Potri.016G029200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.016G029200.1 pacid=42810404 polypeptide=Potri.016G029200.1.p locus=Potri.016G029200 ID=Potri.016G029200.1.v4.1 annot-version=v4.1
ATGGGCAAAAACGTTGTTTCTGATGAAGATGAGGTGGAGTTCGAAGACGAAGAGAGAGAGCCTGTTGATGGAGATAGAATTGACCGCCGCCGCCACCACG
ACGACGACGACGACGAGGATGAAGAAGGGCAGGATGAATATGAGAAGGATGATTTTATAGTGGATGATGTTGAGGAAGAAGAGGAGGCGGATGAAGAAGA
GGAGAGGGCAGATAGTGATGAAGAGAGGCTCAAGAAGAAGAAGAAAAAGAAGAAAAGAGAAGCTGAGGATGTCCTTGATGAAGATGATTACGAGCTCCTG
CGGGACAACAATGTGTACCATCATCGGCCAAAGGATAGCAAAAAATTCAAGCGTTTGAAGAAAGCTCAGAGGGATTCAGATGAGGACCTTTCTGATGACG
AGTTTGATGGAAGCGGTAAAGGTGGGCGAACTGCTGAGGAGAAACTGAAGCGCAGCTTATTTGGTGATGATGAAGGTGTTCCACTTGAGGATATGCCTGA
AGAGGAGGAGCAAGAAGAAGTGGAAGAGGATGCAGACATTGGTGATGAGGATGAGATGGCTGATTTTATCGTGGATGAAGATGATGAGGATGGAACTTTA
GTGAGACGCAAGAAGCTAAAGAAAAAGAAATCTCGGCAGGCATCTGGAGCTTCATCTTCTGCTTTGCAAGAAGCTCAAGAAATATTTGGTGATGTTGATG
AACTCATACAGATGCGCAAGCAAGGTTTAGAATCTAGTGAATGGAGAGAAAGGAGATTGGAAGATGAATTTGAACCAACTGTCCTATTTGAAAAATACAT
GACTGAGAAAGATGATCAGATTCGGATGATTGATATTCCAGAAAGAATGCAGGTATCCGAGGAAAGCACTGGTCCTCCTCCATTGGATGATTTCAGCATA
CTGGAGGAGAGCAACTGGTTATATAGTCAGATTGCAAGCGGAACAGTACCTTTGTTTGCCAAGAATGGACTGTTTATAAACAAGGATGATGTTACCCGAT
TTTTGGAATTACATCACATTCAGAAGTTAGATATACCCTTCATTGCTATGTATCGGAAAGAGGAATGTTTAAGCCTGTTGAAAGATCCCGATCAACATGA
AGACAATGAAAATTATGATGACACTGATAAAAATCCCACATTCAAGTGGCACAAGGTTCTTTGGGCCATCCAGGACTTGGACAGGAAGTGGCTACTTCTT
CAGAAACGAAAGAGTGCCCTCAATTCATACTACAATAAGCGCTTTGAAGAAGAATCTCGACGGATATATGATGAAACGAGACTTAATTTAAATCAGCAGC
TGTTTGAGTCAATTCTTAAATCACTGAAGACTGCTGAATCAGAAAGAGAGGTTGATGATGTCGATGCAAAATTTAACCTGCATTTTCCACCAGGTGAGGT
TGGGGCTGATGAAGGACAGTACAAGAGGCCCATGCGGAGATCACAATACAGTATCTGTAGTAAGGCTGGCCTATGGGAAGTTGCAAGTAAATTTGGTTAC
AGTGCCGAGCAACTGGGAATGCAGTTATCTTTATTAAAGATGGAGGACGAGTTGCAGGATGCCAAGGAAACACCAGAGGAGATGGCTTCGAATTTTACAT
