CHR929 (Potri.016G043500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CHR929 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.016G043500.1 pacid=42809420 polypeptide=Potri.016G043500.1.p locus=Potri.016G043500 ID=Potri.016G043500.1.v4.1 annot-version=v4.1
ATGGACAACAGGAGACAAGCCAAGGACTCGCTTTCTTACTCCAATTTGTTCAATCTTGAGTCTTTGGTGAACTTCAGAGTTCCACAACCAGATGATGAAT
TTGATTATTATGGGAATAGTAGTCAGGATGAGAGCAGAGGTAGCCAAGGTGGAGCAATGTCAAAATTTGTTAATGGGAATCTATCGGAGAGGGAATTGAG
TTCAGGGAAGAGAAAGAGGCGGTATAACAACAGTGAGGGAGAGGAGGAAGACGGATATTCGGGGGCACGCATTACTGAGGAACAGTATCGATCAATGTTA
GGAGAACATATCCAGAAGTACAAAAGGAGGTATAAGGATTCCTTGTCAAGTCCTGCACCTCCTCCTCGGATGGGGATTCCCGTTCCAAAGAGCAGTTTGG
GTGGTTCTAAAACTAGGAAGCTGGGAAGTGAGCAGCGAGGGGGTTTATATGATATGGAAACCACATCTGAATGGGTCAATGACATTGTACCATCAAAACG
CGGAGACTATCATGAGCCAGAGTTCACGCCCAAAATATACTATGAACCTCCTTATTTGGATATTGGGGATGGTGTCACTTATAGGATACCCCCCTCTTAT
GATAAGCTGGCGGCATCATTGAACTTACCAAGCTTCTCAGATATGCGGGTGGAGGAATTTTACTTGAAGGGCACCTTAGATTTGGGTTCCTTGGCAGCAA
TGACAGCCAATGATAAAAGGTTTGGACTTAGAAGCCGAGCGGGAATGGGAGAGCCCCAGTTACAATACGAGTCACTTCAGGGAAGGTTGAAGGCATTGGC
AGCTTCTAACTCTGCTGAGAAATTCAGTCTCAAGATATCCGAAGAGGCATTGAATTCCTCAATCCCAGAGGGGGCAGCTGGAAATATAAAACGCTCTATT
TTGTCAGAGGGCGGTGTAATGCAGGTTTACTATGTTAAAGTTTTGGAGAAAGGAGACACATATGAGATTATTGAGCGGAGTCTACCTAAGAAGCCAAAAA
TAATTAAAGACCCTTCTGTTATTGAGAGGGAGGAAATGGAAAGGATTGGTAAAGTTTGGGTAAACATTGTGAGAAGAGATATACCTAAGCATCATAGGAT
TTTCACTACGTTTCATCGGAAGCAACTAATTGATGCAAAGAGGTTCTCAGAAAACTGTCAAAGAGAGGTCAAATTGAAGGTTAGCAGATCACTCAAAATA
ATGAAGGGTGCTGCAATTCGCACTAGGAAGTTAGCTAGAGACATGCTACTGTTTTGGAAGCGAGTGGATAAGGAGATGGCAGAGGTGAGGAAAAAGGAGG
AAAGAGAAGCTGCAGAAGCTTTGAAGCGTGAACAAGAGCTCAGGGAAGCAAAAAGACAGCAGCAAAGGCTCAATTTTCTCATACAACAAACAGAACTGTT
