Potri.016G069100 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.016G069100.2 pacid=42809430 polypeptide=Potri.016G069100.2.p locus=Potri.016G069100 ID=Potri.016G069100.2.v4.1 annot-version=v4.1
ATGGGTCGACCCGAACCCAGCGTCTTGTTCTCTCAAACTTTCGTTCATCCTCAACTGGATGAGTACGTCGACGAGGTATTGTTCGCCGAACCAATTGTGA
TTACTGCGTGTGAGTTTCTTGAACAGAATGCTTCTTCTGCGTCACAAGCAGTGTCGGTACTGGGGGCTACGTCACCTCCTTCATTTGCTTTGGAGGTGTT
TGTTAAGTGTGAGGGGGAGACAAGATTTAGACGACTCTGCCAGCCATTCCTCTATTCCCATTCGTCATCACATGTGCTAGAAGTTGAGGTGGCTGTTGTA
ACTAACCATCTGGTTGTGAGGGGCAGCTATCGCAGTCTAAGTTTGGTCATATATGGGAACACAGCTGAGGATCTGGGGCAGTTCAGCATTGAATTTGATG
ATAGCTCTTTAACTAATCTTGTTAGTTCTGCTGAGGGAAAGCTCGAGGATCTCCCTTTGGCATTACATTCAACTAATCGTACTGTTGAGGATTCTCTATC
CTCTCTGAATGTATTATCTTTACCTGTTGCTGCTTCACATATATCTGCTGAGGTTAAGCAATTTTTGCAGCTGATTTTGAAGTTGCTGGAGCTCCCGAAC
CTTAGTGATTCTGTACATAGAGTTCTTACTACTGTAGTGAAAGCTGTTTGTTCCTTTGTCACCCGTGATTTGTGTTGTGAAACAGTAAATCAGAAGCACA
TTAAAATGTGTGGATCCAAAAACATTGAAGAGTTTCACCATGTTATCAATGAGGCTAGAAATGAGCTTCTCCAAGTGCTTGGACAAGTCTTAGGAGATGA
ATCGGCTGAACTTTTGGCAGACTGCACATTTCTTGAGTCGGAGGCTGACTTGGCTACTTCTAAACAGCTGGTGGACATGTTAAGCCAGTATTTCTCTTTT
GAAAGGAACTCCACAAATGTTGGAGCTTGCCAGCTTTCACAGAACAAGAGTGTCATTCTGGGTCTGAGCTTGGCTCTTTTATTGTGCTCTGGTAGAGAAA
GCTGCTTTCACTTCGTCAGTAGTGGGGGAATGGAGCAGCTTGCACATATTTTTTCTAATGAGGTGCAGAATTCTTCCGCCATAATTTTACTATCACTTGG
AGTTGTTGAGCAGGCAACAAGGCATCCAATTGGCTGTGAAGGATTTTTAGGTTGGTGGCCTCGTGAAGATGAAAATATCCCATCTGGTACTAGCAAAGGT
TATAGTCAATTGTTGAAGTTGGTCCTACAGAGACCTCAGCATGATGTTGCTTCTCTTGCAACCTATGTCCTTCATCGGTTACGATTTTATGAAGTTGTTT
CTAGATATGAGTTTTCTGTTCTTTCTGCTCTAGGGGGACTTTCTGCACTTGGAAGAGTTACAAGTGTTACGTCAGCAATGCTTAACAGTGCTAAATCTCA
ACTTAAGATGCTCTTGAAATTGATAAATTTGCGTGGTCCAATTGAAGATCCTTCTATAGCAGCCTCTGCAAGTAGGTCATTGATCATTGGTCAAACAGAA
GGATTATTGTCATATAAAGCAACCAGTAACTTGGTTGGTTCATCACATTGCTGCTTTTCAAACTGGGATATTGATTCCCATTTGCTGGCACTTCTCAAGG
AGAGAGGCTTTCTTCCTTTGTCAGCTGCACTGTTGTCGTCCCCCATATTGCGTTCAGAAGCTGTAGATGCCATGGATACATTTGTTGATATCGCATCAAC
TATTGGGGCCATACTTCTTTCACTTCTTATGTGTCGCTCAGGTTTAATTTTTCTCTTAAATTACCCTGAGCTTTGCACTACACTAATTGATGCTTTAAGG
