Potri.015G078300 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G078300.1 pacid=42775751 polypeptide=Potri.015G078275.1.p locus=Potri.015G078300 ID=Potri.015G078300.1.v4.1 annot-version=v4.1
ATGATTCATAATGGAGAAGGGACCCACGACACCATCCCGAGGCCCACCTCTTCTGATCCGTTCGGTTCTATCAAACCGGACGGTGGTGATGCATCGCCTG
CATCAGCATCACAGCATTCCTCATGTGGAGAATCGGAGTTCGAGCGGTACTGTAGCGCCACCTCAGTTATGGGCACACCTAGTATTTGTAGTGGTTCGTT
TGGTCCTTCTTTTAATGATTGTATAAAGTCTGATGTTGAGTCCTTAAAAAGTTTGGATAATTTTAGCCTGGGGCCGAAAAGTTTCCATTTTGGGTTTGAT
GATAATAGGAACTTAGAGGATCAAAAGCTTTCAAATTCGGTTATTGATTGTTTAGATAGCAGTTTTGAAGAAAATGGGATTGATGGGTTGGAAATTCGTG
GTAGTGAAATGGACAGTAAGCGGGAGAGTGTTCGTTTGGGAATTGAGAATGGAGAGAATGATGGTTGTTCAAGTGGGTTGGATGTGGAGGTGGGTTTGGG
GTTTGATGGAGGGGAAGTGGAGAGAGGGGAAGATGGGGGTTCGTCTAGATATGGCTATTCGGAAGATGATGATTCGATGTATGGGTGTGGTTCGGATGAT
GAGAATAGGAAGAATTTGAATTTTCGGAAGACTGTGTTGCTTGGTGAGGAAGGGAAAGTTGGTGATGCGAACCCGTTGATTATGAGTTCATCTGTGGCTT
TTGGGTCGGAGGATTGGGATGATTTTGAGCTAGAAACAAGGGGAGGCATTGGAGCTTCTTTTACCTTGGATAAATTTCAGCAGCCGGAGCAGGGTCAGGA
GACTGATGGAAACTTTTTTAGTTCTACTTCTGTGGCTTTGACTGTGGCACCAGTTGTTGGTGAAACAGAAATAGGGAAAGGTTTGATGGAGGAACATGCT
GGCATCAGAGACAGTGCAGCTGATGGTTCAGGAGAGAAGCTCAATTCTGTTACTAAGGTCCCTTTTGGTGTACAGAATTCTGTCGTGGACCAAGTGGAAG
ATGTGAGAGACATTCCTGTAGCCAGTTGTCAGGTGCAAGGTGGTCATGAATTAGCCAAAGATGATAAAGGTACCTCGATCGTGCCAGTTGGTTTTCCAGG
CTATTGTGAACCACAGGAAGAAGATGTTATAAATATTTCTTTTAACTGCAATCAAGTCCAAGGTGCTAATGATACAACAGAGCTTTATAAAAATTGTCCT
GTCAGTAGTGTCTTTGAGGTGGAGCAGGAACCTCTGGTAGAAAAATCTCCCATAGGGTTAGGAATGGATTTTACAGATCATCATGTGGATGATCTGAATC
CATCTGTGAAATCCGGGGAAGTTGTTTGCACTGATGACAACGTAACTTTGGAAAATGAAGAGGCTGGGAATTTAAAAGTAGAGGCAGACCCCTTTTCTGA
