Potri.015G123400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G123400.2 pacid=42775737 polypeptide=Potri.015G123400.2.p locus=Potri.015G123400 ID=Potri.015G123400.2.v4.1 annot-version=v4.1
ATGGCTTCCCACATCTCCCTCTTTTCCACACCTTTTTTGGTTTTTTCCTTTCTTGCTTGTGCTTCCTTCTTTGCTTCTTTTGCTTACTCGGCTAGTACTG
GAGCTGCTGAAGTAGCAAATGGAAGGACGGAAGCGGAAGCTCTTCTAGAATGGAAAGTCAGTCTTGACAACCAAAGCCAGTCTCTCCTGTCTTCATGGGC
TGGAGACAGCCCTTGCAACTGGTTCGGAATCAGTTGCGACCAGTCTGGTAGTGTCATCAACATAAGTCTGCCCGATTCTAGTTTGAGAGGTACGCTTAAT
AGACTCAGATTCTCTTCCTTTCCTAACTTGACTGTGCTTAACCTTCCCAACAACTCCCTTTATGGTTATGTTCCTTCACATATAGGTAACCTTTCTAACC
TCTCCATCCTTAACTTGGCTTTCAATAGCATTTCTGGCAACATTCCACCAGAAATTGGAAATTTAGTATCTCTCACAATTTTAGCTTTGTCATCTAATAA
ACTAACGGGTACAATACCAGCATCTTTAGAAAATTTAAAAAACTTATCTAAACTTTACCTTTGGAATAATAATCTTTTTGGTTCCATTACTTTTATTGGA
AACCTGACGAGATCTCTCACCATTTTAGTTTTGTCATTTAATAAACTAACTGGTACAATACCAGCATCTTTACAAAATTTAAAAAGCTTATCTAAACTTA
ACCTTTGTAATAATAATCTTTCTGGTTCCATTACTTTTATTGAAAACATAACGAGATCTCTCACCAATTTAGATTTATCATTCAATAAATTAACTGGTAC
AATACCAGCATCTTTAGAAAATTTAAGAAGCTTATATCAACTTGACCTTCAACATAACAATTTTTTTGGTCCCATTATTTTTATCCAAAACCTGACGAGA
TCTCTCACTATTTTAGATTTGTCATTTAATAATTTAACTGGTACAATACCTGCATCTCTAGGCAATTTAAGAAGCTTATCTCAACTTTACTTCCAAGATA
ACAGTCTTTTTGGTCCGATTCCTCCAGAAATGAACAATCTTACACATTTGTATTCGTTAGAAATATATTCTAATAGATTGTCTGGTAATCTACCACGAGA
TGTGTGCCTTGGTGGATTACTTTCATATTTTTCTGCGTCTGAAAATTATTTCACTGGACCTATCCCAAAAAGCTTGAGAAATTGCAGCAGCTTGTTGAGA
CTCAGACTTGAGAGAAACCAACTTAGCGGAAACATATCTGAAGCTTTTGGCACACATCCCCATCTGTACTACATGGGTTTGAGTGATAATGAATTGCATG
GTGAACTTTCATTGAAATGGGAGCAGTTTAACAATCTGGCAGCCTTCAAGATTTCTGGAAACAAGATTTCTGGAGAGATACCCGCAGCTCTTGGAAAGGC
AACTCATCTACAAGCTCTTGACCTGTCGTCAAATCAACTAGTTGGGAGAATTCCCAAAGAATTGCGAAATTTGAAGTTGATTGAACTTGCACTAAACGAT
AACAAGCTTTCAGGTGATATTCCTTTCGATGTGGCATCGCTATCTGATCTGGAAAGGCTTGGCCTGGCCGCGAACAATTTTAGTGCCTCAATTCTTACGC
AGCTTGGCAAGTGCTCAAAACTGGTATTCTTGAATATGAGCAAGAATAGATTCACAGGGATTATTCCTGCTGAGATGGGGTCTTTACAGTCTCTTCAAAA
TCTTGATCTCAGTTGGAATTCCCTCATGGGAGGTATAGCACCGGAGCTTGGACAGCTGCAGCAGCTAGAGTTTCTGAACCTCTCCCATAATATGCTGTCT
GGTCTCATTCCAGCCAGTTTTAGTAGACTGCAAGGCCTCACTAAAGTGGATGTATCCTATAACAAGCTAGAGGGTCCCATTTCTGACATCAAAGCCTTTC
GCGAGGCACCATTTGAAGCAATTCGCAACAACACCAACCTGTGTGGCAATGCAACTGGTTTGGAGGCTTGTTCTGCTCTCATGAAAAACAAAACTGTGCA
CAAGAAGGGCCCCAAAGTTGTCTTTTTGACAGTATTTTCTCTGCTGGGTAGTTTGTTGGGCCTGATTGTAGGTTTTCTTATCTTTTTCCAAAGCAGAAGA
