ATGSL08.2 (Potri.015G089300) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ATGSL08.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G089300.2 pacid=42775274 polypeptide=Potri.015G089300.2.p locus=Potri.015G089300 ID=Potri.015G089300.2.v4.1 annot-version=v4.1
ATGTCTCGGGTTAGCAATAACTGGGAGAGGTTAGTGAGGGCGACGTTGAAGCGAGAACTCGGTCAAGGTCATGAGAGAATGTCCAGTGGCATCGCCGGAG
CTGTGCCAGTTTCGCTTGGCCGTACGACCAATATTGACGCGATTTTGCAGGCTGCCGATGAGATTCAAGATGAGGACCCTAACGTTGCCAGAATTTTGTG
TGAGCAAGCATATAGCATGGCACAAAATTTGGACCCAAGCAGTGATGGTCGTGGTGTTCTCCAATTCAAGACCGGTTTGATGTCTGTTATTAAGCAAAAA
CTTGCCAAAAGGGACGGTGCCCGTATAGATCGCAATCGGGATATTGAACACCTGTGGGAGTTTTACCAGCACTACAAGAGACGTCATAGAGTGGATGACA
TCCAAAGAGAGGAACAAAAGTTCCGGGAATCTGGAAATTTTTCTACTGTTATTCGGGGAGAGTTTGAGCTAAGTTCTCTAGAGATGAAAAAGGTATTTGC
TACCTTGAGGGCCTTAGAAGATGTCATGGAAGCAGTCAGTAAAGATGCAGATCCTCATGGGGCTGGCAGACATATCATGGAGGAGCTACAAAGAATTAAA
ACGGTTGGGGAGCTCACATCGTATAATATTGTCCCATTGGAAGCACCTTCATTAAGTAATGCTATTGGAGTTTTCCCTGAAGTTCGAGGCGCAATGTCTG
CCATTAGATATGCTGAGCATTACCCCAGACTTCCTGCTGGTTTTGTGATTTCTGGAGAACGTGATTTAGATATGTTTGATCTTCTTGAATATGTCTTTGG
TTTTCAGAATGATAATGTAAGGAATCAGCGTGAAAATGTTGTTCTTGCAATAGCAAATGCACAGTCTCGACTTGGCATACCCATCCAAGCAGATCCAAAA
ATTGACGAGAAAGCTATCAATGAGGTTTTCTTGAAGGTGTTGGATAACTACATCAAATGGTGCAAATACTTGCGAAAACGCCTTGCTTGGAATAGTATTG
AAGCGATTAATAGAGACAGGAAGCTTTTTCTGGTATCACTATATTATCTTATATGGGGTGAAGCTGCAAATGTTCGTTTTCTCCCAGAGTGCATTTGCTA
CATATTTCATCATATGGCCAAAGAGTTGGATGCAATATTGGATCATGGTGAAGCTAATCATGCCGCCAGCTGTATAACAGAAAGTGGTTCTGTGTCTTTC
CTTGAGCAAATCATTTGTCCTATATACCAGACAATAGCAGCGGAAGCTGAGAGGAATAATAATGGAAAAGCTGTGCATTCAGCTTGGAGGAATTATGACG
ACTTCAATGAATATTTTTGGTCCCCTGCTTGCTTTGAGTTGAGCTGGCCGATGAAAGAAAATTCCTCATTCTTGTTGAAGCCTAAAAAGTCAAAAAGGAC
TGGTAAAAGCACGTTTGTTGAGCATCGCACATTTCTTCACATATATCGAAGTTTCCATCGGCTCTGGATCTTTTTAGCTCTGATGTTCCAGGCTTTGGCA
ATTATAGCCTTCAACCATGGAGATTTGAGTTTAGATACCTTTAAAGAAATGCTCAGCGTTGGGCCCTCATTTGCCATCATGAACTTCATAGAGAGTTGTT
TAGATGTGCTGCTTATGTTTGGCGCTTACTCCACAGCGAGAGGCATGGCTATCTCAAGACTGGTTATTAGGTTTTTCTGGTGTGGCTTGAGCTCGGTGTT
CGTAACATACTTATATGTGAAGGTTCTGGAGGAAAAAAATCGACAGAACTCGGATTCGTTCCATTTCCGTATATATATACTTGTTCTGGGTGTTTATGCA
