External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G09590 114 / 2e-28
HSC70-5, mtHSC70-2
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
AT5G09600 99 / 9e-25
SDH3-1
succinate dehydrogenase 3-1 (.1.2.3)
AT4G32210 99 / 9e-25
SDH3-2
succinate dehydrogenase 3-2 (.1)
AT4G37910 87 / 4e-19
MTHSC70-1
mitochondrial heat shock protein 70-1 (.1)
AT5G42020 82 / 2e-17
BIP2, BIP
luminal binding protein, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
AT5G28540 81 / 4e-17
BIP1
heat shock protein 70 (Hsp 70) family protein (.1)
AT5G49910 76 / 4e-15
CPHSC70-2EATSHOCKPROTEIN70-2, HSC70-7, cpHSC70-2 ,CPHSC70-2EAT SHOCK PROTEIN 70-2
HEAT SHOCK PROTEIN 70-7, chloroplast heat shock protein 70-2 (.1)
AT1G09080 75 / 6e-15
BIP3
binding protein 3, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
AT4G24280 75 / 8e-15
CPHSC70-1
chloroplast heat shock protein 70-1 (.1)
AT5G02500 69 / 7e-13
AtHsp70-1, AT-HSC70-1, HSP70-1, HSC70-1
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042000
112 / 5e-28
AT5G09590 866 / 0.0
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
Lus10018003
112 / 9e-28
AT5G09590 1065 / 0.0
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
Lus10041999
112 / 2e-27
AT5G09590 1138 / 0.0
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
Lus10009677
110 / 4e-27
AT5G09590 1152 / 0.0
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
Lus10009039
110 / 5e-27
AT5G09590 1147 / 0.0
HEAT SHOCK COGNATE, mitochondrial HSO70 2 (.1)
Lus10017306
82 / 2e-17
AT5G28540 1169 / 0.0
heat shock protein 70 (Hsp 70) family protein (.1)
Lus10032430
82 / 2e-17
AT5G42020 1033 / 0.0
luminal binding protein, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
Lus10023043
82 / 2e-17
AT1G64550 1249 / 0.0
susceptible to coronatine-deficient Pst DC3000 5, general control non-repressible 20, ATP-binding cassette F3, general control non-repressible 3 (.1)
Lus10003202
81 / 1e-16
AT5G42020 1133 / 0.0
luminal binding protein, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
Lus10027341
77 / 2e-15
AT4G24280 1003 / 0.0
chloroplast heat shock protein 70-1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0108
Actin_ATPase
PF00012
HSP70
Hsp70 protein
CL0335
FumRed-TM
PF01127
Sdh_cyt
Succinate dehydrogenase/Fumarate reductase transmembrane subunit
Representative CDS sequence
>Potri.015G078000.1 pacid=42776014 polypeptide=Potri.015G078000.1.p locus=Potri.015G078000 ID=Potri.015G078000.1.v4.1 annot-version=v4.1
ATGGCCACCGCTGCTCTCCTCCGCTCTCTACGACGCCGTGACGTCGCCTCCGCTCCTCTTTCGTCCTACAGAAGCTTGACTAATGATGTGAGTCCAAAGT
GGGCTTCTTCTAATTTCAGCCAAAATTGGGCTGGTTTGTCAAGAGCTTTGAGTGCAAAACCTGCATGTGGAGATTTTATTGGTGTTGATCTGGGTACTGC
AAATTCATGTGCTGCTGTTATGGAAGGGAAGAATCCCAAAGTTTTTGAGAATGCTGAAGGATGTAGGATAACCCCCTCAGCTGTTACCTTCTCCCCAAAG
ATAGTAGCTGGGATCTTTGGAAAGAGCCCAAGCAAGGGAGTAAACCCAGTTGGGGCAGTGGCTATGGGAGCTGCAATTCAGGGTGGAATTTTACGTGGAG
GTGTCAAAGAATTGCGTCTCCTTGATGTTGCTCCCTTGTCACTTGGTATCAAGACACGGGTTTTCTCCACAGGGACTGACAACAAGACCCAGGTTACGGG
TATTCCGCCACCTCCCAGGGGCACGCCTCAAATTGAGGTGACCTCTGACATTGATCAACCTAATACCTCACATATCCTTCGCCCATTATCTCCTCATCTT
CCTATTTATAGCCCCCAGGTTCATTCAACATTTTCCATTGTCAATAGAATCTCCGGGGCTTACCTATCTGCTCTAGTTTTGTTTTTTTGTCTTATTTGTC
TGAAAGCGGGTTCGATTTGCTTCACCTATTATAGTTTCTACCAATTCCTCTTTTATTCATCCAAGCTGATCCTACCTGTTGTCGATGTTACCGCCGCCCT
TGCCCTGACCTATCATCTGTTTTATGGGGTTCGTCATTTACTGCATTGA
AA sequence
>Potri.015G078000.1 pacid=42776014 polypeptide=Potri.015G078000.1.p locus=Potri.015G078000 ID=Potri.015G078000.1.v4.1 annot-version=v4.1
MATAALLRSLRRRDVASAPLSSYRSLTNDVSPKWASSNFSQNWAGLSRALSAKPACGDFIGVDLGTANSCAAVMEGKNPKVFENAEGCRITPSAVTFSPK
IVAGIFGKSPSKGVNPVGAVAMGAAIQGGILRGGVKELRLLDVAPLSLGIKTRVFSTGTDNKTQVTGIPPPPRGTPQIEVTSDIDQPNTSHILRPLSPHL
PIYSPQVHSTFSIVNRISGAYLSALVLFFCLICLKAGSICFTYYSFYQFLFYSSKLILPVVDVTAALALTYHLFYGVRHLLH
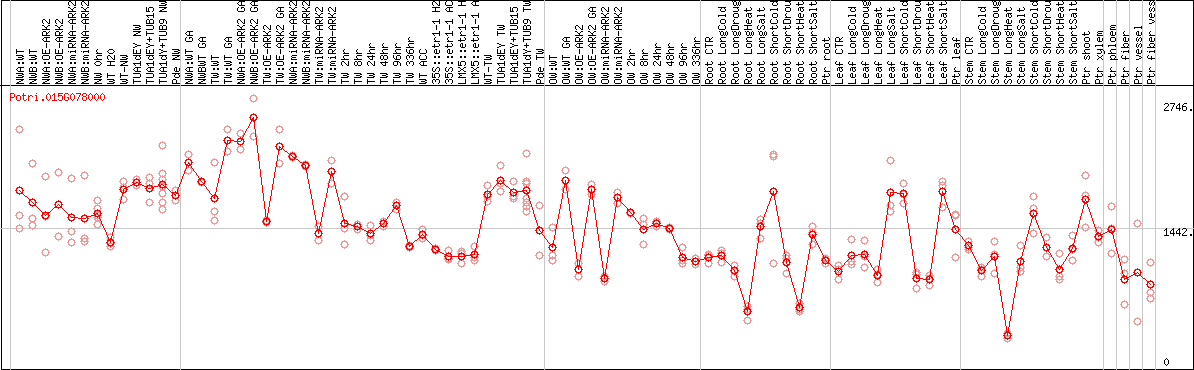
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G078000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.