External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G51780 143 / 1e-42
bHLH
bHLH036
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G51790 141 / 1e-41
bHLH
bHLH120
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G25410 141 / 2e-41
bHLH
bHLH126
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G25400 133 / 5e-39
bHLH
bHLH118
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G62975 112 / 7e-30
bHLH
bHLH125
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G12540 111 / 2e-29
bHLH
bHLH055
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G04150 47 / 3e-06
bHLH
bHLH101
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G20970 42 / 9e-05
bHLH
bHLH162
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G17880 42 / 0.0004
bHLH
bHLH004, MYC4
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT5G46760 40 / 0.001
bHLH
bHLH005, ATR2, MYC3
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G132000
319 / 6e-111
AT5G51790 154 / 6e-47
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G113400
160 / 1e-48
AT4G25410 124 / 6e-35
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G132100
160 / 1e-48
AT4G25410 120 / 3e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.015G134400
139 / 2e-40
AT5G51790 107 / 2e-28
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.011G053400
62 / 2e-11
AT4G20970 160 / 2e-49
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.004G044400
61 / 3e-11
AT4G20970 150 / 3e-46
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G172101
51 / 3e-07
AT1G01260 526 / 0.0
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Potri.019G034700
49 / 4e-07
AT4G20970 104 / 5e-28
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G079100
48 / 8e-07
AT4G20970 105 / 6e-29
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031677
169 / 9e-52
AT5G51780 143 / 2e-42
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027394
169 / 1e-51
AT5G51780 149 / 9e-45
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027395
122 / 7e-34
AT5G51780 138 / 9e-41
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10007049
120 / 1e-32
AT1G62975 138 / 3e-39
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10031676
112 / 8e-30
AT5G51780 119 / 1e-33
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10031675
85 / 2e-19
AT4G25400 94 / 4e-24
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027396
81 / 7e-18
AT4G25400 89 / 6e-22
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10006707
73 / 8e-16
AT1G62975 89 / 8e-22
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10004574
44 / 6e-05
AT1G32640 618 / 0.0
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10004575
44 / 0.0001
AT4G17880 445 / 9e-152
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.015G134300.1 pacid=42776025 polypeptide=Potri.015G134300.1.p locus=Potri.015G134300 ID=Potri.015G134300.1.v4.1 annot-version=v4.1
ATGTTCCCTTTACACCAAGGTGGTGAGCTGTGCTTCAAGATTTCCTCCAGTCCACACCAGCAAGACAATATCCCAGATCTGATTCTTGCTCATCAATATG
CTGAAATCCATGGCAGTACTGATATAACCAACAATATGGAGAAAGGTCGGCGGCGAAATATAATTTCCATGGACAATAATGAGGCTGCTAGAGATAATAA
TAACAATAGTAAGAAGAAGATGATGCATAGAGATATTGAAAGACAAAGGAGGCAAGAAATGACAACCCTTTATGCATCCCTTCGAGCTTTACTTCCTCTT
GAATTTATCAAGGGAAAGCGTTCGATTTCAGATCACATGAACGAGGCAGTGAATTATATAAAGTATCTCCAAAAGAAGATCAAAGAAACTAGTGCCAAAA
GAGATGAGTTAAAGAAACTATCCGATTTCAGTTCTGTTGCCTCACCAAGTGGATGTTCTAACAAATCTTCTTCAAGTTCAGTAGCTCTCCAGCCATATCC
AGGTGGTATTGAGATTACGTTTGATAGTGATTTAATGGGGAGAGATTTGCCCCTATCAAGAGTATTGCAAGTGTTGCTTGAAGAAGGAATTTCTGTTATC
AACTGTGTTTCCACCAAAGTAAATGAAAGATTATTCCACTCTGTCCAAACTGAGGTCAATGATCCTACATGCTTAAATTTATCCGAGCTGTGGCAGAAAT
TAACCCTAGTAGTTTCTTCTACAAGTGATTTATCCAAGTGA
AA sequence
>Potri.015G134300.1 pacid=42776025 polypeptide=Potri.015G134300.1.p locus=Potri.015G134300 ID=Potri.015G134300.1.v4.1 annot-version=v4.1
MFPLHQGGELCFKISSSPHQQDNIPDLILAHQYAEIHGSTDITNNMEKGRRRNIISMDNNEAARDNNNNSKKKMMHRDIERQRRQEMTTLYASLRALLPL
EFIKGKRSISDHMNEAVNYIKYLQKKIKETSAKRDELKKLSDFSSVASPSGCSNKSSSSSVALQPYPGGIEITFDSDLMGRDLPLSRVLQVLLEEGISVI
NCVSTKVNERLFHSVQTEVNDPTCLNLSELWQKLTLVVSSTSDLSK
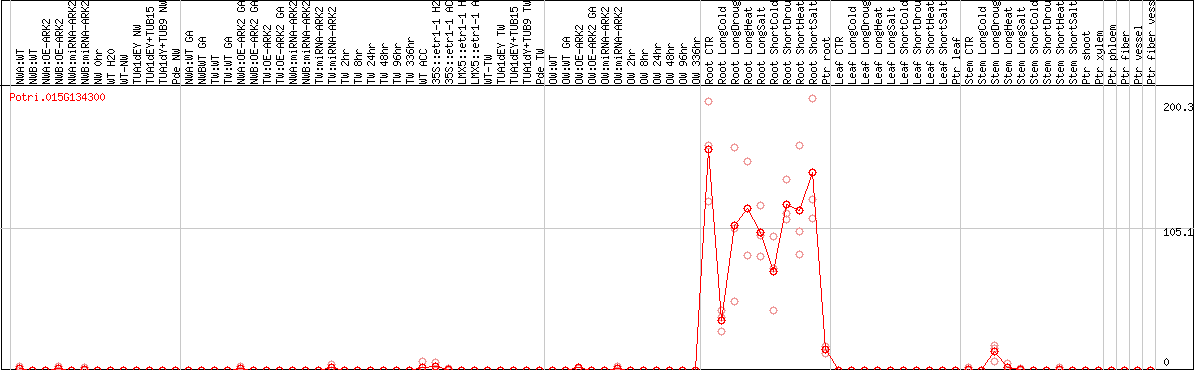
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G134300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.