GTGCAATGTTCGAAAGTCCCCAAACAGTTCTTAAAGGGGCTAGGCACATGGCTGCAGTTGAGATAAGTTGTGAGCCCTGTGTCAGGAGATATGTCCGTTT
CATTTTTATGGACAATGCTGTTGTGTCTACAAGTCCAACGGCAGATGGGAATGCTGCCATTGATTCCTTCCATCAGTTTGCGGGTGTCAAGTGGTTGCGT
GAAAAGCCAATCAAGATGTTTGAGGATGCACAATGGCTCCTTATCCAAAAGGCTGAAGAGGAGAAACTTCTTCAAGTCACTGTTAAGCTTCCACAAAAAG
TTATGGATCAATTGATTGAGGACTGTAATGGTCGCTACTTGAGTGTTGGTGTCAGTAAATATGCTCAACTGTGGAATGAGCAAAGGAGTTTGATACTGAA
GGATGCACTCTTTGGTTTTTTGTTACCTTCAATGGAGAAAGAGGCTAGATCCTTGTTGGCTAGCAGAGCTAAAAACTGGTTGCTTTATGAATATGGGAAG
GTTCTGTGGAATAAAGTCTCTGTGGGACCATATCAACGGAAAGAAAGTGATGTTTCCATGGACGATGAAGCTGCACCTAGGGTGATGGCCTGCTGTTGGG
GTCCTGGAAAACCAGCAACTACTTTTGTGATGTTAGATTCATCTGGGGAGGTTTTAGATGTGCTTTACACTGGGTCCCTTACATTAAGATCTCAAAATGT
TAATGATCAGCAACGCAAGAAAAATGATCAGCAACGTGTTCTGAAATTTATGACAGACCACCAACCTCACGTGGTGGTTCTGGGAGCTGCACACTTGTCT
TGTACTAAACTGAAAGATGATATTTATGAGATTATCTTTAAAATGGTGGAGGAAAATCCTAGAGATGTTGGACATGAGATGGATGAACTGAGCGTTGTGT
ATGGGGACGAATCTCTCCCTCGTTTATATGAAAATTCTCGGATTTCTTCTGATCAACTACCTGGGCAGTCAGGCATTGTTAAACGGGCTGTTGCTCTTGG
CCGTTGTCTGCAAAATCCTTTGGCTATGGTTGCAACACTATGTGGTCCAGCCAGGGAGATTTTATCTTGGAAACTAAACCCTTTGGAGAACTTTCTTACC
CCTGATGAGAAATACTTGGTGATTGAACAAGTTATGGTAGATGCCACAAACCAGGTGGGCCTTGACATTAATTTGGCCACAAGCCATGAATGGTTATTCG
CTCCTTTGCAGTTCATTTCTGGGCTTGGACCTAGAAAGGCAGCATCTTTACAGAGGTCTCTAGTAAGAACTGGAGCGATTTTCACTCGCAAGGACTTTGT
TACAGCTCACGGTCTTGGTAAAAAGGTGTTTGTCAATGCAGTTGGTTTTTTGCGTGTCCGGAGGAGTGGCCTTGCTGCTAGCAGCAGCCAGTTCATTGAT
GTGTTAGATGATACGAGGATACATCCAGAATCCTATGGTCTCGCACAGGAGTTGGCAAAAGTTGTTTATGAAAAGGACAGTGGTGATGCAAACGACGATG
ACGATGCATTAGAGATGGCAATAGAATATGTTAGAGAGCGGCCTAATTTATTGAAAACATTTGCTTTTGATTTGTATTTTAAAGACAATAAGCGGGACAA
CAAGAAGGAGACTTTCAAAGATATAAAAATGGAACTGATACAAGGTTTCCAGGATTGGCGTAAACAGTATAAAGAACCAACCCAGGACGAGGAATTTTAT
ATGATTTCAGGTGAAACTGAGGATACCCTTGCTGAAGGGAGAGTTGTACAAGCCACGGTTCGCAGGGTGGTTGGTGGAAAGGCAATATGTGCACTGGAAA
CTGGTCTGACTGGGATTCTTACGAAGGAGGACTATGCTGATGATTGGAGGGATATACCTGAGTTGTCTGACAAGCTGCGCGAAGATGATATACTGACTTG
