Potri.015G079800 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Potri.015G079800.5 pacid=42776295 polypeptide=Potri.015G079800.5.p locus=Potri.015G079800 ID=Potri.015G079800.5.v4.1 annot-version=v4.1
ATGACATCAAGTATGTTGACCGGAGATCGCAGATATGCTCCTGCAAGAAGAGGTGGTGGCATGACAAGTTTGGGGAAAATCGCTGTTCCCAAACCAATAA
ACCTACCCAGCCAAAGGTTAGAAAATCATGGTCTGGACCCGAATGTCGAAATTGTGCCCAAGGGTACCTATAGCTGGGGAACCAGATCATCTTCTTCCAC
TCCTAATGCCTGGGGTTCTTCAACGCTGTCTCCAAATACTGATGGTGGTAGTGGTTCACCAAGCCATCTGAGTGGTCGCCCCTCATCTGGTGGAAGTGGC
ACTCGACCATCAACTGCAGGCAGTGATAGAACCCACGATCCCATTGCTAGTGCATGGGGTACAAATTCTAGGCCATCTTCAGCATCAGGAGCTTTGACAT
CGAATCAAACATCATTTACATCATTGCGTCCTTGCAGTGCAGAAACAAGACCTGGTAGCTCGCAGTTATCCCGTTTCGCTGAGCCCTTATCTGATAATTC
AGTTGCTTGGGTTGCAACTGGAACGGCAGAAAAGTTGGGAGGGACATCCTCTAAGAATGAGGGGTTTTCTTTGACTTCAGGAGATTTTCCAACACTTGGT
TCAGAGAAAGAAAACTCTGGAAAAAACACGGAGTCCCAAGACCATGATTCTTACAGTCGTCCTGGTTCATCATCGGGTGGAGTAGCTCCAGGGAAAGAGA
GTGCTGAAAATTCTGCAGGTGATGCTTCTATAAACACAAATGCAAAGATGGAACCTGCAAATTCTTGGAGAAGAGAGAATCCCATGTGTGGTGAAGATGG
GTTGAGGCCTAGCATGGAGAAGTGGCACCCTGATCACCAATTGTATCCTAATTCTAATATTCGCCCACAGAATTATGATTCTTGGCATGGCCCCCCTGTA
AATAATCCTCCAGGTGGTGTTTGGTACAGAGGACCTCCAGGTGGTCCACCTTTTGCACCGCCAATAGCACCTGGTGGCTTTCCCATGGAGCCATTTCCTT
ATTACTGTCCGCAGATTCCACCTACTGCTCTTGCCAACCCACAACAAGGTCCTCCACCAGGACCTGGACCAAGAGGACCCCATCCAACAAATGGAGATAT
GTACAGACCCCACATGCATGATGCTTTTATGCGTCCAGGCATGCCATTTCGTCCTGGCTTTTATCCTGGACCAGTTCCTTACGAGGGCTATTACGCTTCT
CACATGGGCTACTGCAACTCCAATGATCGAGATATTCAGTTTATGGGAATGGCAGTGGGTCCTGCTCCATATAACAGATTTTCAGGTCAAAATGCTCCTG
ATCCTGCAAATTCCCATGGAAGGCCAGCTGGATATGGTCCTCCTAGTGGTCACACAATGGTTCCAGAACAACTCGAATCTGGTCATCCACAAGATACTCG
