External link
Symbol
Pt-ORG2.3
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G56970 162 / 9e-49
bHLH
ORG2, bHLH038
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G56980 126 / 7e-35
bHLH
ORG3, bHLH039
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G04150 121 / 3e-33
bHLH
bHLH101
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G41240 108 / 2e-28
bHLH
bHLH100
basic helix-loop-helix protein 100 (.1.2)
AT1G71200 50 / 3e-07
bHLH
bHLH160
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G51780 47 / 2e-06
bHLH
bHLH036
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G20970 46 / 6e-06
bHLH
bHLH162
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G56960 45 / 3e-05
bHLH
bHLH041
basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT5G46760 44 / 6e-05
bHLH
bHLH005, ATR2, MYC3
Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT4G09820 42 / 0.0003
bHLH
BHLH42, TT8, bHLH042
TRANSPARENT TESTA 8, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G037600
312 / 8e-108
AT3G56970 184 / 9e-58
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.006G037200
266 / 8e-90
AT3G56970 168 / 2e-51
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.006G148800
54 / 4e-08
AT5G56960 204 / 4e-59
basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Potri.001G113400
46 / 7e-06
AT4G25410 124 / 6e-35
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.015G134300
45 / 2e-05
AT5G51780 143 / 1e-42
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G132000
44 / 6e-05
AT5G51790 154 / 6e-47
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G066500
41 / 0.0005
AT4G00050 303 / 5e-99
unfertilized embryo sac 10, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039006
110 / 1e-28
AT3G56970 142 / 1e-40
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027300
108 / 8e-28
AT3G56970 139 / 2e-39
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10010453
91 / 2e-21
AT3G56970 126 / 8e-35
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10003823
75 / 2e-15
AT3G56970 129 / 1e-35
OBP3-RESPONSIVE GENE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10041493
48 / 4e-06
AT3G25710 201 / 2e-62
TARGET OF MONOPTEROS 5, basic helix-loop-helix 32 (.1)
Lus10017152
48 / 4e-06
AT5G56960 196 / 4e-56
basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
Lus10011172
45 / 8e-06
AT4G20970 144 / 7e-44
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10031677
46 / 1e-05
AT5G51780 143 / 2e-42
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027396
45 / 2e-05
AT4G25400 89 / 6e-22
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10027394
44 / 6e-05
AT5G51780 149 / 9e-45
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.016G037300.1 pacid=42810084 polypeptide=Potri.016G037300.1.p locus=Potri.016G037300 ID=Potri.016G037300.1.v4.1 annot-version=v4.1
ATGTTAGAAGAATTATCTCCCATCAGTTTGTTCTCAACATTTGGATGGCCCTTGGAGGAAGCCATAAGCCATGAACAGCTCTACAGCTTTAGAGATGGTG
AAACTCCAGAGTCATTTACTCACTTCCCTCCATCTCAGCCAGATGTAAGACAGCTTGATCGCTCCACATCATTCACGGCCCACAGTGGAAGCGGTGACCC
TAGCATGGCTAAGAAGCTTAACCACAACGCTAGCGAACGTGACCGTCGCAAAAAGATCAACAGTTTGTATTCTTCACTCCGTTCACTACTTCCTGCAGCC
GATCAAAGGAAGAAATTAAGCATACCGTATACAGTTTCACGTGTGCTTGTATACATACCAAAACTTCAACAACAAGTGGAGAGACTGATTCAAAGGAAGG
AGGAGCTTCTATCGAAGTTATCTAGGCAAGCTGACGATTTAACTCATCAAGAAAATCAAAGAAAAGGCACCATGTATAGCTCTTTATCATCGGTATCGGC
GAGCCGGCTCAGTGACAGGGAAGTTGTCATTCATATCTCAACTAACAAGCTCCATAGAAGTTCATTATCAGAAATCTTGGTTAATTTAGAGGAGGCTGGA
CTTCTTCTACTAAATTCTTCTTCCTTCGAGTCCTTTGGAGGCAGAGTCTTCTATAATTTACACCTTCAGGCCATGGAAGGAACTTACACAGTAGAGTGCG
AGGCCTTGAATGAGAGGCTTGTGTCCTTGTGCGAGAAGAGGGAGTCATTGTTTCCATTAAATTCAAGTTCTCCATATTCCAACTGTGTATTCTAG
AA sequence
>Potri.016G037300.1 pacid=42810084 polypeptide=Potri.016G037300.1.p locus=Potri.016G037300 ID=Potri.016G037300.1.v4.1 annot-version=v4.1
MLEELSPISLFSTFGWPLEEAISHEQLYSFRDGETPESFTHFPPSQPDVRQLDRSTSFTAHSGSGDPSMAKKLNHNASERDRRKKINSLYSSLRSLLPAA
DQRKKLSIPYTVSRVLVYIPKLQQQVERLIQRKEELLSKLSRQADDLTHQENQRKGTMYSSLSSVSASRLSDREVVIHISTNKLHRSSLSEILVNLEEAG
LLLLNSSSFESFGGRVFYNLHLQAMEGTYTVECEALNERLVSLCEKRESLFPLNSSSPYSNCVF
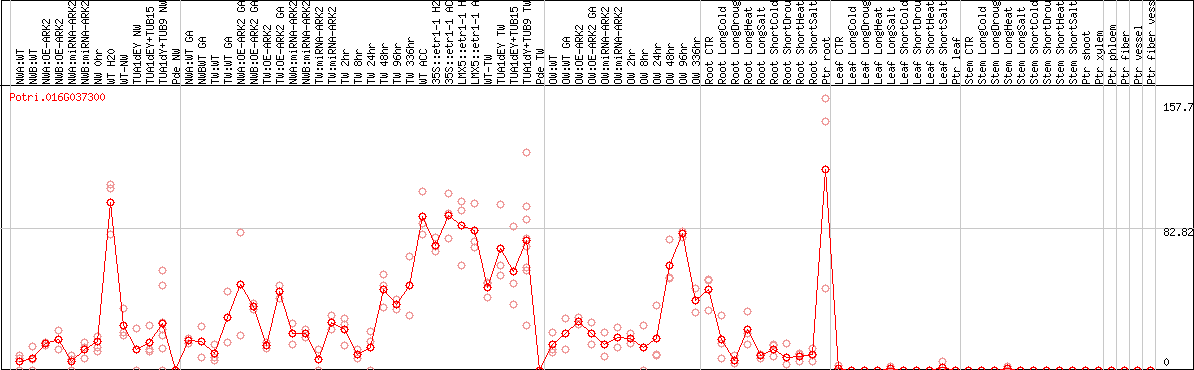
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G037300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.