CAAGATCAAATCAATTCAAAAGAATAGATATCAAGTGTTCTTGGTTTGCAAAGACAGTGAGATGAGAAGTAACCGATATCGGCAGGTACAAAACCTTGAT
CTCTATTTTCATGAAGACCAAAGCAGTATGAGGAGTGAGCAGGAGAAGGTTCGCAAAGAGAGGGAGCTTGCAAAGAAACATTTTAAGCCTAGAATGATTG
TTCATCCACGCTTCCAGAATATAACTGCTGATGAAGCCATGGAGTTCTTATCCGACAAGGATCCCGGAGAAAGTATTATTCGTCCTAGTTCTCGTGGACC
TTCGTATTTAACTTTAACTCTTAAAGTTTATGATGGAGTTTATGCTCACAAAGACATAGTCGAAGGGGGGAAGGAACACAAAGATATCACAAGTTTACTT
CGTATTGGGAAGACATTGAAAATAGGAGAGGATTCTTTTGAGGATTTGGATGAGGTAATGGACCGCTATGTTGATCCCTTGGTGGGCCATCTAAAGTCAA
TGCTAAATTATCGCAAGTTCAGAAGTGGCACTAAAGCAGAAGTTGATGAGCTCCTAAGGATTGAGAAATCACAACAGCCTACTAGGATAGTTTATTCGTT
TGGCATTTCTCATGAACACCCAGGGACTTTCATTCTCACTTATATAAGGAGTACAAATCCACATCACGAGTATGTTGGTCTATATCCTAAGGGATTCAAG
TTTCGAAAGCGCATGTTTGAGGACATTGACCGGCTTGTGGCATATTTTCAAAAGCATATAGATGATCCCTTACATGAATCAGCACCATCAATCAGGTCAG
TTGCTGCCATGGTCCCAATGCGAAGCCCTGCAACAAGGGGTTCTTCCTGGGGTGGTTCAACGGATGAAGATGGCTGGAGAGGGCAATCTTTTGACAGAGA
TCGTTCTTCTGGTCCAGGTTCCAGGACAGGTCGAAATGATTATAGAAGTGGTGGTAGTCGAGATGGTCATCAGAATGGGCCCCCAAGACCATTTAGTGGG
CGAGGACGTGGTCGCGGATCGTATAACAGCACCAGAGGAAATAATAGTGGCAATGAGAGGCAAGATTCTGGTTATGACAAGCCCAGATGGGATTCAGGTA
CCAAGGATAACGATGAAGGTTGGGGCAGTTTCCCAGGTGCCAAAGTCCAAAACTCTCCTGGCAGGGAAGCCTTTCCTGGTGGCTGGGGAACTGGTGCAAG
TGGTGGTGATAATGGTGGTCGGGGCCATGGAAATGGTGGTGACACTGATAGTGGGAATGCTGGCTGGGGTAATACTAGCTTGAAAAGAGATTCAGCACAA
GGGGGCTGGGGTGCTGGTGGAGGTGGAAGTGGTGGTGGTGACTCTGGTCATGGAACTGGTGGTGGAGGCACTGACAATGGGAATTCTGGGTGGGGTAATA
GCTCTAAAAAAGGTGGGACACAAAGTGGCTGGGGTTCTGGTGGAAGAGGCGGTGGCGGCGATGTGGGTCAAGGAAGTGGTGGAGTCACTGACGATGGGAA
ATGGGGTACGGCCTCTAAGAGAGATTCTTCACAATCACAAGCCGACAATGGATGGTCAGGAGGTGGTGGTGGTTGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.016G029200.1 pacid=42810404 polypeptide=Potri.016G029200.1.p locus=Potri.016G029200 ID=Potri.016G029200.1.v4.1 annot-version=v4.1
