External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G22830 130 / 2e-36
SQE2
squalene epoxidase 2 (.1)
AT4G37760 129 / 4e-36
SQE3
squalene epoxidase 3 (.1)
AT1G58440 114 / 1e-30
SQE1, XF1
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
AT5G24150 76 / 4e-17
SQP1, SQE5
SQUALENE MONOOXYGENASE 5, FAD/NAD(P)-binding oxidoreductase family protein (.1), FAD/NAD(P)-binding oxidoreductase family protein (.2)
AT5G24160 72 / 9e-16
SQE6
squalene monoxygenase 6 (.1)
AT5G24140 71 / 2e-15
SQP2
squalene monooxygenase 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G120900
148 / 3e-43
AT1G58440 745 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Potri.002G114500
135 / 2e-38
AT1G58440 761 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Potri.005G146700
134 / 4e-38
AT1G58440 766 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Potri.007G007600
134 / 7e-38
AT4G37760 795 / 0.0
squalene epoxidase 3 (.1)
Potri.012G121136
111 / 8e-30
AT1G58440 564 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Potri.019G014378
110 / 3e-29
AT1G58440 603 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Potri.019G014376
110 / 4e-29
AT1G58440 575 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011732
134 / 7e-38
AT1G58440 785 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Lus10039649
134 / 9e-38
AT1G58440 791 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Lus10007718
131 / 7e-37
AT1G58440 828 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
Lus10018654
131 / 9e-37
AT1G58440 827 / 0.0
SQUALENE EPOXIDASE 1, FAD/NAD(P)-binding oxidoreductase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF08491
SE
Squalene epoxidase
Representative CDS sequence
>Potri.015G121201.1 pacid=42775937 polypeptide=Potri.015G121201.1.p locus=Potri.015G121201 ID=Potri.015G121201.1.v4.1 annot-version=v4.1
ATGACTGTGGCTCTATCACACATTGTGCTACTACGTGATCTTCTTCGACCTCAACTCAATGATACATCTTCCTTTGCAAATATCCCCAGTCCTTCTACTC
CCTGGCAAGTGGCACCTACCATAAGCACATTGGCAGGTGCCCTATACAAGGTGTTTAGGGCATCACCTGATCATCCAGCTAGAAACGAAATGTGCCAAGC
ATGTTTTGACTACATGAGCCTTGAAGGAGTATTTTCGAATGGACTAATAGCTCTACTTTCTGGTCTAGACCCTCGACCGTTGAGCTCGGTTCTCCATTTC
TTTGCAGTTGCTATCTATGGTGTTGGCCGCCTAAAGCTTCCCTTTAAGTCTAGTCTGCACTGTGCTTTCCCGCTTATGTAG
AA sequence
>Potri.015G121201.1 pacid=42775937 polypeptide=Potri.015G121201.1.p locus=Potri.015G121201 ID=Potri.015G121201.1.v4.1 annot-version=v4.1
MTVALSHIVLLRDLLRPQLNDTSSFANIPSPSTPWQVAPTISTLAGALYKVFRASPDHPARNEMCQACFDYMSLEGVFSNGLIALLSGLDPRPLSSVLHF
FAVAIYGVGRLKLPFKSSLHCAFPLM
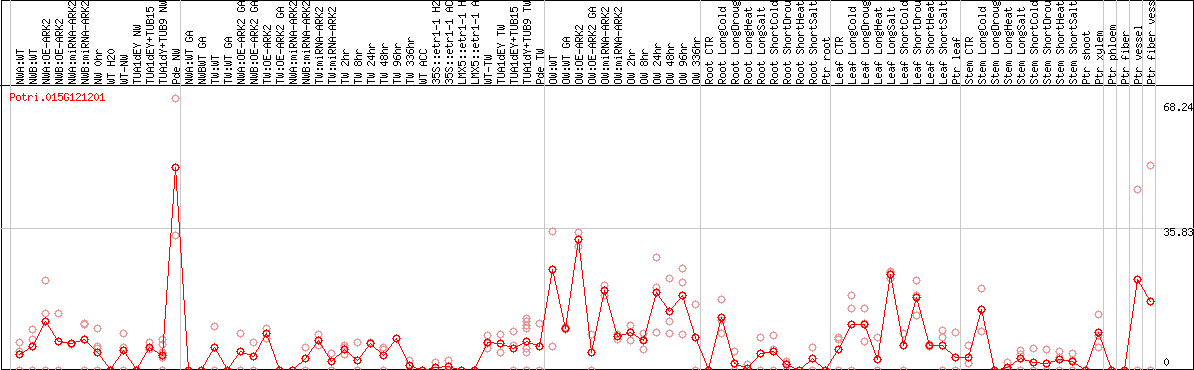
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G121201 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.