TGGACCATTCAAAGTCCTTCTGAAGCAGCATGATGGTTTAGAGGGAAAAGACAAAGAACAGAAGTGGGATGACATGATGGCAACCAATGCATCTTATCCT
GGGAAGGCAGGCCATCAAAGAAAGTCATCATGGGAGAATGGTTGGAGTGCAGATGAGAAGAACAATAAAGAGAGGAATACAAGAAGAATTGGCGAAGAAT
TTTCTTCCGAGGCCAATGGCAACCAAGGAGGTGTTAAAGTAAAACCTCTTGAACATGTAGGAAATTGGAAGGCAGCTGATGACAGTTCAGTTAAGAAACT
GGAACCTGCAGCATCTGGTTTTCCAGAAGTTTCAACCGCCCCAAAGGATCCTAGTCTAATCAGGAAAATAGAAGGTTTAAATGCGAAGGCCAGGGCCTCT
GATGGAAGGCAAGAAGTTAAATTTTCTTCCAGTAGGGAGGAGCATAAGAATAGATTACAAGGTGGTAATGCCAGGTCTAATCATTCTGCAAATGAAGCTG
GTAATAGTTATGCATCTCTTGAAAGAACTCATGTCTGTGGCATTAGCGACACTGCTTCCCATGAAGACCGCATTTCTGCTGCAGATAAGAGCCATGAGGT
AACAGATGCTATCGGAACAGCAAGCTCCAGGAGATCAACTCATGGTATGCATGGGAGACCTGATCATCATGGTAAAGGGAGATTCAGTACCCAAGAGGCT
GAAGGGTGGCGGAGAAGATCTCATGTTGCAGACTTGTCGAGTGTATTGTCGTCTTCTCATTTTGAAAGTTCTAATGTTCATAGGCAAGACCATAGTCCTG
CAGAGGCAACTGAAAAGTCTGGGTCATATCATCAAGGGAAGGATGATGGAGAATCTGTGCTACCTCACCCTGATCCCAGTGATAGCCAGGTGCAACGTGC
TAAGATGAAAGAGCTAGCCATTCAACGGGTTAAACAACGCGAGAAGGAAGAGGAGGAACGTGCTAGAGATCAAAAGGCCAAGGCACTTGCAAAATTGGCA
GAGTTGAATAAGCGGACTAAGGCAGCGGAGAGTTTAAGCGAGGTTCTGCCAGGGATGCCGAAGGCTACCCACAAGGAATCTGTTGTGATACATGACCAAT
TGGAGCCATTGCAACAGGATGTTAGTCGTGCTGATGGTGATCACCCTGATAATGCCCCACAGACTTATGACAATAGAGCTTCTAAGCAGAAGCGTGTGAG
CTACAGACAAAAGCAGAATGGTCCCTTGGAAAAGACTTGCAATGATAAGTTAATGACTTCTATAATTGAGGCACCAAAAAATGTTACTGATGTAGCAGCC
AATGCTCCTGTTTCTATCGAGGGTGCAACTGAAATGACCACAAGCCCTGAATCGACTTTGCCCATCAACCCAACTGCCACGACCGAGTCTTCAGTACATC
ATGGTAGAAGAAAGAACAGAAATGGAAAGAACAAGTACAAAGTGGAGGAGGCATCATCTATGGCAGTAGTTGTAACGCCAACCCTATCAAAAGAAATAAC
TGCTTTAGACATTTCAGTTGAAAGTAGCAAGTCCAAGGCTTCTGAATCTGTGTCAGATCCAAGCTCCCAGACTGATTCTAGAGATGGAAATCAGTCTTTG
GATCATCGCACATCATCACCAAATGAAGAAGTTCAAGGCAGAGTAAATAATCAGTGGAAATCTCAATATTCTCGCAGGATGCCGAGGAATCCACAAGCTA