MGKNVVSDEDEVEFEDEEREPVDGDRIDRRRHHDDDDDEDEEGQDEYEKDDFIVDDVEEEEEADEEEERADSDEERLKKKKKKKKREAEDVLDEDDYELL
RDNNVYHHRPKDSKKFKRLKKAQRDSDEDLSDDEFDGSGKGGRTAEEKLKRSLFGDDEGVPLEDMPEEEEQEEVEEDADIGDEDEMADFIVDEDDEDGTL
VRRKKLKKKKSRQASGASSSALQEAQEIFGDVDELIQMRKQGLESSEWRERRLEDEFEPTVLFEKYMTEKDDQIRMIDIPERMQVSEESTGPPPLDDFSI
LEESNWLYSQIASGTVPLFAKNGLFINKDDVTRFLELHHIQKLDIPFIAMYRKEECLSLLKDPDQHEDNENYDDTDKNPTFKWHKVLWAIQDLDRKWLLL
QKRKSALNSYYNKRFEEESRRIYDETRLNLNQQLFESILKSLKTAESEREVDDVDAKFNLHFPPGEVGADEGQYKRPMRRSQYSICSKAGLWEVASKFGY
SAEQLGMQLSLLKMEDELQDAKETPEEMASNFTCAMFESPQTVLKGARHMAAVEISCEPCVRRYVRFIFMDNAVVSTSPTADGNAAIDSFHQFAGVKWLR
EKPIKMFEDAQWLLIQKAEEEKLLQVTVKLPQKVMDQLIEDCNGRYLSVGVSKYAQLWNEQRSLILKDALFGFLLPSMEKEARSLLASRAKNWLLYEYGK
VLWNKVSVGPYQRKESDVSMDDEAAPRVMACCWGPGKPATTFVMLDSSGEVLDVLYTGSLTLRSQNVNDQQRKKNDQQRVLKFMTDHQPHVVVLGAAHLS
CTKLKDDIYEIIFKMVEENPRDVGHEMDELSVVYGDESLPRLYENSRISSDQLPGQSGIVKRAVALGRCLQNPLAMVATLCGPAREILSWKLNPLENFLT
PDEKYLVIEQVMVDATNQVGLDINLATSHEWLFAPLQFISGLGPRKAASLQRSLVRTGAIFTRKDFVTAHGLGKKVFVNAVGFLRVRRSGLAASSSQFID
VLDDTRIHPESYGLAQELAKVVYEKDSGDANDDDDALEMAIEYVRERPNLLKTFAFDLYFKDNKRDNKKETFKDIKMELIQGFQDWRKQYKEPTQDEEFY
MISGETEDTLAEGRVVQATVRRVVGGKAICALETGLTGILTKEDYADDWRDIPELSDKLREDDILTCKIKSIQKNRYQVFLVCKDSEMRSNRYRQVQNLD
LYFHEDQSSMRSEQEKVRKERELAKKHFKPRMIVHPRFQNITADEAMEFLSDKDPGESIIRPSSRGPSYLTLTLKVYDGVYAHKDIVEGGKEHKDITSLL
RIGKTLKIGEDSFEDLDEVMDRYVDPLVGHLKSMLNYRKFRSGTKAEVDELLRIEKSQQPTRIVYSFGISHEHPGTFILTYIRSTNPHHEYVGLYPKGFK
FRKRMFEDIDRLVAYFQKHIDDPLHESAPSIRSVAAMVPMRSPATRGSSWGGSTDEDGWRGQSFDRDRSSGPGSRTGRNDYRSGGSRDGHQNGPPRPFSG
RGRGRGSYNSTRGNNSGNERQDSGYDKPRWDSGTKDNDEGWGSFPGAKVQNSPGREAFPGGWGTGASGGDNGGRGHGNGGDTDSGNAGWGNTSLKRDSAQ
GGWGAGGGGSGGGDSGHGTGGGGTDNGNSGWGNSSKKGGTQSGWGSGGRGGGGDVGQGSGGVTDDGKWGTASKRDSSQSQADNGWSGGGGGW
|
DESeq2's median of ratios [POPLAR]
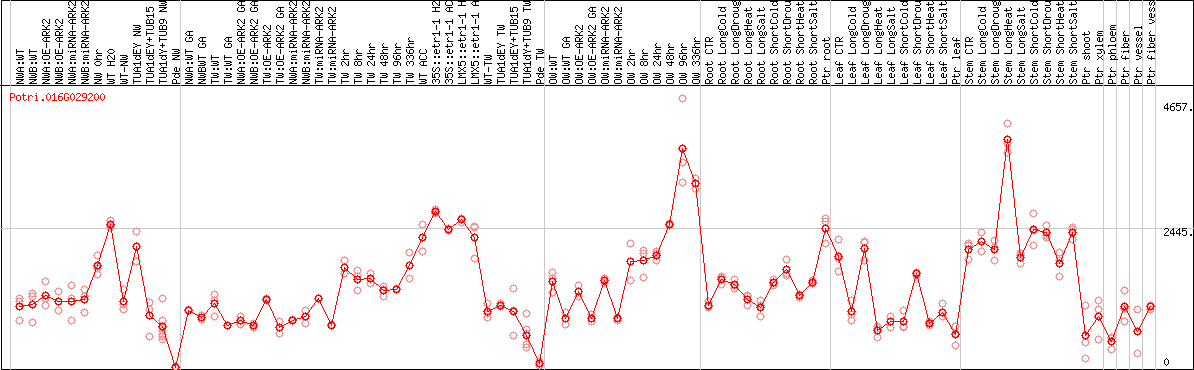
Coexpressed genes
Potri.016G029200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