GGTGTGGGTGGTATGAATAGGGAAGAATGTGTTCCTCTACGTTATGCATCTGTTTTATTGTCCAAGGGTTTCGTTTGCAGTCCACATGAAGTTGGAGTGA
TTGTTGAGACGCATTTGAGAGTGGTTAATGCTATTGATCGTTTGCTAATATCAACTCCACATCCCGAAGAGTTTCTATGGGTATTGTGGGAGCTTTGTGG
CCTTTCCAGGTCAGATTGTGGGCGCCAGGCTTTGTTGGTTCTGGGATATTTTCCTGAGGCCATTTCAATCTTGATTGAAGCCTTACATTCTGTCAAGGAG
TCAGAGCCAGTTGCTAGTGGAGCTTCTCCAATAAATCTTGCGATTTTTCACTCGGCTGCAGAGATTTTTGAAGTCATTGTCACTGATTCAACTGCATCTT
CCCTTGATTCTTGGATTGGACATGCTATGGAACTTCATAAAGCGTTACATTCTTCTTCCCCTGGTTCCAATCGGAAAGATACTCCAACTAGACTGTTGGA
GTGGTTTGATGCTGGTGTAGTTTATCATAAAAACGGGGCTATTGGTCTTCTACGGTATTCTGCTGTGTTGGCTTCTGGTGGAGATGCCCATTTAACCTCC
ACCAGCATTTTGGTGGCTGATTTGACAGATGTTGAGCAAGTTGTTGGAGATGCATTGGGTGGTTCGGACATAAATGTTATGGATAATCTTGGAAAGTTGA
TCTCTGATAAGTCTTTTGAGGATAATCCTCTTCGTGACTCATCCATAACACAGATGACAACAGCCATTCGGATACTGGCCTTTGTGTCAGAAAACTCAAC
TGTTGCTGCTGCTCTGTATGATGAAGGTGCTCTCATAGTTATTTATGCAATTCTGATCAAGTGCAGCTTGATGCTTGAAAGGTCATCAAACAGTTATGAT
TACCTTGTTGATGAGGGTACAGAACGCAATTCTACATCTGATTTGCTATTGGAACGTAATCGTGAGCAGAGCTTGGTTGATCTTTTGGTACCTACTCTGG
TATTGCTGATCAATCTTTTGCAGAAATTACAAGAAGCTAAAGAGCAGCACAGGAATACAAAACTAATGAATGCCCTTTTGCGACTGCACAGAGAAGTTAG
TCCAAAGCTAGCTGCAAGTGCTGCAGACTTATCATCTCCCTATCCTGATTCTGCACTTGGTTTTGGAGCTGTCTGCCATCTTGTTGTATCAGCACTTACT
TGTTGGCCCTTATATGGCTGGACTCCTGGGCTTTTCCACTCTCTTCTTGCAAATGTTCAAGCTACTTCACTGTTGGCCTTGGGTCCAAAGGAAACCTGCA
GTTTGCTGTGTCTTCTGAATGATTTATTTCCTGAAGAAGGTGTCTGGCTTTGGAAAAATGGAATGCCCATGTTAAGTGCGCTAAGAAAATTGGCTGTTGG
GACATTATTGGGGCCACAGAAGGAGAAACAGGTTGACTGGTATCTGGAGACTTCACACCGTGAGAAACTGCTTAACCAACTGACACCACATCTCGATAAA
ATTGCACAGATTATCGAACATTATGCTATATCTGCATTGGTGGTCATTCAAGACATGCTTCGGGTTTTCATAATTCGAATTGCTTGCCAAAAGATTGAGT
ATGCTTCTTTGCTTCTGCAGCCCATTTTATGCTGCATCCGCAATCATCTTTCTGACTTGACTTCTCCATCAGAGATTGATGCTTACAAGGTTTACAGATA
TCTTGATTTTCTTGCGAGCATATTGGAACATCCATGTGCAAAGGAACTGTTATTGGAGGAAGGCATTGCTGAGATGCTAACACAAGTGCTGGAAAGGTGT
TTAGTTGCTATCGGTTCAGATGGAAAACAGATCTCAGATAGTAAAATTTCAGCCAAGTCTGGGTTTACTTTGATCAGTTGGTGCTGTCCCGTGTTCAAGT
CCTTCTCACTTCTTTGTGTTCCTCGAACACCTTTACCATACCCCGTGAGACATGATTTGCACAGTTCTGCAAGTTTAAGTGCTAAAGACTGTTCATTGAT
TCTACCATATCTGCTTAAGTCCTGCCAGGTCTTACCGGTTGGAAAAGAGTTGCTTTCTTGCCTTGCATTCTTTAAAGATTTAGGTTCCTGCAATGAAGGT