ATAAATCAACTGAGAAATTCCAAAGTGGTGATGCTGTTATCTGGGCTCCTGTGCGGTCACACAATAAAATTGAGGCTACTGATGAAGCCAGTCAGAAGAC
TCTGGCTGATGCCATTAGTGAACCCATGAAGAGTGATCAGCAAGTGCAGAATAATACAAGAAATAAACGGGCAGAAATGGAAAGATACATTCCAAAGTCT
GTAGCCAAAGAAATGGCTCAGCAAGGAAGTAGTCCCCATTCAGCGGCGCCATTAATCAATCAGATTACTCCAGATGAGACTGCTGGAAGACCTGAATCTC
GTTCTCTGGGCAATGAAAGTTCTCAAAGCCCTGCCACAGGTATGGGAAAGGTAGTATCCATTTTAGAATCTAAGAATGGGGATGGCAGGCAAAATAAGTC
AGGAAAAAGGAATGGATCATGGCGCCAACGGGGTTCGTCTGAATCAACTATGTTTTTTACAAGTAAAAATGTTCAAAAATCAATTGAGCATCAAGTCCAG
AAACCTGATGTAAGTTCAGTGAAAGAGCAACTGGGTCATTATGATGAATGGAGCGACTCTGATGGTTGGAACATTCCTGAAAAATCTGAGGTGCCCATCA
CTGTCCCTGCTATAAAAGATCATGGTGCAACAGCCAGAGCTAGGCGGCCATCATACAGGGGACATAAAAGTAGCCATGATCCTGATGAGAGGAGAATTCA
TACTGGGGACGCTGAAAAGGTTCATGTCCAAACCTTAGGTTCAGAGATGCATCAAGCAGACTCAGCTGCAACTTCAAAAGAAAATCGTGCTGTTGGGGAA
CGACCAGCATCTCATTGGCAACCCAAATCCCAGGCAATTTCAGCAACTACTAATCCAGGAAGCAGGGCCAGTGGTGGTCAGAACACAGGCTCTGAAGTTG
GTAGGGGTAATAAAAAGGATTCCACTTCTCAAAATGGTATGCCAGTTCTGCCTCAACCTGACAAGGATATTGCTGCAGAGGCCCAGTCCCATCCTGATGG
CTCTCTGTCTGCAAGGAGCAATTTGGAAGAAGATCCTAGCACTGGGCATCAAGAGGTTAAGAAGGAAAGGAAGATAGCATCACATAAAGGACATCCGGCT
GAACCATCTCCTTTGAATATGGATTTTCAACAACGGGTGTCTTCAGGGTTCCGAAAAAATGGAAACCAAAACAGCCGCTTTGGCAGAGAGCATGATTCTC
GTGGTGGCGAGTGGAGTGGTCCTGGGAAAGATAACGAGCACCATAACCGTGAAAGGCAGAGACAGAATTCACATTATGAGTACCAGCCAGTAGGGCCTCA
ATATAATAACAAAGCTAACAATTATGAATCTTCAAAAGATGGTTCTCATAATTCTGTTGCAAGGTCCAGGGAGAGAGGTCAGAGTCACTCAAGGCGTGGT
GGGGGGAATTCTCATGGGCGGCAACCTGGTGGTGCTCGAGGTGATGCCAATTATGATTAG
|
|||||||||||||||
|
AA sequence
|
>Potri.015G079800.5 pacid=42776295 polypeptide=Potri.015G079800.5.p locus=Potri.015G079800 ID=Potri.015G079800.5.v4.1 annot-version=v4.1
