Potri.015G145300 [POPLAR]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G145300.2 pacid=42775164 polypeptide=Potri.015G145300.2.p locus=Potri.015G145300 ID=Potri.015G145300.2.v4.1 annot-version=v4.1
ATGCCTGGGGATTCAATTACTGAAACATCTGGTGGTTATGGTGGTGGAGATGAGAACCGGCCTCTCAGGAATTTCAAGCTGAATGAATCAACTTTTTTGG
CTTCCTTGATGCCTAAGAAAGAGATTGGTGCTGATCACTTCGTTGAAGCTCACCCTCAATATGATGGCCGTGGTGTCATCATTGCTATCTTTGATTCTGG
AGTTGATCCGGCTGCCTCTGGATTACAAGTGACATCAGATGGGAAGCCGAAAGTGCTTGATGTTATTGATTGTACTGGGAGCGGAGATATTGATACTTCA
AAAGTGGTGAAGGCTGATGCAGATGGTTGCATTCAAGGAGCTTCAGGTGCTTCTTTGGTTGTCAATTCTTCATGGAAGAATCCTTCTGGTGAATGGCATG
TGGGTTACAAGTTTTTGTATGAGCTACTCACTGACACACTGACTTCTCGTTTGAAGAAAGAGAGGAAAAAGAAGTGGGACAAGAAAAACCAAGAAGAAAT
CGCCAAGGCTGTAAAGCATCTGGACGAATTCAACGAGAAACACTCCAATCCAGAGGAAGCAGACTTGAAAAGGGTTCGTGAAGACCTCCAAGCCAGAATT
GATTTGCTACGGAAACAGGCTGATAGCTATGATGACAAGGGGCCTGTCATAGATGCTGTTGTATGGCATGATGGAGATCTGTGGAGGGCTGCTCTTGACA
CACAGAGTGTTGAGGATGACTCAGATTGTGGACAGCTAGCTAACTTCGTACCGCTTACAAATTATAGGATTGAAAGGAAGCATGGAGTGTTCAGCAAGTT
AGATGCCTGTGCATTTGTTCTTAATGTTTATAGTGACGGAAACATCTTAAGTATTGTAACGGATTGCTCGCCTCATGGAACTCATGTTGCTGGTATTGCT
GCAGCCTTCCACCCGAAGGAGCCGTTATTGAATGGAATTGCACCCGGGGCCCAGTTGATATCTTGCAAAATTGGCGACACACGTTTAGGTTCAATGGAGA
CAGGCACTGGCTTGATCCGGGCCCTAATAGCTGCTGTGGAGCATAAATGTGATCTCATCAACATGAGTTATGGGGAACCAACGTTACTGCCAGATTATGG
CCGCTTCGTTGACCTTGTTAATGAAGTGGTGAACAAGCACCGTCTTATATTTGTAAGTAGTGCTGGCAATGGTGGTCCAGCATTAAGTACTGTTGGTGCT
CCTGGTGGCACAACTTCAAGCATTATAGGTGTTGGTGCATATGTCTCTCCTTCCATGGCTGCAGGTGCTCATTCTGTGGTTGAGCCTCCTTCTGAAGGAC
TTGAATACACATGGTCAAGCCGAGGACCAACTTCTGATGGAGATCTTGGGGTCAGTATTAGTGCTCCTGGAGGGGCTGTTGCTCCTGTCCCTACCTGGAC
TCTTCAAAAACGCATGCTTATGAATGGAACATCAATGGCATCTCCATCTGCTTGTGGTGGAGTTGCATTACTTATTAGTGCAATGAAGGCTGAGGGTATT
CCTGTAAGTCCATACAGTGTAAGGAAGGCCCTTGAGAATACATCCGGTCCTGTAGGTGAATTGCCAGCAGATAAACTATCCACTGGACAGGGGCTTATGC
AAGTTGACAGAGCCCACGAATATATAAGGCAGTCTCGGAATATTCCATGTATTTGCTATGAAATAATGGTCAATCAATCTGGAAAATCAACTCCTACTTC
