1)
AT1G29920 116 / 3e-31
AB165, LHCB1.1, CAB2
LIGHT HARVESTING CHLOROPHYLL A/B-BINDING PROTEIN 1.1, chlorophyll A/B-binding protein 2 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001644
439 / 4e-158
AT3G47470 437 / 3e-157
light-harvesting chlorophyll-protein complex I subunit A4 (.1)
Lus10021663
438 / 7e-158
AT3G47470 434 / 2e-156
light-harvesting chlorophyll-protein complex I subunit A4 (.1)
Lus10011361
229 / 4e-75
AT3G61470 414 / 6e-148
photosystem I light harvesting complex gene 2 (.1)
Lus10006416
227 / 6e-74
AT3G61470 404 / 6e-144
photosystem I light harvesting complex gene 2 (.1)
Lus10021730
210 / 1e-67
AT1G45474 369 / 2e-130
photosystem I light harvesting complex gene 5 (.1.2)
Lus10042657
207 / 8e-67
AT1G45474 371 / 3e-131
photosystem I light harvesting complex gene 5 (.1.2)
Lus10039912
197 / 9e-63
AT1G19150 362 / 2e-127
photosystem I light harvesting complex gene 6 (.1)
Lus10012297
169 / 2e-51
AT1G61520 474 / 2e-171
photosystem I light harvesting complex gene 3 (.1.2.3)
Lus10016074
169 / 2e-51
AT1G61520 474 / 2e-171
photosystem I light harvesting complex gene 3 (.1.2.3)
Lus10040391
152 / 2e-45
AT3G54890 371 / 1e-131
photosystem I light harvesting complex gene 1 (.1.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00504
Chloroa_b-bind
Chlorophyll A-B binding protein
Representative CDS sequence
>Potri.015G062200.2 pacid=42776041 polypeptide=Potri.015G062200.2.p locus=Potri.015G062200 ID=Potri.015G062200.2.v4.1 annot-version=v4.1
ATGGCCACCGTCTCAACACAAGCTTCTGCTGCCGTTTTCCGGCCATGTGCTTATAAGTCGAGGTTTCTCGCTGGGGCTCCAAGTAAACTGAACAGAGAAT
TGTCCATCAAACCAATGGCATCGTCATCTCCTAGTTTCAAGGTTGAAGCCAAGAAAGGAGAGTGGTTACCTGGTTTGCCTTCTCCATCTTATCTAAATGG
CAGCCTTCCAGGTGATAATGGGTTTGACCCCCTCGGGCTTGCTGAGGACCCTGAGAACTTGAGATGGTTCGTTCAGGCTGAGCTTGTGAATGGTCGGTGG
GCCATGTTGGGCGTTGCAGGAATGCTACTGCCAGAAGTTTTCACAAAGATTGGAATCATCAATGCTCCTCAGTGGTATGATGCTGGCAAAGCAGAATACT
TTGCGTCTTCATCAACCCTCTTTGTGATCGAGTTCATACTGTTCCATTACGTCGAGATAAGAAGGTGGCAAGATATTAAGAACCCAGGATGCGTTAACCA
AGATCCCATCTTCAAACAATACAGCTTGCCTCCAAATGAATGCGGGTACCCCGGCGGCATCTTCAACCCCCTCAACTTTGCACCCACAATTGAGGCCAAA
GAGAAAGAGCTTGCCAATGGGAGATTGGCAATGTTGGCATTCTTAGGATTTGTGATTCAGCACAATGTGACCGGCAAAGGACCGTTCGACAACCTCTTGC
AGCACATCTCAGACCCGTGGCACAACACCATCGTCCAAACATTTAGCGGGCAGTAG
AA sequence
>Potri.015G062200.2 pacid=42776041 polypeptide=Potri.015G062200.2.p locus=Potri.015G062200 ID=Potri.015G062200.2.v4.1 annot-version=v4.1
MATVSTQASAAVFRPCAYKSRFLAGAPSKLNRELSIKPMASSSPSFKVEAKKGEWLPGLPSPSYLNGSLPGDNGFDPLGLAEDPENLRWFVQAELVNGRW
AMLGVAGMLLPEVFTKIGIINAPQWYDAGKAEYFASSSTLFVIEFILFHYVEIRRWQDIKNPGCVNQDPIFKQYSLPPNECGYPGGIFNPLNFAPTIEAK
EKELANGRLAMLAFLGFVIQHNVTGKGPFDNLLQHISDPWHNTIVQTFSGQ
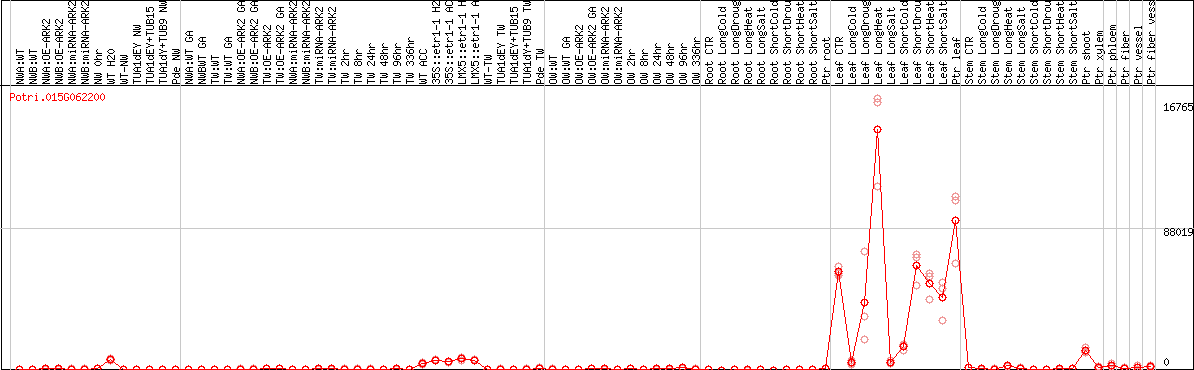
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G062200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.