External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G41340 268 / 8e-93
ATUBC4, UBC4
ubiquitin conjugating enzyme 4 (.1)
AT1G63800 266 / 7e-92
UBC5
ubiquitin-conjugating enzyme 5 (.1)
AT2G46030 257 / 1e-88
UBC6
ubiquitin-conjugating enzyme 6 (.1)
AT2G02760 97 / 5e-26
ATUBC2
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
AT1G16890 97 / 1e-25
UBC36 ,UBC13B
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
AT1G78870 97 / 1e-25
UBC35 ,UBC13A
UBIQUITIN CONJUGATING ENZYME 13A, ubiquitin-conjugating enzyme 35 (.1.2.3)
AT5G62540 96 / 2e-25
UBC3
ubiquitin-conjugating enzyme 3 (.1)
AT1G14400 96 / 3e-25
ATUBC1, UBC1
UBIQUITIN CONJUGATING ENZYME 1, ubiquitin carrier protein 1 (.1.2)
AT5G53300 94 / 1e-24
UBC10
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
AT5G41700 94 / 2e-24
ATUBC8, UBC8
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002359
301 / 3e-105
AT1G63800 275 / 5e-95
ubiquitin-conjugating enzyme 5 (.1)
Lus10042136
286 / 1e-99
AT1G63800 267 / 2e-92
ubiquitin-conjugating enzyme 5 (.1)
Lus10014763
254 / 5e-87
AT1G63800 294 / 6e-103
ubiquitin-conjugating enzyme 5 (.1)
Lus10036372
246 / 5e-84
AT1G63800 286 / 9e-100
ubiquitin-conjugating enzyme 5 (.1)
Lus10032274
239 / 1e-81
AT1G63800 278 / 5e-97
ubiquitin-conjugating enzyme 5 (.1)
Lus10024638
201 / 5e-66
AT1G63800 229 / 2e-77
ubiquitin-conjugating enzyme 5 (.1)
Lus10037203
98 / 5e-26
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10036727
98 / 5e-26
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10030786
96 / 3e-25
AT2G02760 305 / 2e-108
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10017654
96 / 4e-25
AT1G16890 311 / 5e-111
UBIQUITIN CONJUGATING ENZYME 13B, ubiquitin-conjugating enzyme 36 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.015G073400.1 pacid=42776178 polypeptide=Potri.015G073400.1.p locus=Potri.015G073400 ID=Potri.015G073400.1.v4.1 annot-version=v4.1
ATGTCTTCTCCAAGCAAGAGGAGAGAGATGGATGTCATGAAGTTGATGATGAGTGATTACAATGTGGAGACTATAAATGATGGACTGAATGATTTTAATG
TGGAGTTTCATGGTCCTAAAGAAAGCCTTTATGAAGGTGGTGTCTGGAAAATCCATGTTGAGCTTCCTGATGCATATCCTTACAAGTCGCCTTCTATTGG
GTTTGTGAACAAGATATTCCACCCAAATGTTGATGAACTTTCTGGTTCTGTATGCTTGGATGTAATCAACCAGTCATGGAGTCCAATGTTTGATCTTTTA
AATGTTTTTGAAGTTTTCTTGCCACAGCTCTTGATTTATCCAAATCCTTCAGACCCCCTCAATGGTGATGCAGCTTCATTGATGATGAAGGATAAAGAAC
AATATGATCAGAAAGTGAAAGAGTATTGTGAAAGATATGCAAAAAAGGAGCATATTATGAACTCCACGGGCGAAGAGCTGAGTGACGAGGAGGATGTCAG
TGATGAAGAAAGTGGATCCAGTGATGAGGAAGATGAGATTGCTGGGCATGTAGACCCTTAA
AA sequence
>Potri.015G073400.1 pacid=42776178 polypeptide=Potri.015G073400.1.p locus=Potri.015G073400 ID=Potri.015G073400.1.v4.1 annot-version=v4.1
MSSPSKRREMDVMKLMMSDYNVETINDGLNDFNVEFHGPKESLYEGGVWKIHVELPDAYPYKSPSIGFVNKIFHPNVDELSGSVCLDVINQSWSPMFDLL
NVFEVFLPQLLIYPNPSDPLNGDAASLMMKDKEQYDQKVKEYCERYAKKEHIMNSTGEELSDEEDVSDEESGSSDEEDEIAGHVDP
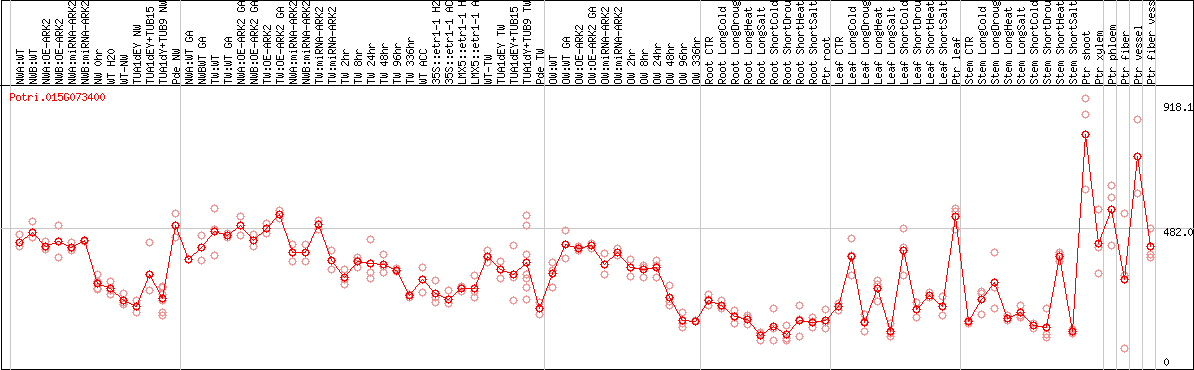
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G073400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.