GCTTTGCGCCTCTTCCTTGCTCTGTTGCTAAAGTTTCCAGCTTGTCATGCTCTGTCTGATATGTCTGATCAATCATTCTTCCAATTTTTCAAGTGGATTT
ATCAGGAAAGATACTATGTTGGTCGTGGTCTTTTTGAGAAGATGAGTGATTACTGCAGATATGTCCTCTATTGGTTGGTGATCTTTGCGTGCAAATTTAC
ATTTGCTTACTTTCTCCAGCAGATTAGACCCCTTGTCAAACCAACCAATACCATCAGGGCACTTCCATCTTTACCGTACTCATGGCATGATTTGATCTCA
AAGAATAATAACAATGTCTTGACCATAGCTAGCCTATGGGCTCCTGTTGTGGCGATCTATATCATGGACATTCACATATGGTACACTATCTTATCAGCAA
TTGTTGGGGGCGTGATGGGTGCACGAGCTCGTTTAGGCGAGATACGCAGTATTGAAATGGTTCATAAACGTTTTGAGAGCTTCCCAGCGGCCTTTGTCAA
GAATCTTGTCTCTCCACAAGCACAAAGGATGGTGCTCAACAGACATGCATCTCAAGAAGCTCAGGATATGAACAAAGCATATGCTGCACTGTTTGCTCCT
TTCTGGAATGAGATAATCAAAAGCTTACGCGAGGAAGATTATATCAGCAATAGGGAGATGGACTTGCTTTCTATTCCGAGCAACACAGGAAGTCTAAGAT
TAGTTCAATGGCCACTGTTTCTTCTTAGCAGCAAGATTTTATTGGCAGTGGATTTGGCTTTGGACTGCAAAGATACACAAGCGGATCTTTGGAATAGAAT
TTCTAAGGATGAATACATGGCATATGCAGTTCAGGAATGCTACTACAGTGTTGAAAAAATCTTGCATTCCCTGGTTGACGGTGAAGGAAGACTCTGGGTT
GAAAGGATCTTTCGTGAGATCAACAACAGTATATTGGAGGGATCACTTGTCATTACTTTACGTCTTGAGAAGCTTCCTCATGTGTTGTCAAGATTTATTG
CACTGTTTGGACTTCTGATACAAAATGAGACTCCTGTACTTGCCAATGGTGCTGCAAAGGCTGTATATGCCGTGTATGAGGCTGTTACACATGATCTGCT
GTCATCGGATCTAAGAGAGCAACTTGATACATGGAACATTTTAGCTAGAGCAAGAAATGAACGACGCCTGTTCTCCAGGATAGAGTGGCCAAAGGATCCA
GAAATTAAGGAGCAGGTGAAAAGGTTGCAACTTCTTCTCACCGTAAAGGACTCTGCAGCCAACATTCCTAAGAATCTTGAAGCAAGAAGAAGATTAGAGT
TTTTCTCAAATTCACTATTCATGGATATGCCTTCGGCAAAGCCAGTTTCTGAAATGACACCATTTAGTGTCTTCACGCCATACTACAGTGAGACTGTGCT
TTATAGTTCCTCTGAGTTGCGGGTGGAAAATGAGGATGGAATATCCATTCTTTTCTATCTTCAAAAAATCTTTCCAGATGAATGGGAGAATTTCTTGGAG
CGGATTGGTAGAGCAGAGTCTACCGGGGATGCAGATCTTCAAGAAAATTCTGGTGATTCCTTGGAGCTTAGGTTCTGGGCATCTTATCGTGGACAGACTC
TAGCTCGGACTGTACGTGGTATGATGTATTATAGGAGAGCTCTGATGCTGCAGAGTTATCTAGAAAGGAGGTCACAGGGAGTTGATGATTATTCTCAGAC
AAACTTTTCTACATCACAAGGTTTTGAGTTATCACATGAAGCACGGGCACAAGCAGATTTGAAGTTCACTTATGTGGTCTCCTGCCAGATTTATGGGCAA
CAGAAGCAGCGAAAGGCTGTAGAGGCAGCTGATATTTCTCTTTTGCTGCAGCGGAATGAGGCACTTCGAGTTGCTTTCATACATGTGGAGGAGAGTGATA
GTGCTGATGGACAGGTTTCCCATGAATTTTATTCGAAGCTTGTGAAAGCTGATATACATGGAAAAGATCAGGAAATATATTCAATCAAACTTCCTGGAAA
