External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G57080 287 / 1e-98
RPB5B, NRPE5
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, B, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
AT2G41340 276 / 2e-94
RPB5D
"RNA polymerase II fifth largest subunit, D", RNA polymerase II fifth largest subunit, D (.1)
AT3G54490 209 / 5e-68
RPB5E
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
AT3G22320 134 / 7e-39
RPB5A, NRPD5, NRPB5, ATRPABC24.3
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, A, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
AT5G57980 111 / 4e-30
RPB5C
"RNA polymerase II fifth largest subunit, C", RNA polymerase II fifth largest subunit, C (.1)
AT3G16680 42 / 3e-05
DNA binding;DNA-directed RNA polymerases (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G025500
252 / 7e-85
AT3G54490 249 / 1e-83
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Potri.006G161600
144 / 6e-43
AT3G22320 365 / 2e-130
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, A, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
Potri.T127006
91 / 3e-23
AT3G54490 92 / 5e-24
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Potri.003G200450
89 / 1e-22
AT3G54490 92 / 4e-24
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Potri.001G179300
71 / 1e-15
AT3G54490 92 / 6e-24
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018022
330 / 2e-115
AT3G57080 280 / 6e-96
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, B, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
Lus10042019
294 / 2e-101
AT3G57080 239 / 4e-80
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, B, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
Lus10024169
206 / 2e-66
AT3G54490 221 / 7e-73
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Lus10039554
183 / 3e-57
AT3G54490 184 / 6e-58
"RNA polymerase II fifth largest subunit, E", RNA polymerase II fifth largest subunit, E (.1)
Lus10013639
142 / 7e-42
AT3G22320 342 / 3e-121
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, A, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
Lus10001335
142 / 7e-42
AT3G22320 342 / 3e-121
RNA POLYMERASE II FIFTH LARGEST SUBUNIT, A, Eukaryotic rpb5 RNA polymerase subunit family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01191
RNA_pol_Rpb5_C
RNA polymerase Rpb5, C-terminal domain
CL0236
PDDEXK
PF03871
RNA_pol_Rpb5_N
RNA polymerase Rpb5, N-terminal domain
Representative CDS sequence
>Potri.016G038500.1 pacid=42809265 polypeptide=Potri.016G038500.1.p locus=Potri.016G038500 ID=Potri.016G038500.1.v4.1 annot-version=v4.1
ATGGAGCAAAATTTACTAACGGAGAGACCGAATGAAGAAGAAGGAGGAGCTCATCAAGAAAAAGAAAATGGGAGTTTTGGAATGGAGTCTCTGGGGAGGT
GCTTGAGCAGCTTTGTTGATGAAGGTAGCACAGAGAGCCACAGATACTATCTTTCCAGAAGAACTGTGCTTGAGATGTTAAAAGATAGAGGCTATTCTGT
TCCAAGTTCAGAGATTGACATTTCTTTGCAAGATTTTAGAGGCGTTTATGGTCAAAACCCTGATATTGAACTCCTCAAGTTTTCTGCCACCCATAAATCA
GATCCTTCAAAAAGGATGCTGGTGATTTTCTGTGGCCTTGGTGTGGTTAAAGTTGGCATGATTCGTCTCATTACGGTTCAGATTACTGACAGAGACAGTT
TGACTGGTCTGATATTAGTCTTGCAAAACAACATAACTAATCAAGCTATGAAAGCTCTGGATCTTTTCAAATTCAAGATTGAAATATTCCAGATAACTGA
CTTGCTTGTTAATATCACAAAGCATATATTAAAGCCAAAGCATCAGGTGCTAAGTGAGCAGGCAAAACAAAGGCTCTTGAAGAAGTACAGTATAGAAGAA
AAGCAGCTTCCTCGACTGTTAAAGAAAGATGCAATTTCACGATACTATGGCCTCGAGAGGGGACAGGTGGTGAAGGTCACCTATGATGGTGATATTACGG
GGTCACATGTAACTTATCGCTGTGTCTGGTGA
AA sequence
>Potri.016G038500.1 pacid=42809265 polypeptide=Potri.016G038500.1.p locus=Potri.016G038500 ID=Potri.016G038500.1.v4.1 annot-version=v4.1
MEQNLLTERPNEEEGGAHQEKENGSFGMESLGRCLSSFVDEGSTESHRYYLSRRTVLEMLKDRGYSVPSSEIDISLQDFRGVYGQNPDIELLKFSATHKS
DPSKRMLVIFCGLGVVKVGMIRLITVQITDRDSLTGLILVLQNNITNQAMKALDLFKFKIEIFQITDLLVNITKHILKPKHQVLSEQAKQRLLKKYSIEE
KQLPRLLKKDAISRYYGLERGQVVKVTYDGDITGSHVTYRCVW
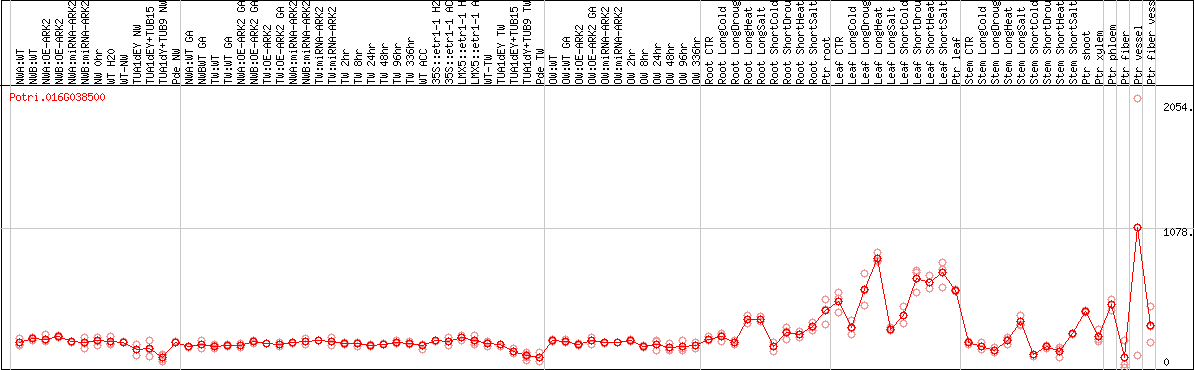
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G038500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.