CAAAGTGCTTGTGTGACGACTTTGCATCACATCAACACCAGCATTGAGGAACATGAATCTGGAAAAGGACAGGAAAGGAATGGGAATTATAATTTGGATG
ACATTGAATGGAGAAAGCATCCTCCTTTGCTTTCTTGCTGGATTAGGTTGTTGGAATCAGTGGATTCAAAAGATGACGCCTCAATTTGTGCACTTGAGGC
TGTTACTACATTATCTATTGGTGCTTTGTGCTTTTGCTTGGATAGTAAATGTAATTTGAATTTGAATGGGGTGGCTGCGATAAAAAAACTTTTCGGTATC
CATGATGATATGGATGGGACAGATAGCTCTCCAGAGAACATAGGTTTCATACTGGAAATGATTACACTTCTGAGTTCAAAGTTAAATGATGATGACTATC
TAGCTACAGATATGAGGGAAAGTTTGTATCAAGCTTCAGATTCTGCGAAGTCATTGCTGCTATTGTTGCAGAAGCCCACTGGTTCAGTTACAATAGATGA
CATTATGAGCAGTGAAGGCATTCAATCTCTACCATCAAATGAGCTTCTGGTTCATTCAAGAATAAATCAGATGGCAGATGGCACTGCTGAAAAGTTTGAT
GGTTACCTTTACCTAGGAGGCCTTGGAGATAAATTTCTGTGGGAATGTCCAGAAACCTTACCTGATAGATTGTCGCAGAATCCTTCTATGAAAAGGAAAT
TAGCATCATTGGATGGTTCTGGTAAGCGTGTCAAGGGAGAAACTTCAGTGGCTGAAGCTACTGTACAAAATGCATTTTCACGGGGAATGGGTTCATCAAC
TGCTCCTTCTGGTCCGACTCGCAGGGACACCTTTAGGCAGCGCAAGCCAAATACCAGTAGGCCTCCATCCATGCACGTTGACGACTATGTTGCTAGAGAA
AGAAGTGTTGATGGTGTTAGCAATTCAAATGTTATAGCAGTCCAGCGAGTGGGATCTACAGGTGGAAGACCTCCGTCAATTCATGTGGATGAATTTATGG
CCAGGCAAAGAGAACGCCAGAATCCAATGGTAGCTGTTGTTGGGGAACCTTCAGCAAAGGTTAAGAATGCAACCCCTGCGAATGATGTAGACAAGGAAAA
AGACAACAAGTCTAAGCAATTGAAAACTGTTCTGGATGATGATCTTCAGGGAATAGATATTGTTTTTGATGGTGAGGAGTCTGAATCTGATGACAAATTG
CCTTTTCCTCAGCCTGATGATAATCTGGAGCAGCTAGCCCCTGTCATAGGCGACCAAAGTTCTCCCCACTCGATTGTTGAAGAGACTGAGAGTGATGTTA
ATGGAAACAATCAATTTTCTCATTCACACACACCATTAGCATCTCATGTTGATGAGAACACCCAAAGTGAATTCTCTTCAAGGATGTCTGTTTCACGACC
TGAAATGCCGTTGACACGAGAACCAAGTGTTTCTTCGGATAAGAAGTTTTTTGAGCAACCTGATGATGCAAAGAATACCATTAAGACTTCTGCTGGCTTT
GATTCTATTTCAGCTGCAAGTACTTCAGGTTTTCCTCATCAGATTCCAGTTGATTCAAGGATGCCTCCACAAAATTTCTATATGAAGAACAGCTTGCAGC
ATTCTTCAGGATCTCGAGGACTTTATGACTCTAAAATTCCTCTGAATCAGCCACCTTTGCCTCCAATGCCACCTCCAGCAATGTCATCCATGATACCTCA
GAATCATGATCCAGGCCCAACTCAGTCATCCCCCTATGTTAACTCTGGAACAGAGGTGCAGCCTCCACTTCCTGCAGCCTTCCAAGTTCAGTCAGATTAT
CTATCTGCATTTGGTAGCAATCCTTCCATTCAGATGCCAGATTCCAAGTATTCACGTGCTTCTATCTCTTCTCCCAGTGGATCTGCCGGGCCTCACCCAC
CACTTCCTCCTACACCACCTCCCTTCTCATCTAGTCCATATAATTTACCTTCTTTAAATCCTTCTACTTCTCAATCTTCAGTATATACTGTTGGAACAAA
