Potri.015G086400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G086400.1 pacid=42774938 polypeptide=Potri.015G086400.1.p locus=Potri.015G086400 ID=Potri.015G086400.1.v4.1 annot-version=v4.1
ATGACAATCAAAGCCACGCCTATCATCAAAGATGGCTGCCTCATGGTGAGAGGCAAGGTGGTTCTGAGCAGAGTTCCACAAAACATTCTTGTTTCTCCGG
CAAGCAATGGCTCTGCTTTCTTTGGTGCTACATCTCCATCTCCTAGTTCTCGCCATGTGTTTAGCCTTGGAGTTCTTGAGAAGTACAGGTTCTTGTGCCT
TTTTAGAGTCAAAATCTGGTGGATGATACCTCGTGTAGGGAAATCCGGTAGTGAAATTCCCATGGAAACTCAAATGCTTTTGTTGGAAGCAACAGAGGAA
TCTGCCCTGAATGACGAGGTGAATTCTTCTGAAACATCGACAGATAACACTTTCTACATTCTTTTCTTGCCTGTCTTGGATGGACTATTTCGATCTAGTT
TGCAAGGAACTTCGGAAAATGAGCTCCACTTCTGCGTTGAAAGCGGAGATGCTAATGTCCAAACCTCACAAGCACTTGAAGCAGTATTTGTGAACTCAGG
GGAGAACCCTTTTGAGCTTATTAAGAATTCTGTCAAGATACTGGAGCAACACAAGGGGACCTTTTGCCACATTGAGAACAAGAAGATTCCTGCACATTTA
GACTGGTTTGGCTGGTGTACTTGGGATGCTTTTTACACTCAAGTAAATCCACAAGGAATCAAAGAGGGTCTTCAAAGTTTCTTAGAAGGTGGGTGTTCAC
CGAAGTTTCTTATTATTGATGATGGATGGCAAGACACAGTGAATGAATTTCGCAAGGAAGGTGAACCTCTTATCGAAGGGACACAGTTTGCTACTAGATT
GGTAGATATAAAAGAAAATGGAAAATTCAGAAGTTCAGGTCCAGATGAAGGCTGCACCGATTTGCATGAATTTATTGATACCATCAAAGAGAAATATGGA
TTAAAATTTGTTTATATGTGGCACGCTTTAGCTGGTTATTGGGGAGGGGTTCTTCCATCATCTGATTCAATGAAAAAGTACAACCCGAAACTTGTATATC
CAATTCAGTCACCTGGTAATGTGGGGAACATGAGAGACATTGCAATGGACAGCTTGGAGAAATATGGAGTTGGAGTCATTGATCCCAGTAAGATATTTGA
TTTCTACAATGATCTTCATAGCTATCTGGCAAGCAATGGTGTAGATGGAGTCAAGGTGGACGTCCAGAATCTGATTGAAACTTTAGGTTCTGGATGTGGA
GGTCGAGTGACGTTGACTAGACAGTATCAGGAAGCACTTGAGAGATCTATTTCAAGAAACTTTAAAGAAAATAATCTAATTTGTTGCATGAGTCACAACT
CTGACTCAATTTACAGTTCGAAGAGAAGTGCAATTGCAAGAGCATCAGAGGATTTCATGCCAAGAGAGCCAACATTTCAGACCTTGCACATTGCGTCTGT
AGCTTTCAACAGCTTTCTTCTTGGGGAGATTGTTGTTCCAGACTGGGACATGTTCCATAGCAAACATGATACCGCTGATTTCCATGGTGCTGCAAGAGCA
TTGGGTGGATGTGCAGTATATGTAAGTGACAAGCCAGGCATTCATGACTTCAAAATCCTTAAGAAGCTCGTGTTGCCTGACGGGTCTATCCTGAGAGCTA