TCCAAAGCTTGGAGAAGGGAAGCCTGAAAACCAGAACCATGCTATAATTTTCACTCGTGGAGAAGCCATTCAGACTATTGACATGAACCAGGACAACTAC
CTTGAAGAGGCAATGAAAATGAGAAATCTTCTTGAGGAATTTCGAGCAAATCATGGCATTCGACCTCCCACCATCCTTGGTGTAAGGGAAAATGTTTTCA
CTGGAAGTGTTTCATCGTTAGCCTGGTTCATGTCCAATCAAGAAACAAGCTTTGTAACATTGGGCCAACGTGTGTTAGCATATCCACTCAAGGTCCGCAT
GCACTATGGCCATCCAGATGTATTTGACCGTGTATTTCATATAACTCGAGGTGGCATCAGTAAGGCATCACGTGTGATCAATATTAGTGAAGATATATTT
GCAGGGTTTAACACCACATTGAGGCAAGGAAATATTACTCACCATGAATATATTCAGGTTGGAAAGGGAAGAGACGTTGGGTTAAACCAGATTGCCCTTT
TTGAAGGGAAAGTGGCAGGAGGAAATGGCGAGCAAGTTCTAAGCAGGGATGTGTACAGGCTTGGGCAACTATTTGATTTCTTCAGAATGCTTTCTTTCTA
CTTTACAACTGTTGGGTACTATGTGTGCACTATGATGACAGTCCTCACAGTATATGTATTCTTGTATGGAAGGGCTTACCTGGCTTTCTCTGGACTTGAT
AATGCTATATCAGTTAGTGCAAAGAAGATGGGAAATACAGCTCTGGATGCTGCTCTTAATGCACAGTTCTTGGTCCAGATTGGAGTTTTTACTGCAATCC
CCATGATAATGGGTTTTATACTTGAACTTGGATTGCTTAAGGCTGTTTTCAGCTTTATTACAATGCAACTTCAGTTGTGTTCAGTCTTCTTCACGTTTTC
ATTGGGCACCAGAACTCATTATTTTGGACGCACTATCCTCCATGGAGGTGCAAAATACAGAGCAACTGGTAGAGGTTTTGTGGTACGTCACATCAAGTTT
GCTGAAAATTACAGGCTCTACTCAAGAAGTCATTTTGTTAAAGCGCTTGAAGTTGCCCTGCTTCTCATAGTTTACATAGCATATGGGTACACAGATGGAG
GTGCTTTGTCATTTGTTTTGTTGACACTTAGCAGTTGGTTTCTTGTCATCTCATGGCTATTTGCTCCTTACATATTTAATCCCTCTGGTTTTGAGTGGCA
AAAGACTGTTGATGATTTTGAAGATTGGACTAGCTGGCTTTTATATAAAGGTGGTGTAGGGGTGAAGGGAGATAATAGCTGGGAATCATGGTGGGAAGAA
GAACAGGCGCATATCCAAACTTTGAGAGGTCGTATATTGGAAACAATATTAAGCTTGAGATTCCTCATTTTTCAATATGGCATCGTATACAAGCTCCATC
TCACAGGAAAGGATAGATCTATTGCGATTTATGGCTTCTCTTGGGTCGTATTGGTTTGCTTTGTCATGATATTTAAGGTCTTCACCTATAGCCCAAAAAG
ATCTACCAGTTTCCAGCTATTGATGCGATTCATGCAAGGAATCGCGTCCTTAGGACTTGTTGCAGCTCTTTGCCTTATTGTTGCCTTCACTGATTTGTCA
ATTCCTGATCTTTTTGCGAGCTTTCTTGCATTTATTGCCACTGGATGGACTATCCTATCTATTGCTATCGCGTGGAAAAGGATAGTTTGGAGTTTAGGCT
TGTGGGATTCAGTGCGTGAGTTTGCCAGGATGTATGATGCTGGGATGGGCGTGTTAATTTTTGTTCCAATTGCATTTTTATCATGGTTTCCATTCGTTTC
TACCTTCCAGTCTCGCCTGCTCTTCAACCAAGCATTCAGCCGTGGGCTGGAGATCTCACTTATCCTTGCTGGAAACAAAGCTAACGTGGATAGATCGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G089300.2 pacid=42775274 polypeptide=Potri.015G089300.2.p locus=Potri.015G089300 ID=Potri.015G089300.2.v4.1 annot-version=v4.1