CAGTCATTTCATGTCAAACAAACCAAATTCACAGCCGTCAGAAGCTTTGCCAATTGCAGATGAAAAGACAGATGATCAAGTAATGGATTGTAGCACTGCT
GAAGCTGGACCTGATCCTGAAGAAGATCCTGAGGATGCTGAATTGAGAAAAGAGGCTCTCAAAGCTGCTCAAGATGCAGTCTCCAAGCAGAAACTGTTAA
CAAGTGCTTTTGATAGTGAGTGTTCAAAGCTGCGTGAAGTTGCTGATATTGAGGGACCTATAACTGATGCCTCAGTTGCAGGATCTAGCAACATTGATTT
GCAGACACCCTCCACCATGCCTGTTACATCAACAGTTAAGACACCAGAGTTGTTTAAAGGCAGCCTTAAAGAATATCAGTTGAAGGGTCTTCAGTGGCTG
GTTAATTGTTATGAGCAGGGCTTAAATGGTATACTGGCTGATGAGATGGGCCTTGGAAAGACCATTCAAGCCATGGCATTCTTGGCTCATTTGGCTGAGG
AAAAAAATATCTGGGGTCCTTTTCTGATTGTTGCTCCTGCTTCTGTCTTGAATAACTGGGCTGATGAAATCAGTCGTTTCTGCCCTGACTTGAAAACCCT
TCCATATTGGGGTGGGCTTCAAGAGCGGATGGTACTTCGGAAAAACATTAATCCAAAACGCCTTTATCGAAGGGAGGCTGGTTTTCACATTCTCATTACC
AGCTATCAATTGCTAGTGTCTGATGAGAAGTATTTTCGGCGTGTGAAATGGCAATATATGGTACTAGATGAGGCCCAGGCTATTAAGAGTGCCAACAGTA
TAAGATGGAAGACACTGCTCAGTTTTAATTGTCGTAATCGCCTACTGCTGACTGGTACACCAATTCAGAATAACATGGCAGAATTGTGGGCCCTCCTACA
TTTCATCATGCCCACATTATTTGACAGCCACGAACAGTTCAATGAGTGGTTTTCTAAAGGAATTGAGAATCATGCAGAACATGGAGGTACTTTAAATGAG
CACCAGCTAAACAGATTGCATGCAATTCTGAAGCCTTTTATGCTGCGACGTGTGAAGAAAGATGTGGTTTCTGAGTTAACTAGGAAAACAGAAGTTACTG
TACACTGCAAGTTGAGCTCTCGACAACAAGCTTTCTATCAAGCTATTAAGAACAAGATATCTCTTGCTGAGTTGTTTGATAGCAATCGTGGACATCTTAA
TGAGAAGAAAATTATGAATCTCATGAACATAGTCATTCAGCTGAGAAAGGTTTGTAACCATCCAGAGTTGTTTGAAAGGAATGAGGGAATCACATATTTC
TATTTTGGAGAGATTCCTAATTCTTTCCTGCCCTCTCCTTTTGGGGAGTTGGAGGACATACATTATTCAGGAGGTCGAAATCCTATTACGTATAAGATTC
CAAAAGTAGTCCACAATGAGATTGTCCAAAGTTCTGAGGTACTTTGCTCAGCAATTGGGCGTGGTTTTGGCAGAGAGTCATTTCAAAAGCATTTTAATAT
ATTCTCATCAGAAAATGTGTATCGATCTGTATTTGCACTGGACAATAGCTCAGATAGCTTGCTTATTAAGAGTGGGACGTTTGGTTTTTCTCATTTGATG
GATTTATCCCCAGCGGAGGTTGCATTTTTGGCAATTAGTTCATTTATGGAGAGGCTGTTGTTTTTTATTATGAGATGGGGCCGGCGATTTTTGGATGGAA