MTSSMLTGDRRYAPARRGGGMTSLGKIAVPKPINLPSQRLENHGLDPNVEIVPKGTYSWGTRSSSSTPNAWGSSTLSPNTDGGSGSPSHLSGRPSSGGSG
TRPSTAGSDRTHDPIASAWGTNSRPSSASGALTSNQTSFTSLRPCSAETRPGSSQLSRFAEPLSDNSVAWVATGTAEKLGGTSSKNEGFSLTSGDFPTLG
SEKENSGKNTESQDHDSYSRPGSSSGGVAPGKESAENSAGDASINTNAKMEPANSWRRENPMCGEDGLRPSMEKWHPDHQLYPNSNIRPQNYDSWHGPPV
NNPPGGVWYRGPPGGPPFAPPIAPGGFPMEPFPYYCPQIPPTALANPQQGPPPGPGPRGPHPTNGDMYRPHMHDAFMRPGMPFRPGFYPGPVPYEGYYAS
HMGYCNSNDRDIQFMGMAVGPAPYNRFSGQNAPDPANSHGRPAGYGPPSGHTMVPEQLESGHPQDTRGPFKVLLKQHDGLEGKDKEQKWDDMMATNASYP
GKAGHQRKSSWENGWSADEKNNKERNTRRIGEEFSSEANGNQGGVKVKPLEHVGNWKAADDSSVKKLEPAASGFPEVSTAPKDPSLIRKIEGLNAKARAS
DGRQEVKFSSSREEHKNRLQGGNARSNHSANEAGNSYASLERTHVCGISDTASHEDRISAADKSHEVTDAIGTASSRRSTHGMHGRPDHHGKGRFSTQEA
EGWRRRSHVADLSSVLSSSHFESSNVHRQDHSPAEATEKSGSYHQGKDDGESVLPHPDPSDSQVQRAKMKELAIQRVKQREKEEEERARDQKAKALAKLA
ELNKRTKAAESLSEVLPGMPKATHKESVVIHDQLEPLQQDVSRADGDHPDNAPQTYDNRASKQKRVSYRQKQNGPLEKTCNDKLMTSIIEAPKNVTDVAA
NAPVSIEGATEMTTSPESTLPINPTATTESSVHHGRRKNRNGKNKYKVEEASSMAVVVTPTLSKEITALDISVESSKSKASESVSDPSSQTDSRDGNQSL
DHRTSSPNEEVQGRVNNQWKSQYSRRMPRNPQANKSTEKFQSGDAVIWAPVRSHNKIEATDEASQKTLADAISEPMKSDQQVQNNTRNKRAEMERYIPKS
VAKEMAQQGSSPHSAAPLINQITPDETAGRPESRSLGNESSQSPATGMGKVVSILESKNGDGRQNKSGKRNGSWRQRGSSESTMFFTSKNVQKSIEHQVQ
KPDVSSVKEQLGHYDEWSDSDGWNIPEKSEVPITVPAIKDHGATARARRPSYRGHKSSHDPDERRIHTGDAEKVHVQTLGSEMHQADSAATSKENRAVGE
RPASHWQPKSQAISATTNPGSRASGGQNTGSEVGRGNKKDSTSQNGMPVLPQPDKDIAAEAQSHPDGSLSARSNLEEDPSTGHQEVKKERKIASHKGHPA
EPSPLNMDFQQRVSSGFRKNGNQNSRFGREHDSRGGEWSGPGKDNEHHNRERQRQNSHYEYQPVGPQYNNKANNYESSKDGSHNSVARSRERGQSHSRRG
GGNSHGRQPGGARGDANYD
|
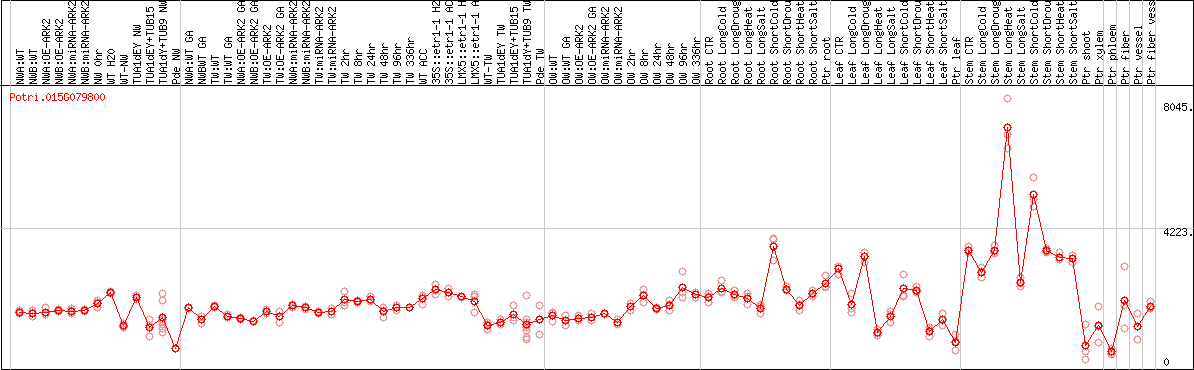
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G079800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
