External link
Symbol
CTRNP.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G24770 290 / 4e-97
CP31, ATRBP33, ATRBP31, RBP31
ARABIDOPSIS THALIANA RNA BINDING PROTEIN, APPROXIMATELY 31 KD, 31-kDa RNA binding protein (.1)
AT5G50250 276 / 3e-92
CP31B
chloroplast RNA-binding protein 31B (.1)
AT2G37220 172 / 1e-51
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G52380 150 / 1e-42
CP33, PDE322
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
AT1G60000 139 / 2e-39
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT3G52150 122 / 1e-32
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT3G53460 102 / 1e-24
CP29
chloroplast RNA-binding protein 29 (.1.2.3.4)
AT3G23830 97 / 2e-24
AtGRP4, GR-RBP4, GRP4
glycine-rich RNA-binding protein 4 (.1.2)
AT2G35410 97 / 4e-23
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G13850 89 / 2e-21
ATGRP2, GR-RBP2
glycine rich protein 2, glycine-rich RNA-binding protein 2 (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G090200
451 / 2e-160
AT4G24770 310 / 7e-105
ARABIDOPSIS THALIANA RNA BINDING PROTEIN, APPROXIMATELY 31 KD, 31-kDa RNA binding protein (.1)
Potri.001G340800
277 / 8e-93
AT4G24770 322 / 2e-110
ARABIDOPSIS THALIANA RNA BINDING PROTEIN, APPROXIMATELY 31 KD, 31-kDa RNA binding protein (.1)
Potri.016G090700
180 / 7e-55
AT2G37220 302 / 4e-103
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.016G068300
149 / 3e-42
AT3G52380 264 / 5e-87
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
Potri.008G172100
147 / 3e-42
AT1G60000 288 / 5e-98
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.010G065600
146 / 8e-42
AT1G60000 287 / 9e-98
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.006G127200
143 / 2e-40
AT2G37220 269 / 5e-90
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.009G065900
113 / 3e-29
AT3G52150 237 / 2e-78
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.001G141300
108 / 2e-27
AT2G35410 248 / 2e-81
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017962
249 / 3e-81
AT5G50250 274 / 5e-92
chloroplast RNA-binding protein 31B (.1)
Lus10041952
231 / 6e-74
AT5G50250 272 / 2e-90
chloroplast RNA-binding protein 31B (.1)
Lus10002222
191 / 6e-59
AT3G53460 263 / 1e-87
chloroplast RNA-binding protein 29 (.1.2.3.4)
Lus10023191
190 / 2e-58
AT3G53460 263 / 2e-87
chloroplast RNA-binding protein 29 (.1.2.3.4)
Lus10026514