TCTTGGACTTGCTCATGAAAGACATAGAAAATGACCATAGCAATTACCTTGAAAAACATAAAGTTAGAGCTGTTACACGGATGTTATTGATGCCATCAAG
ATCTGAGACAGATATCCTAAGGAGGAAAATGGCAACAGGCCCTGCTGATACTCCATTTGAAGCTCTAGTTAATTCTCATCAGGACAGGCTTCTGTCCAAC
ATCAAACTTCTCCACTCAACTTACACGTTCATCCCACGGACCAGAGCACCACCGATCGGTGGCCAATGCTCAGATAGAAACTTTGCCTACCAAATGATGG
AAGAATTACACCAACCCATGGTTAAGAGGTTGTTAACTGGGTTTGCACGCACATCCACATTTAATGGACCTAGAAAGCCAGAGCCTCTTCATCCTTTGAT
TCAGGAGATTGATTCTGAACTACCTGTTTCACAACCTGCCCTTCAGTTGACATACAAGATTTTTGGTTCTTGTCCCCCCATGCAGAGCTTTGACCCTGCA
AAGTTGCTCACGGATTCTGGAAAGCTTCAAACACTTGATATATTACTGAAACGATTGCGAGCAGAAAACCACCGAGTTCTATTGTTTGCTCAAATGACAA
AAATGCTGAATATTCTCGAGGACTACATGAATTATAGGAAATATAGATATCTTAGACTTGATGGATCTTCCACCATTATGGATCGTCGGGACATGGTCAG
AGACTTCCAGCTTAGGAATGATATTTTTGTTTTCTTGCTAAGTACCAGAGCTGGTGGGTTGGGTATTAATTTGACAGCTGCAGACACTGTTATCTTTTAT
GAAAGCGATTGGAATCCTACACTAGATTTACAGGCAATGGATAGAGCACATCGGCTGGGCCAGACGAAGGATGTCACTGTCTACCGGCTCATTTGTAAAG
AGACAGTTGAGGAAAAGATTTTACAAAGAGCAAGTCAGAAAAATACTGTGCAGCAGCTTGTTATGACAGGTGGTCATGTTCAGGATGATCTCTTGGCTCC
TGAGGATGTGGTGTCCTTGCTTCTGGATGATGCCCAGCTGGAGCAGAAGTTGAGAGAAATTCCACTGCAGGCCAGGGATAGACAAAAGAAAAAACCAACG
AAGGCAATAAGGGTAGATGCTGAAGGGGATGCAACTTTCGAAGATTTGACAGAAACTGTTGCTCAGGGCACGGGAAATGAACAATCCGAGGATGCAGAGA
AATTGAAATCACCCAATAGCAATAAGAGGAAAGCTGCTTCTGACAAGCAGATTACATCAAAGCCAAGGAATTCACAGAAGAATGAACCAAATTCTTCACC
GATGGACTATGAACTGGATGATCCTTTTCCAAATTCTGAGCCACAGTCACAGAGACCCAAGCGGTTGAAGAGGCCAAAGAAGAGTGTAAATGAGAAACTC
GAACCAGCATTTACTGCCACACCCTCAATCGATTCATCACAGATCCAATATCCACCTACAAATAATCTTGCTTCAACATATTCTAACACTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.016G043500.1 pacid=42809420 polypeptide=Potri.016G043500.1.p locus=Potri.016G043500 ID=Potri.016G043500.1.v4.1 annot-version=v4.1