GACATGCTGGCCGTCCCACAAGAGATTGCTTATTTGAAGATCCTGTCATGGATGCGAAAAGTTTGTTGAAGATATGGAACTTGAACAAGTTAACTGGGGT
CATCGGTGTTTTTAATTGCCAAGGAGCAGGAAGTTGGCCAATGAAACAAGAAGCTGAAGAAATACCTACTGTGCCTTCCGGGCCTTCATCCCTTTCAGGT
CATGTGAGTCCTATTGATGTTGAATTTCTCGACGATATTGCCGGTGAAGATTGGAATGGAGATTGTGCAATCTATGCTTTCAATTCAGGATCTCTTTCCA
TGCTGCCAAAGAAAGGGATTCTTGAGGTGTCCCTGACCACTCTAAAATACGAGATATATACCATCTCTCCAATTAAGGTTTTTGGTCAGAATCTTCAGTT
TTCTCCGATAGGATTGCTTGACATGTACAACTCAGGAGGAGCAGTGGAAGCTGTGAACTGCATTATCGATGTTTCATCTTACACAATCAAAGTCAATGGC
CGAGGCGGTGGCCGATTTGGAGCCTACTCCAACACTAAACCAACCTTTTGCAGGGTGGACATGAAAGAAGAAGAGTTCACTTACAACGACAAAAATGGAT
TATTGATAGTCAAACTTGAATGCACAGGCAATTTAAGAGAGATTGAATTCATATATTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G086400.1 pacid=42774938 polypeptide=Potri.015G086400.1.p locus=Potri.015G086400 ID=Potri.015G086400.1.v4.1 annot-version=v4.1
MTIKATPIIKDGCLMVRGKVVLSRVPQNILVSPASNGSAFFGATSPSPSSRHVFSLGVLEKYRFLCLFRVKIWWMIPRVGKSGSEIPMETQMLLLEATEE
SALNDEVNSSETSTDNTFYILFLPVLDGLFRSSLQGTSENELHFCVESGDANVQTSQALEAVFVNSGENPFELIKNSVKILEQHKGTFCHIENKKIPAHL
DWFGWCTWDAFYTQVNPQGIKEGLQSFLEGGCSPKFLIIDDGWQDTVNEFRKEGEPLIEGTQFATRLVDIKENGKFRSSGPDEGCTDLHEFIDTIKEKYG
LKFVYMWHALAGYWGGVLPSSDSMKKYNPKLVYPIQSPGNVGNMRDIAMDSLEKYGVGVIDPSKIFDFYNDLHSYLASNGVDGVKVDVQNLIETLGSGCG
GRVTLTRQYQEALERSISRNFKENNLICCMSHNSDSIYSSKRSAIARASEDFMPREPTFQTLHIASVAFNSFLLGEIVVPDWDMFHSKHDTADFHGAARA
LGGCAVYVSDKPGIHDFKILKKLVLPDGSILRARHAGRPTRDCLFEDPVMDAKSLLKIWNLNKLTGVIGVFNCQGAGSWPMKQEAEEIPTVPSGPSSLSG
HVSPIDVEFLDDIAGEDWNGDCAIYAFNSGSLSMLPKKGILEVSLTTLKYEIYTISPIKVFGQNLQFSPIGLLDMYNSGGAVEAVNCIIDVSSYTIKVNG
RGGGRFGAYSNTKPTFCRVDMKEEEFTYNDKNGLLIVKLECTGNLREIEFIY
|
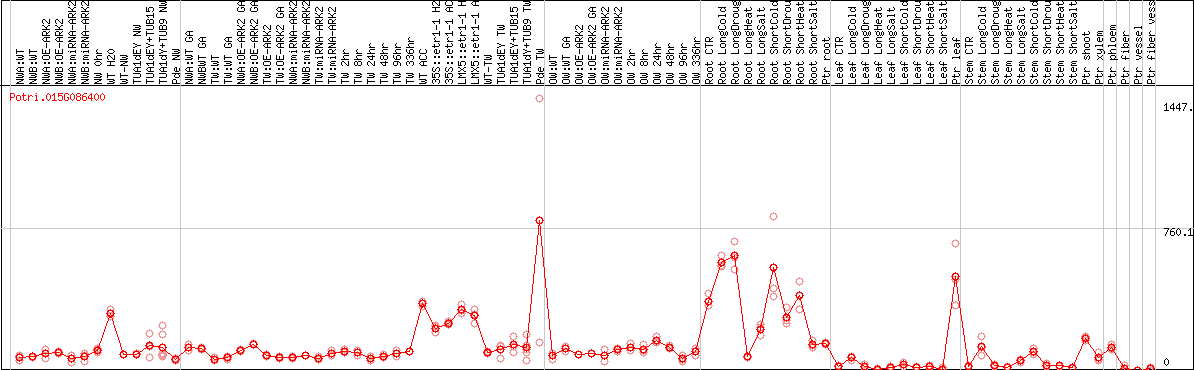
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G086400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