173 / 4e-52
AT2G37220 284 / 4e-96
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10013801
149 / 1e-40
AT3G53460 258 / 2e-82
chloroplast RNA-binding protein 29 (.1.2.3.4)
Lus10029372
143 / 4e-40
AT3G52380 269 / 6e-89
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
Lus10016174
141 / 2e-39
AT3G52380 275 / 3e-91
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
Lus10023723
135 / 5e-37
AT1G60000 259 / 1e-85
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10035678
99 / 1e-23
AT2G35410 231 / 2e-74
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.015G086500.1 pacid=42775901 polypeptide=Potri.015G086500.1.p locus=Potri.015G086500 ID=Potri.015G086500.1.v4.1 annot-version=v4.1
ATGTCTGCCACAGCTGCTTCCACCTTGAAGCCTTTGCTAATGGCAGAAACATGCATATGTTCATTCCCTTCAATTTTCACTTCAAAACCCCCACTAAAAC
CTCTCCCTATCTCACATAGACCCATCAAGCTCCAACTCTCATACTCTCACTCTTTATCAACGTTATCTGTAAAACCCAAAACCCATTTATCTTTAACAAT
CCCTTTTGTGGCCCAGACCTCAGATTGGGCTCAACAAGAAGAAGAAAACAACACCACTATCACTTTAACTGAGTCAGAACAAGAAGACTCCATTTTGGAA
AATGAAGAAAGTAATGATTTTGAAGGTAGAGTATCCGATTGGGAGGCTGAAGGAGAGGATGCAGCTGCTTCCGAAACCGAAGCTGTTAGAGGTGAAGGAG
AGAGAGGTGACGAGGAGGGGTTTGTGGAGCCACCGGAGGAAGCTAAGATTTATGTAGGGAATTTGCCTTATGATGTTACTAGTGAGAAGTTGGCTATGCT
ATTTGATCAAGCTGGAACTGTTGAGATTTCTGAGGTTATTTACAACACGGAAACTGATACAAGTCGTGGCTTTGGGTTTGTGACGATGAGTACCGTTGAA
GAGTCTGACAAGGCTATAGAAATGTTCAATCGTTATAACTTAGATGGAAGGCTCTTGACTGTAAACAAGGCTGCTCCTAGAGGATCAAGGCCAGAACGCC
CTCCTCGTGTGTCTGAACCCAGCTATAGAATCTATGTGGGCAACCTACCATGGGGAGTGGATAGTGGTCGTCTTGAGGAAGTCTTTAGCGAGCATGGTAA
AGTTGTGAGTGCTCAGGTTGTTTCTGACTGGGAGACTGGCCGTTCACGTGGTTTTGGCTTTGTAACTATGTCCTCAGAGAGCGAGTTGAATGATGCCATA
GCCGCACTTGATGGACAGGAACTGGACGGGAGAGCAATTAGAGTAAATGTTGCTGCGGAAAGACCAAGGCGCAGCTCCTTTTGA
AA sequence
>Potri.015G086500.1 pacid=42775901 polypeptide=Potri.015G086500.1.p locus=Potri.015G086500 ID=Potri.015G086500.1.v4.1 annot-version=v4.1
MSATAASTLKPLLMAETCICSFPSIFTSKPPLKPLPISHRPIKLQLSYSHSLSTLSVKPKTHLSLTIPFVAQTSDWAQQEEENNTTITLTESEQEDSILE
NEESNDFEGRVSDWEAEGEDAAASETEAVRGEGERGDEEGFVEPPEEAKIYVGNLPYDVTSEKLAMLFDQAGTVEISEVIYNTETDTSRGFGFVTMSTVE
ESDKAIEMFNRYNLDGRLLTVNKAAPRGSRPERPPRVSEPSYRIYVGNLPWGVDSGRLEEVFSEHGKVVSAQVVSDWETGRSRGFGFVTMSSESELNDAI
AALDGQELDGRAIRVNVAAERPRRSSF
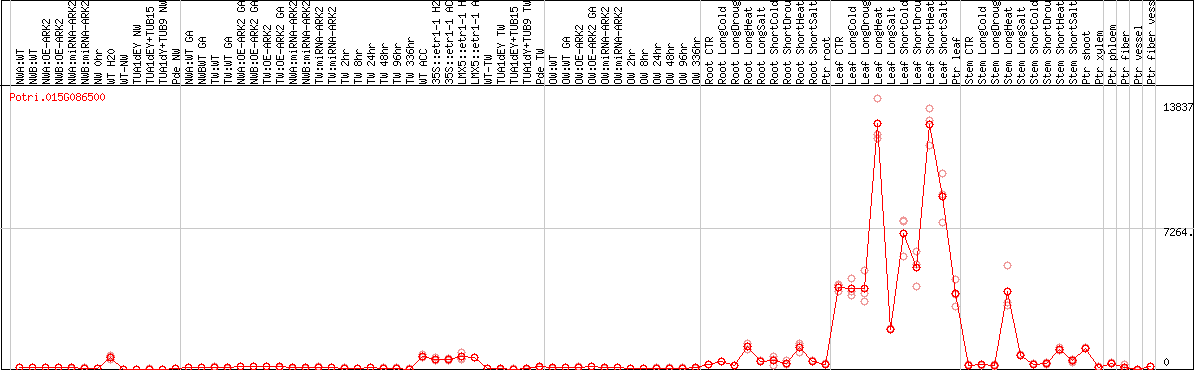
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G086500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.