MDNRRQAKDSLSYSNLFNLESLVNFRVPQPDDEFDYYGNSSQDESRGSQGGAMSKFVNGNLSERELSSGKRKRRYNNSEGEEEDGYSGARITEEQYRSML
GEHIQKYKRRYKDSLSSPAPPPRMGIPVPKSSLGGSKTRKLGSEQRGGLYDMETTSEWVNDIVPSKRGDYHEPEFTPKIYYEPPYLDIGDGVTYRIPPSY
DKLAASLNLPSFSDMRVEEFYLKGTLDLGSLAAMTANDKRFGLRSRAGMGEPQLQYESLQGRLKALAASNSAEKFSLKISEEALNSSIPEGAAGNIKRSI
LSEGGVMQVYYVKVLEKGDTYEIIERSLPKKPKIIKDPSVIEREEMERIGKVWVNIVRRDIPKHHRIFTTFHRKQLIDAKRFSENCQREVKLKVSRSLKI
MKGAAIRTRKLARDMLLFWKRVDKEMAEVRKKEEREAAEALKREQELREAKRQQQRLNFLIQQTELFSHFMSNKPNSQPSEALPIADEKTDDQVMDCSTA
EAGPDPEEDPEDAELRKEALKAAQDAVSKQKLLTSAFDSECSKLREVADIEGPITDASVAGSSNIDLQTPSTMPVTSTVKTPELFKGSLKEYQLKGLQWL
VNCYEQGLNGILADEMGLGKTIQAMAFLAHLAEEKNIWGPFLIVAPASVLNNWADEISRFCPDLKTLPYWGGLQERMVLRKNINPKRLYRREAGFHILIT
SYQLLVSDEKYFRRVKWQYMVLDEAQAIKSANSIRWKTLLSFNCRNRLLLTGTPIQNNMAELWALLHFIMPTLFDSHEQFNEWFSKGIENHAEHGGTLNE
HQLNRLHAILKPFMLRRVKKDVVSELTRKTEVTVHCKLSSRQQAFYQAIKNKISLAELFDSNRGHLNEKKIMNLMNIVIQLRKVCNHPELFERNEGITYF
YFGEIPNSFLPSPFGELEDIHYSGGRNPITYKIPKVVHNEIVQSSEVLCSAIGRGFGRESFQKHFNIFSSENVYRSVFALDNSSDSLLIKSGTFGFSHLM
DLSPAEVAFLAISSFMERLLFFIMRWGRRFLDGILDLLMKDIENDHSNYLEKHKVRAVTRMLLMPSRSETDILRRKMATGPADTPFEALVNSHQDRLLSN
IKLLHSTYTFIPRTRAPPIGGQCSDRNFAYQMMEELHQPMVKRLLTGFARTSTFNGPRKPEPLHPLIQEIDSELPVSQPALQLTYKIFGSCPPMQSFDPA
KLLTDSGKLQTLDILLKRLRAENHRVLLFAQMTKMLNILEDYMNYRKYRYLRLDGSSTIMDRRDMVRDFQLRNDIFVFLLSTRAGGLGINLTAADTVIFY
ESDWNPTLDLQAMDRAHRLGQTKDVTVYRLICKETVEEKILQRASQKNTVQQLVMTGGHVQDDLLAPEDVVSLLLDDAQLEQKLREIPLQARDRQKKKPT
KAIRVDAEGDATFEDLTETVAQGTGNEQSEDAEKLKSPNSNKRKAASDKQITSKPRNSQKNEPNSSPMDYELDDPFPNSEPQSQRPKRLKRPKKSVNEKL
EPAFTATPSIDSSQIQYPPTNNLASTYSNT
|
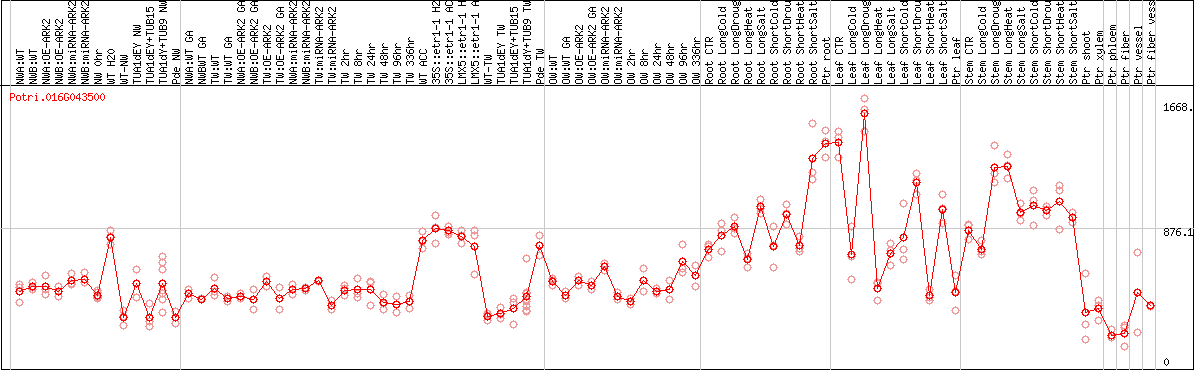
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G043500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