MSRVSNNWERLVRATLKRELGQGHERMSSGIAGAVPVSLGRTTNIDAILQAADEIQDEDPNVARILCEQAYSMAQNLDPSSDGRGVLQFKTGLMSVIKQK
LAKRDGARIDRNRDIEHLWEFYQHYKRRHRVDDIQREEQKFRESGNFSTVIRGEFELSSLEMKKVFATLRALEDVMEAVSKDADPHGAGRHIMEELQRIK
TVGELTSYNIVPLEAPSLSNAIGVFPEVRGAMSAIRYAEHYPRLPAGFVISGERDLDMFDLLEYVFGFQNDNVRNQRENVVLAIANAQSRLGIPIQADPK
IDEKAINEVFLKVLDNYIKWCKYLRKRLAWNSIEAINRDRKLFLVSLYYLIWGEAANVRFLPECICYIFHHMAKELDAILDHGEANHAASCITESGSVSF
LEQIICPIYQTIAAEAERNNNGKAVHSAWRNYDDFNEYFWSPACFELSWPMKENSSFLLKPKKSKRTGKSTFVEHRTFLHIYRSFHRLWIFLALMFQALA
IIAFNHGDLSLDTFKEMLSVGPSFAIMNFIESCLDVLLMFGAYSTARGMAISRLVIRFFWCGLSSVFVTYLYVKVLEEKNRQNSDSFHFRIYILVLGVYA
ALRLFLALLLKFPACHALSDMSDQSFFQFFKWIYQERYYVGRGLFEKMSDYCRYVLYWLVIFACKFTFAYFLQQIRPLVKPTNTIRALPSLPYSWHDLIS
KNNNNVLTIASLWAPVVAIYIMDIHIWYTILSAIVGGVMGARARLGEIRSIEMVHKRFESFPAAFVKNLVSPQAQRMVLNRHASQEAQDMNKAYAALFAP
FWNEIIKSLREEDYISNREMDLLSIPSNTGSLRLVQWPLFLLSSKILLAVDLALDCKDTQADLWNRISKDEYMAYAVQECYYSVEKILHSLVDGEGRLWV
ERIFREINNSILEGSLVITLRLEKLPHVLSRFIALFGLLIQNETPVLANGAAKAVYAVYEAVTHDLLSSDLREQLDTWNILARARNERRLFSRIEWPKDP
EIKEQVKRLQLLLTVKDSAANIPKNLEARRRLEFFSNSLFMDMPSAKPVSEMTPFSVFTPYYSETVLYSSSELRVENEDGISILFYLQKIFPDEWENFLE
RIGRAESTGDADLQENSGDSLELRFWASYRGQTLARTVRGMMYYRRALMLQSYLERRSQGVDDYSQTNFSTSQGFELSHEARAQADLKFTYVVSCQIYGQ
QKQRKAVEAADISLLLQRNEALRVAFIHVEESDSADGQVSHEFYSKLVKADIHGKDQEIYSIKLPGNPKLGEGKPENQNHAIIFTRGEAIQTIDMNQDNY
LEEAMKMRNLLEEFRANHGIRPPTILGVRENVFTGSVSSLAWFMSNQETSFVTLGQRVLAYPLKVRMHYGHPDVFDRVFHITRGGISKASRVINISEDIF
AGFNTTLRQGNITHHEYIQVGKGRDVGLNQIALFEGKVAGGNGEQVLSRDVYRLGQLFDFFRMLSFYFTTVGYYVCTMMTVLTVYVFLYGRAYLAFSGLD
NAISVSAKKMGNTALDAALNAQFLVQIGVFTAIPMIMGFILELGLLKAVFSFITMQLQLCSVFFTFSLGTRTHYFGRTILHGGAKYRATGRGFVVRHIKF
AENYRLYSRSHFVKALEVALLLIVYIAYGYTDGGALSFVLLTLSSWFLVISWLFAPYIFNPSGFEWQKTVDDFEDWTSWLLYKGGVGVKGDNSWESWWEE
EQAHIQTLRGRILETILSLRFLIFQYGIVYKLHLTGKDRSIAIYGFSWVVLVCFVMIFKVFTYSPKRSTSFQLLMRFMQGIASLGLVAALCLIVAFTDLS
IPDLFASFLAFIATGWTILSIAIAWKRIVWSLGLWDSVREFARMYDAGMGVLIFVPIAFLSWFPFVSTFQSRLLFNQAFSRGLEISLILAGNKANVDRS
|
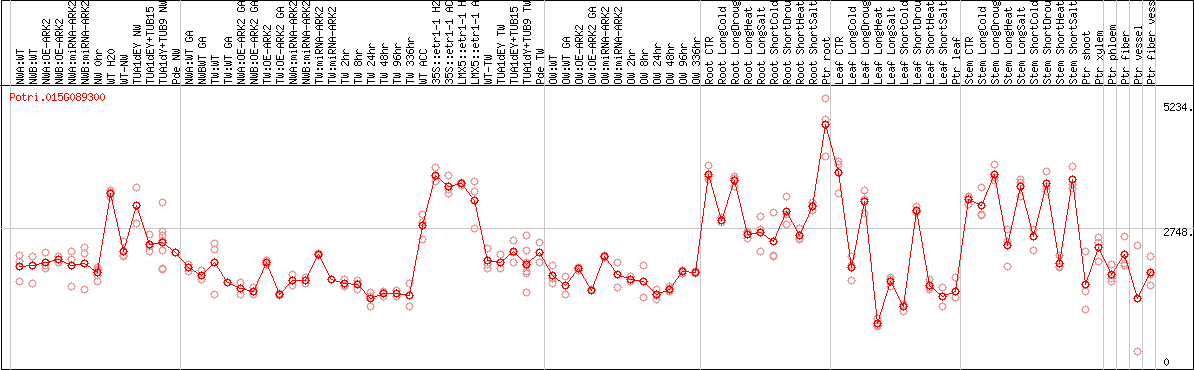
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G089300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