TACAACCAATCAACTTTGTTCCCGCACAGCTGAATATTCAGAAAATGCAAGTGCAGAATTCATTGTGGACCAAAAACTAAATTCAACTCAGTCAATGCTT
GAAAATAACATGAAGAAAGCATCAGAAAATGCTCCTGGTTCTGTGATTCCTTACAAAGACCATCCTGCGGTGGTTAAGGCAGAGAATTTTGAGCTCATTG
AATTTTACGATGAGATTGTTAATGAAATGGAAGAAATTTTACTTGATTCTGTTGAGTCTCCAGGGGCCAGGTTTCCTCGGGGTAATCACATGTTTCAGTC
ACAGCTATTAGTGTCAACTGCTTCTACTTCTGGTACTGATGAGGCTTATATGCTAATTACACAGCCACAGAGAATTGATAGGGTTGAAGTAGTAGGTGCA
AAGCAGAAGAAGGGAGATGTCTCACTTAGCGAAAGGTTGGTTGGAGTGAAGGAATATACTGCATATATAATAAGGGTATGGAGTGGCAAAAACCAGTGGG
AGGTTGAACGCAGGTATCGTGATTTCTATACACTCTATCGTAGGCTGAAATCATTATTTGCTGACCAGGGCTGGACTCTTCCTTCTCCATGGTCCTCTGT
GGAAAAGGAATCTAGAAAAATATTCGGGAATGCATCTCCAGATGTTGTTTCTGAGAGAAGTGTTCTTATTCAGGAGTGTTTACATTCAACTATCCATTCT
GGGTTCTTTTCCAGTCCCCCAAGTGCACTGGTTTGGTTCTTGTTTCCTCGAGATTCATTTCCTAGTTCCCCAGCGGCAAGAACACTAGTTCCTCAGTCTG
TATTTTCTAACAGAGGGGAAGATGCAGGAAATATTTCCACTTTAGGGAAGACAATATCACTTATTGTTGAAATTCGGCCATTCAAATCTACTAAACAGAT
GCTAGAAGCACAACATTATACTTGTGCCGGATGCCACAACCATTTTGATGATGGGATGACATTAATGCGGGACTTTGTACAGACACTTGGGTGGGGAAAG
CCTCGACTTTGCGAGTACACTGGTCAGTTATTTTGTTCCTCATGCCATACAAATGAGACTGCAGTTTTGCCAGCAAGAGTTCTACATTATTGGGATTTTA
TTCAATACCCAGTTTCTCAGTTGGCCAAGTCATACCTGGATTCCATCCATGAACAGCCCATGCTTTGTGTAAGTGCGGTCAATCCTTTCCTATTCTCCAA
GGTCCCAGCCTTGCACCATATCATGGATGTCAGGAAAAAAATTGGAACTATGCTTTCATATGTTCGATGTCCATTCTGCAGGACCATCAATGAAGGACTC
GGTTCTCGAAGATATCTTCTTGAAGGCAATGATTTTTTTGCCCTTAGAGACCTCATAGATCTCTCAAAAGGGGCTTTTGCAGCACTACCAGTGATGGTGG
AAACTGTTTCAAGGAAAATCCTGGAGCATATAACAGAGCAATGCCTCATATGCTGTGATGTGGGAGTCCCCTGCAGTGCCCGTCAAGCTTGTAATGATCC
ATCCTCACTCATTTTTCCTTTTCAGGAAGGCGAAATTGAAAGGTGTGCTTCTTGCGAATCAGTATTCCACAAGCCCTGCTTCAGCAAGCTTACAAATTGC
