External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G16470 456 / 1e-165
PAB1
proteasome subunit PAB1 (.1.2)
AT1G79210 453 / 2e-164
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT5G66140 168 / 1e-51
PAD2
proteasome alpha subunit D2 (.1)
AT3G51260 164 / 5e-50
PAD1
20S proteasome alpha subunit PAD1 (.1.2)
AT1G53850 156 / 3e-47
PAE1, ATPAE1
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
AT3G14290 154 / 2e-46
PAE2
20S proteasome alpha subunit E2 (.1)
AT3G22110 154 / 3e-46
PAC1
20S proteasome alpha subunit C1 (.1)
AT5G35590 153 / 5e-46
PAA1
proteasome alpha subunit A1 (.1)
AT2G05840 149 / 2e-44
PAA2
20S proteasome subunit PAA2 (.1.2)
AT5G42790 146 / 5e-43
ARS5, ATPSM30, PAF1
ARSENIC TOLERANCE 5, proteasome alpha subunit F1 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017135
466 / 1e-169
AT1G16470 454 / 1e-164
proteasome subunit PAB1 (.1.2)
Lus10036210
463 / 3e-168
AT1G16470 450 / 3e-163
proteasome subunit PAB1 (.1.2)
Lus10018318
437 / 5e-150
AT2G35130 801 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10041890
173 / 2e-53
AT5G66140 442 / 9e-160
proteasome alpha subunit D2 (.1)
Lus10028437
167 / 1e-51
AT5G66140 417 / 8e-150
proteasome alpha subunit D2 (.1)
Lus10027669
158 / 7e-48
AT2G05840 473 / 1e-171
20S proteasome subunit PAA2 (.1.2)
Lus10042145
157 / 3e-47
AT3G22110 474 / 3e-172
20S proteasome alpha subunit C1 (.1)
Lus10004235
150 / 1e-45
AT3G22110 375 / 2e-134
20S proteasome alpha subunit C1 (.1)
Lus10037454
155 / 8e-44
AT1G53850 464 / 8e-163
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
Lus10003936
155 / 1e-43
AT1G53850 463 / 9e-162
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF00227
Proteasome
Proteasome subunit
CL0052
NTN
PF10584
Proteasome_A_N
Proteasome subunit A N-terminal signature
Representative CDS sequence
>Potri.015G122400.3 pacid=42775535 polypeptide=Potri.015G122400.3.p locus=Potri.015G122400 ID=Potri.015G122400.3.v4.1 annot-version=v4.1
ATGGGTGATAGCCAGTACTCCTTTTCTCTCACTACTTTCAGTCCTACAGGAAAGCTAGTACAGATCGAGCACGCATTGACTGCCGTTGGATCAGGCCAAA
CTTCTCTCGGAATCAAAGCTGCAAACGGTGTTGTTATTGCAACTGAGAAGAAATTGCCTTCTATTTTAGTCGATGAATCATCTGTTCAGAAAATACAGAA
TCTGACACCTAATATTGGAGTTGTCTACAGTGGAATGGGTCCTGACTTTCGAGTTTTAGTTCGGAAAAGTAGGAAGCAGGCAGAACAATATCATAGATTG
TACAAGGAACCAATTCCTGTCACACAACTTGTGAGGGAAACTGCCGCTGTCATGCAAGAGTTTACTCAATCTGGAGGTGTAAGGCCTTTTGGAGTATCTT
TGCTAGTTGCTGGTTTTGATGACAAAGGTCCACAGCTGTACCAGGTTGATCCCTCAGGTTCCTACTTCTCATGGAAAGCCTCAGCAATGGGAAAAAATGT
TTCAAATGCAAAAACATTTCTTGAGAAGAGATACACTGATGATATGGAGCTTGATGATGCTGTACACACTGCTATCTTGACACTGAAAGAGGGATTTGAA
GGACAGATCTCTGGGAAAAATATTGAAATTGGGATAATTGGTGCTGACAAGCAATTCAGGGTACTAACTCCAGCTGAAATTGATGATTATCTGGCTGAAG
TGGAATAG
AA sequence
>Potri.015G122400.3 pacid=42775535 polypeptide=Potri.015G122400.3.p locus=Potri.015G122400 ID=Potri.015G122400.3.v4.1 annot-version=v4.1
MGDSQYSFSLTTFSPTGKLVQIEHALTAVGSGQTSLGIKAANGVVIATEKKLPSILVDESSVQKIQNLTPNIGVVYSGMGPDFRVLVRKSRKQAEQYHRL
YKEPIPVTQLVRETAAVMQEFTQSGGVRPFGVSLLVAGFDDKGPQLYQVDPSGSYFSWKASAMGKNVSNAKTFLEKRYTDDMELDDAVHTAILTLKEGFE
GQISGKNIEIGIIGADKQFRVLTPAEIDDYLAEVE
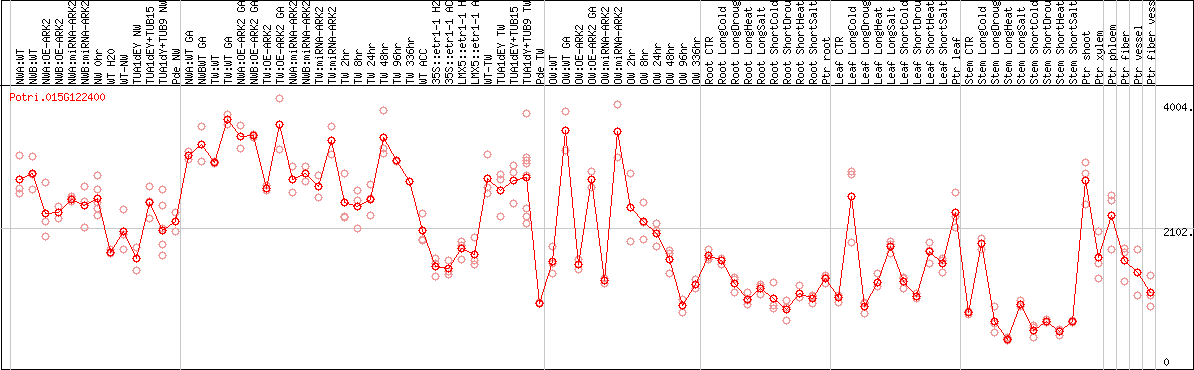
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G122400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.