ACGAGGCATCTACCTGAGGGAGGCTAGTGCTTGCCAACAACCTACAGAGTGGACGGTGCAAGTCCAACCAAAGTTTCATGAAGGTGCAAGTAACTTGGAA
GAATTGGTTCCCTTCGAGGAATGCATTGAGTTACACTCTACTGAAAAAGTTGTTGTGAGGGCTCCTGAGTATCTCCTTCTCACTAACAATGGGCGTAGCT
TCAACATAGTGGTTAATCCCACCAAGCTAAGTGAAGGTCTACATTATTATGAAGTATACGGGGTTGATTGCAAAGCACCATGGCGTGGTCCCATTTTCAG
GATCCCAGTCACCATAACAAAGCCTATGACAGTGAAGAATCACCCTCCATTCATTTCATTTTCTAGGATGTCATTTCTACCAGGGCACATAGAAAGGAGA
TATATTGAAGTGCCATTTGGTGCTACTTGGGTTGAGGCAACTATGAAAACATCAGGGTTTGATACTACTCGACGATTTTTTGTCGACACTGTTCAGATTT
GTCCCTTACAAAGACCTATGAAGTGGGAGAGTGTTGTAACATTTTCTTCTCCTACTGCCAAAAGTTTTGCATTTCCTGTTGTGGGTGGTCAGACAATGGA
ATTAGCTGTTGCTCAGTTTTGGTCAAGTGGGATAGGAAGTCATGAAACCACCATTGTAGACTTTGAGATTTTGTTTCATGGGATTGCCATCAATAAAGAG
GAAATCATTCTTGATGGAAGTGAAGCACCTATAAGAATTGATGCTGAAGCTCTATTGTCATCTGAGAATCTCGTCCCTGCTGCTACTCTGAATAAGATAA
GAGTTCCATATCGACCCGTCGATGCTAAACTTGGTACTCTTACAGAGAATCGCGACAAACTACCCTCTGGAAAACAGACACTGGCACTGACATTAACTTA
CAAGTTTAAATTGGAAGATGGAGCTGAAGTGAAACCTCAAGTTCCATTGCTTAACAATCGCATATATGACACAAAATTTGAATCACAATTTTATATGGTT
TCTGACACTAACAAGCGTGTATATGCCATGGGTGATGTCTATCCTAGTGCTACAAAACTTCCCAAGGGTGAATACAATTTACGCCTATATCTACGGCATG
ACAATATGCAATATTTGGAAAAGATGAAGCAGTTGCTATTGTTTATTGAGAGGAATTTGGATGACAAGGATGTCATTCGATTGAACTTTTTCTCTGAACC
TGATGGTCCTGTGATGGGAGATGGTGCTTTTAAGTCCTCAGTTTTAGTTCCAGGGAAGAAAGAAGCAATTTACTTGGGCCCACCAGTGAAAGACAAGCTC
CCGAAGAATGCTCCACAAGGATCTGTACTGCTTGGAGCAATCTCATATGGAAAGCTATCACTTGCTGGCCAGGAAGGGGAGGAAAGTTCACAAAAGAACC
CTGTATCTTACCAAATCTCTTATGTAGTGCCACCAAATAAGGTTGATGAGGACAAGGGAAAAAGTTCTTCAACTAGCCTGAAAACTGTGTCTGAGCGTTT
AGAAGAAGAGGTTCGAGATGCAAAGATAAGAGTTCTTTCAAGCCTAAAACAAGACACTGATGAAGAACGCTCGGAATGGAAAAAGCTATCCACTTCTCTT
AAGTCTGATTACCCAAACTACACTCCACTACTTGCTAAGATTTTGGAAGGCTTGCTTTCACAGAGTAAGGTGGAAGATAAAATTCATCATCATGAAGATG
TTATGGATGCAGCAGATGAGGTAATTGATAGTATTGACAAAGATGAGTTGGCAAAGTTCTTTTCACTCAAGAGTGATCCAGAAGATGAGGAGACAGAGAA
AAAGAAAAAGGCAATGGAGACGACCAGGGATGAATTGGCCGAGGCACTGTACCAAAAAGGGTTGGCATTGGTGGAGAATGAATCTCTAAAGGTAAGGAAA
