External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09270 189 / 2e-60
ATGSTU8
glutathione S-transferase TAU 8 (.1)
AT2G29420 185 / 7e-59
GST25, ATGSTU7
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
AT2G29490 181 / 2e-57
GST19, ATGSTU1
GLUTATHIONE S-TRANSFERASE 19, glutathione S-transferase TAU 1 (.1)
AT2G29450 181 / 3e-57
ATGSTU1, AT103-1A, ATGSTU5
ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE TAU 1, glutathione S-transferase tau 5 (.1)
AT2G29470 176 / 3e-55
GST21, ATGSTU3
GLUTATHIONE S-TRANSFERASE 21, glutathione S-transferase tau 3 (.1)
AT2G29440 172 / 1e-53
GST24, ATGSTU6
GLUTATHIONE S-TRANSFERASE 24, glutathione S-transferase tau 6 (.1)
AT2G29460 171 / 2e-53
GST22, ATGSTU4
GLUTATHIONE S-TRANSFERASE 22, glutathione S-transferase tau 4 (.1)
AT2G29480 170 / 6e-53
GST20, ATGSTU2
GLUTATHIONE S-TRANSFERASE 20, glutathione S-transferase tau 2 (.1)
AT1G17180 157 / 5e-48
ATGSTU25
glutathione S-transferase TAU 25 (.1)
AT1G78320 156 / 1e-47
ATGSTU23
glutathione S-transferase TAU 23 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001491
246 / 5e-83
AT3G09270 185 / 5e-59
glutathione S-transferase TAU 8 (.1)
Lus10007897
238 / 2e-79
AT2G29420 187 / 1e-59
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10041654
227 / 4e-75
AT2G29420 175 / 1e-54
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10021102
197 / 2e-63
AT3G09270 245 / 2e-82
glutathione S-transferase TAU 8 (.1)
Lus10007896
194 / 3e-62
AT2G29490 171 / 2e-53
GLUTATHIONE S-TRANSFERASE 19, glutathione S-transferase TAU 1 (.1)
Lus10021103
192 / 1e-61
AT3G09270 250 / 1e-84
glutathione S-transferase TAU 8 (.1)
Lus10040727
189 / 4e-60
AT2G29420 216 / 6e-71
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10016471
183 / 5e-58
AT2G29420 234 / 8e-78
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10040764
182 / 1e-57
AT2G29420 229 / 6e-76
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10035241
182 / 2e-57
AT2G29420 188 / 5e-60
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0497
GST_C
PF00043
GST_C
Glutathione S-transferase, C-terminal domain
CL0172
Thioredoxin
PF02798
GST_N
Glutathione S-transferase, N-terminal domain
Representative CDS sequence
>Potri.016G023200.1 pacid=42810403 polypeptide=Potri.016G023200.1.p locus=Potri.016G023200 ID=Potri.016G023200.1.v4.1 annot-version=v4.1
ATGGCAGAAGAAGTTAAGGTGTTTAGAACATGGTCAAGTCCATTTGCTCTGAGAGTCATCTGGGCACTGAAATTGAAGGGTGTTGAGTTTGATACCATAT
ATGAAGATCTCTCCAACAAGAGCCCTTTACTCTTGCAATACAATCCTATCCACAAGAAGGTTCCCGTGCTTGTCCACAATGGTAAAGTCATCTGTGAATC
ACTTGTCATTCTAGAATACATCGACGAGACATGGAAGCAAAATCCTCTGTTGCCTGAAGATCCTCACCAGCAAGCCAGTGCTCGTTTCTGGGCCAAGTTT
GGTGATGATAAGGTTTTGCAATCAATTGTGTGGGGTGTGCTTCTGAAGGAGGGAAAAGAGCTAGAAGAAGGGGTCCTTGCATCCTTGGAGAACTTGAAAT
ATTTAGAAGAAGAGATAAGAGGAAAGAAATTCTTTGGTGGGGAGACTATTGGGCTAGCGGATATTGCATTAGGATGGCTTGCTTATTACCTTGATATCTT
TGAGGAGATACTAGGTCTAAAACTGATAGACCAGGAAAAGTTTCCATCCTTAGCGGCATGGAAGCAGGAATTTGCAAATGCTCCAATCATCCATGAGAAC
TGGCCGGATAGGGACAAGCTGGTTAACAAGTTTGTTGCCATGCGTGAGGCCAAACTTGGAAAAGAAACACCTAAATGA
AA sequence
>Potri.016G023200.1 pacid=42810403 polypeptide=Potri.016G023200.1.p locus=Potri.016G023200 ID=Potri.016G023200.1.v4.1 annot-version=v4.1
MAEEVKVFRTWSSPFALRVIWALKLKGVEFDTIYEDLSNKSPLLLQYNPIHKKVPVLVHNGKVICESLVILEYIDETWKQNPLLPEDPHQQASARFWAKF
GDDKVLQSIVWGVLLKEGKELEEGVLASLENLKYLEEEIRGKKFFGGETIGLADIALGWLAYYLDIFEEILGLKLIDQEKFPSLAAWKQEFANAPIIHEN
WPDRDKLVNKFVAMREAKLGKETPK
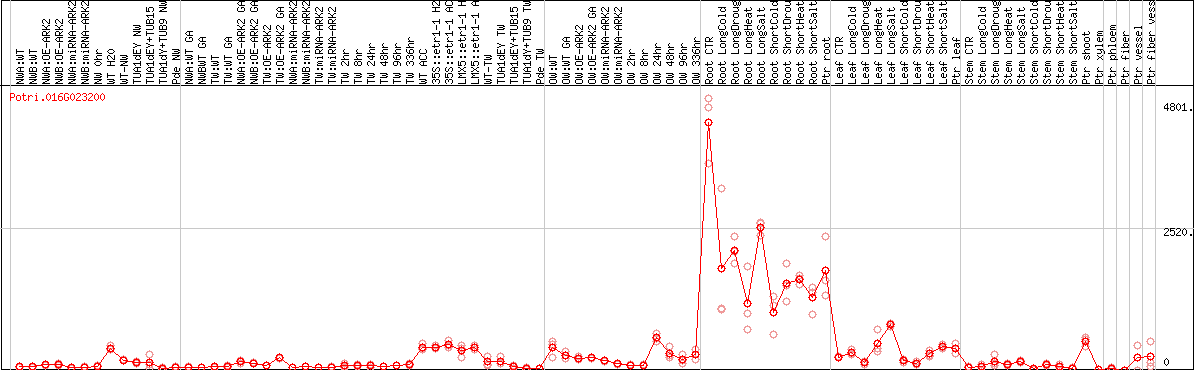
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G023200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.