TTTTGTGGGGCACACCTCAGGACAGACGAGGTCATGGAGTCAACCAGTAGCTTGAGCCGTAAGGCCAGTGGTCTCATTCTGGGAAGGAGATCGGGCTCTG
CTATGGGTCTAGGATTGTTCTCCGAATTATTTTCGAAGGCAAATCCGGAAAAAGTGAAGGATCACAAGGATAATGATGCTTTTATTTTAATGGGTTCTTT
GCCAAGCAATTTTCTTTAA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.015G078300.1 pacid=42775751 polypeptide=Potri.015G078275.1.p locus=Potri.015G078300 ID=Potri.015G078300.1.v4.1 annot-version=v4.1
MIHNGEGTHDTIPRPTSSDPFGSIKPDGGDASPASASQHSSCGESEFERYCSATSVMGTPSICSGSFGPSFNDCIKSDVESLKSLDNFSLGPKSFHFGFD
DNRNLEDQKLSNSVIDCLDSSFEENGIDGLEIRGSEMDSKRESVRLGIENGENDGCSSGLDVEVGLGFDGGEVERGEDGGSSRYGYSEDDDSMYGCGSDD
ENRKNLNFRKTVLLGEEGKVGDANPLIMSSSVAFGSEDWDDFELETRGGIGASFTLDKFQQPEQGQETDGNFFSSTSVALTVAPVVGETEIGKGLMEEHA
GIRDSAADGSGEKLNSVTKVPFGVQNSVVDQVEDVRDIPVASCQVQGGHELAKDDKGTSIVPVGFPGYCEPQEEDVINISFNCNQVQGANDTTELYKNCP
VSSVFEVEQEPLVEKSPIGLGMDFTDHHVDDLNPSVKSGEVVCTDDNVTLENEEAGNLKVEADPFSDTTNQLCSRTAEYSENASAEFIVDQKLNSTQSML
ENNMKKASENAPGSVIPYKDHPAVVKAENFELIEFYDEIVNEMEEILLDSVESPGARFPRGNHMFQSQLLVSTASTSGTDEAYMLITQPQRIDRVEVVGA
KQKKGDVSLSERLVGVKEYTAYIIRVWSGKNQWEVERRYRDFYTLYRRLKSLFADQGWTLPSPWSSVEKESRKIFGNASPDVVSERSVLIQECLHSTIHS
GFFSSPPSALVWFLFPRDSFPSSPAARTLVPQSVFSNRGEDAGNISTLGKTISLIVEIRPFKSTKQMLEAQHYTCAGCHNHFDDGMTLMRDFVQTLGWGK
PRLCEYTGQLFCSSCHTNETAVLPARVLHYWDFIQYPVSQLAKSYLDSIHEQPMLCVSAVNPFLFSKVPALHHIMDVRKKIGTMLSYVRCPFCRTINEGL
GSRRYLLEGNDFFALRDLIDLSKGAFAALPVMVETVSRKILEHITEQCLICCDVGVPCSARQACNDPSSLIFPFQEGEIERCASCESVFHKPCFSKLTNC
FCGAHLRTDEVMESTSSLSRKASGLILGRRSGSAMGLGLFSELFSKANPEKVKDHKDNDAFILMGSLPSNFL
|
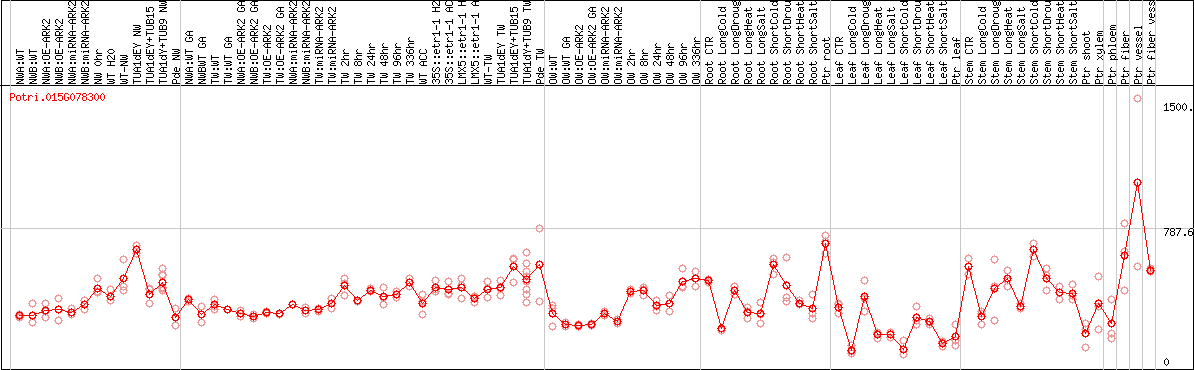
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G078300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