GCTGAAACAGAAGGCACAAAAGATTTGTTCGAGGACAATTTCAAAGGGCTGCAAAAATGGGTTGATGCAAAGTCCTCTAAATATGGCACCCTCTTAGTGC
TACGTGAAAGACGACGTGGGAGGCTAGGAGCAGCACTAAAGGCATTGAATGAAATGATGCAAGATAATGGAGATCCACCCAAGAAGAAGCTGTATGAACT
GAAGCTCTCGTTGCTTGATGAGATTGGATGGAAACATTTATCAACATATGAGAAAGAGTGGATGCTCGTGCGGTTCCCACCAAGCTTGCCTCTTTTTTAA
|
||||||||||||||||||||
|
AA sequence
|
>Potri.015G145300.2 pacid=42775164 polypeptide=Potri.015G145300.2.p locus=Potri.015G145300 ID=Potri.015G145300.2.v4.1 annot-version=v4.1
MPGDSITETSGGYGGGDENRPLRNFKLNESTFLASLMPKKEIGADHFVEAHPQYDGRGVIIAIFDSGVDPAASGLQVTSDGKPKVLDVIDCTGSGDIDTS
KVVKADADGCIQGASGASLVVNSSWKNPSGEWHVGYKFLYELLTDTLTSRLKKERKKKWDKKNQEEIAKAVKHLDEFNEKHSNPEEADLKRVREDLQARI
DLLRKQADSYDDKGPVIDAVVWHDGDLWRAALDTQSVEDDSDCGQLANFVPLTNYRIERKHGVFSKLDACAFVLNVYSDGNILSIVTDCSPHGTHVAGIA
AAFHPKEPLLNGIAPGAQLISCKIGDTRLGSMETGTGLIRALIAAVEHKCDLINMSYGEPTLLPDYGRFVDLVNEVVNKHRLIFVSSAGNGGPALSTVGA
PGGTTSSIIGVGAYVSPSMAAGAHSVVEPPSEGLEYTWSSRGPTSDGDLGVSISAPGGAVAPVPTWTLQKRMLMNGTSMASPSACGGVALLISAMKAEGI
PVSPYSVRKALENTSGPVGELPADKLSTGQGLMQVDRAHEYIRQSRNIPCICYEIMVNQSGKSTPTSRGIYLREASACQQPTEWTVQVQPKFHEGASNLE
ELVPFEECIELHSTEKVVVRAPEYLLLTNNGRSFNIVVNPTKLSEGLHYYEVYGVDCKAPWRGPIFRIPVTITKPMTVKNHPPFISFSRMSFLPGHIERR
YIEVPFGATWVEATMKTSGFDTTRRFFVDTVQICPLQRPMKWESVVTFSSPTAKSFAFPVVGGQTMELAVAQFWSSGIGSHETTIVDFEILFHGIAINKE
EIILDGSEAPIRIDAEALLSSENLVPAATLNKIRVPYRPVDAKLGTLTENRDKLPSGKQTLALTLTYKFKLEDGAEVKPQVPLLNNRIYDTKFESQFYMV
SDTNKRVYAMGDVYPSATKLPKGEYNLRLYLRHDNMQYLEKMKQLLLFIERNLDDKDVIRLNFFSEPDGPVMGDGAFKSSVLVPGKKEAIYLGPPVKDKL
PKNAPQGSVLLGAISYGKLSLAGQEGEESSQKNPVSYQISYVVPPNKVDEDKGKSSSTSLKTVSERLEEEVRDAKIRVLSSLKQDTDEERSEWKKLSTSL
KSDYPNYTPLLAKILEGLLSQSKVEDKIHHHEDVMDAADEVIDSIDKDELAKFFSLKSDPEDEETEKKKKAMETTRDELAEALYQKGLALVENESLKVRK
AETEGTKDLFEDNFKGLQKWVDAKSSKYGTLLVLRERRRGRLGAALKALNEMMQDNGDPPKKKLYELKLSLLDEIGWKHLSTYEKEWMLVRFPPSLPLF
|
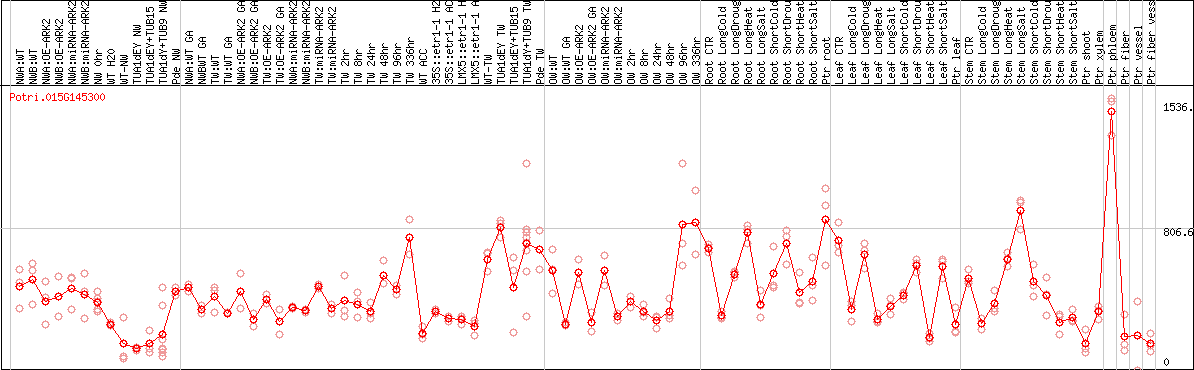
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G145300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