AAGAAAAGGTTAGTGGAAACACCACAAAGAGATGTTACCGCGAGATGGTGCCCTGGTGGGGATCTGCGCTACGAGGACATCATTGAGGCTACTGAGGAAT
TCGACTCCAAATATTGCATTGGTACTGGAGGGTATGGAGTTGTATACAAAGCTGTGCTTCCATCAGAACAAGTCCTTGCCGTGAAGAAATTCCACCAAAC
ACCAGAAGTTGAGATGAGCAGCTTGAAAGCTTTTAGAAGCGAGATTGATGTCTTAATGGGTATACGGCATCGAAATATTGTGAAGCTGTATGGTTTCTGC
TCCCATGCCAAACACTCATTTTTGGTATATGAATTTGTGGAAAGGGGAAGTTTAAGAAAGGTACTGAACGAGGAGGAACAATCAGCAAAGATGGATTGGG
ATAAAAGGATGAATCTTATCAAAGGCGTAGCCAATGCTTTATCCTACATGCACCATGACTGCTCCCCTCCGATTATTCATAGAGACATTTCCAGCAACAA
TGTTCTTTTGGATTCAGAATATGAGGCTCATGTCTCTGACTTCGGCACAGCCAGGCTCTTAATGCCTGACTCGTCCAATTGGACATCATTTGCTGGCACC
TTTGGGTACACAGCTCCAGAGTTGGCATACACAATGAAAGTGGATGAGAAGTGTGATGTGTATAGCTTTGGAGTATTAACATTGGAAGTAATGATGGGAA
AGCATCCTGGCGATTTCATCTCATCTCTGATGTTATCTGCCTCCACCTCCTCATCATCACCAATTGGTCACAATACAGTCTTGAAGGATGTCTTAGACCA
GCGCCTCCCACCTCCTGAAAACGAGCTTGCGGACGGTGTGGCACATGTTGCAAAACTGGCGTTTGCTTGCTTGCAGACCGATCCTCACTATCGGCCAACA
ATGCGACAGGTTTCTACAGAGCTTACAACTCGATGGCCTCCATTGCCAAAGCTATTCTCCACGATGGAATTGGAAGATATAATGGTTCACAGAAATGATA
TTGGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G123400.2 pacid=42775737 polypeptide=Potri.015G123400.2.p locus=Potri.015G123400 ID=Potri.015G123400.2.v4.1 annot-version=v4.1
MASHISLFSTPFLVFSFLACASFFASFAYSASTGAAEVANGRTEAEALLEWKVSLDNQSQSLLSSWAGDSPCNWFGISCDQSGSVINISLPDSSLRGTLN
RLRFSSFPNLTVLNLPNNSLYGYVPSHIGNLSNLSILNLAFNSISGNIPPEIGNLVSLTILALSSNKLTGTIPASLENLKNLSKLYLWNNNLFGSITFIG
NLTRSLTILVLSFNKLTGTIPASLQNLKSLSKLNLCNNNLSGSITFIENITRSLTNLDLSFNKLTGTIPASLENLRSLYQLDLQHNNFFGPIIFIQNLTR
SLTILDLSFNNLTGTIPASLGNLRSLSQLYFQDNSLFGPIPPEMNNLTHLYSLEIYSNRLSGNLPRDVCLGGLLSYFSASENYFTGPIPKSLRNCSSLLR
LRLERNQLSGNISEAFGTHPHLYYMGLSDNELHGELSLKWEQFNNLAAFKISGNKISGEIPAALGKATHLQALDLSSNQLVGRIPKELRNLKLIELALND
NKLSGDIPFDVASLSDLERLGLAANNFSASILTQLGKCSKLVFLNMSKNRFTGIIPAEMGSLQSLQNLDLSWNSLMGGIAPELGQLQQLEFLNLSHNMLS
GLIPASFSRLQGLTKVDVSYNKLEGPISDIKAFREAPFEAIRNNTNLCGNATGLEACSALMKNKTVHKKGPKVVFLTVFSLLGSLLGLIVGFLIFFQSRR
KKRLVETPQRDVTARWCPGGDLRYEDIIEATEEFDSKYCIGTGGYGVVYKAVLPSEQVLAVKKFHQTPEVEMSSLKAFRSEIDVLMGIRHRNIVKLYGFC
SHAKHSFLVYEFVERGSLRKVLNEEEQSAKMDWDKRMNLIKGVANALSYMHHDCSPPIIHRDISSNNVLLDSEYEAHVSDFGTARLLMPDSSNWTSFAGT
FGYTAPELAYTMKVDEKCDVYSFGVLTLEVMMGKHPGDFISSLMLSASTSSSSPIGHNTVLKDVLDQRLPPPENELADGVAHVAKLAFACLQTDPHYRPT
MRQVSTELTTRWPPLPKLFSTMELEDIMVHRNDIG
|
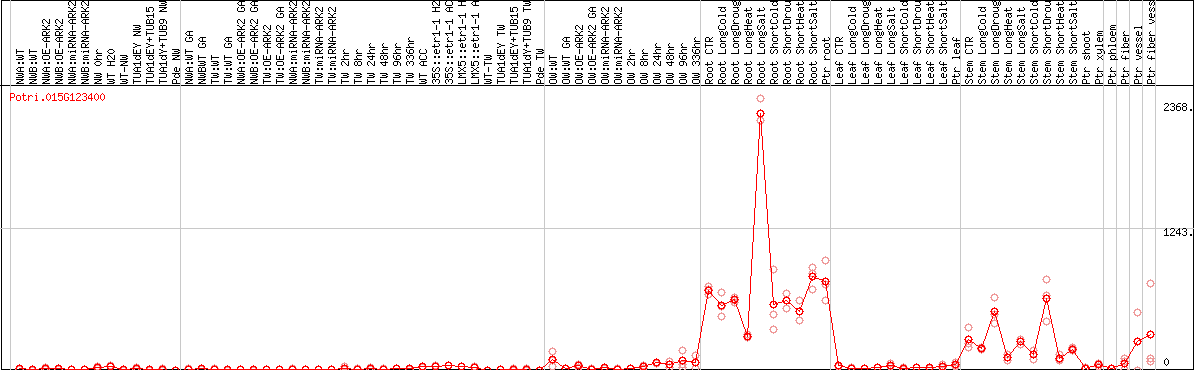
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G123400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