TGAGCTTCCCCAGACCTCTACTTCACCACCAATTGATCCGAGATTAGGAAATCTCTCTGTGTCGGGTGCTGGTTTAACTTCTTATATGCCACCTCCTCTA
ATGCCACCCATGGTTTTCAGTAGGCCAGCTACAATCCCTGTGACTCCTTATGGAAGCATCCCCACCCAGCAGCAAGGCGAGAGTCCAAATGTCTTGCAAA
ATCTCTCCATTCCTCAGCCTTCTGTTCAGTCAATACATCAGCTGCAGCCTCTTCAGCCTCCGCTTCGACGTCCACCACAGCCACCTCAACATCTTTGGTC
ACTTGCACAATCTTCTCAGCAGCTGGAACAAGGGGGGTCTTTGCAAAGTTCTATTCAAATGCAAGGCCATCAATTACAAATGCTGCAACAACAACAACTC
CCTTCTGTGCATGCTCATTATCAAGCTCAGCAGCAAGAGCTTTCTCAGTCAAGGCAGCAGCTAGTTGAGCATGCACAGCCACATGTTATACATCAGCAGG
GAGATGTTTCATCCCAGCAGCAACAAGATCTTGGAATGTCGTTACAGGAGTACTTCAAGGATCCAAAAGCTATCACGTCTTTATTGAGTAACAAGGAAGA
GCTGTGCCGGCTTTTGGAGCAAAATCCAAAATTAATGCAGATGCTTCAGGAAAGACTAGGCCAGCAATAG
|
|||||||||||||||
|
AA sequence
|
>Potri.016G069100.2 pacid=42809430 polypeptide=Potri.016G069100.2.p locus=Potri.016G069100 ID=Potri.016G069100.2.v4.1 annot-version=v4.1
MGRPEPSVLFSQTFVHPQLDEYVDEVLFAEPIVITACEFLEQNASSASQAVSVLGATSPPSFALEVFVKCEGETRFRRLCQPFLYSHSSSHVLEVEVAVV
TNHLVVRGSYRSLSLVIYGNTAEDLGQFSIEFDDSSLTNLVSSAEGKLEDLPLALHSTNRTVEDSLSSLNVLSLPVAASHISAEVKQFLQLILKLLELPN
LSDSVHRVLTTVVKAVCSFVTRDLCCETVNQKHIKMCGSKNIEEFHHVINEARNELLQVLGQVLGDESAELLADCTFLESEADLATSKQLVDMLSQYFSF
ERNSTNVGACQLSQNKSVILGLSLALLLCSGRESCFHFVSSGGMEQLAHIFSNEVQNSSAIILLSLGVVEQATRHPIGCEGFLGWWPREDENIPSGTSKG
YSQLLKLVLQRPQHDVASLATYVLHRLRFYEVVSRYEFSVLSALGGLSALGRVTSVTSAMLNSAKSQLKMLLKLINLRGPIEDPSIAASASRSLIIGQTE
GLLSYKATSNLVGSSHCCFSNWDIDSHLLALLKERGFLPLSAALLSSPILRSEAVDAMDTFVDIASTIGAILLSLLMCRSGLIFLLNYPELCTTLIDALR
GVGGMNREECVPLRYASVLLSKGFVCSPHEVGVIVETHLRVVNAIDRLLISTPHPEEFLWVLWELCGLSRSDCGRQALLVLGYFPEAISILIEALHSVKE
SEPVASGASPINLAIFHSAAEIFEVIVTDSTASSLDSWIGHAMELHKALHSSSPGSNRKDTPTRLLEWFDAGVVYHKNGAIGLLRYSAVLASGGDAHLTS
TSILVADLTDVEQVVGDALGGSDINVMDNLGKLISDKSFEDNPLRDSSITQMTTAIRILAFVSENSTVAAALYDEGALIVIYAILIKCSLMLERSSNSYD
YLVDEGTERNSTSDLLLERNREQSLVDLLVPTLVLLINLLQKLQEAKEQHRNTKLMNALLRLHREVSPKLAASAADLSSPYPDSALGFGAVCHLVVSALT
CWPLYGWTPGLFHSLLANVQATSLLALGPKETCSLLCLLNDLFPEEGVWLWKNGMPMLSALRKLAVGTLLGPQKEKQVDWYLETSHREKLLNQLTPHLDK
IAQIIEHYAISALVVIQDMLRVFIIRIACQKIEYASLLLQPILCCIRNHLSDLTSPSEIDAYKVYRYLDFLASILEHPCAKELLLEEGIAEMLTQVLERC
LVAIGSDGKQISDSKISAKSGFTLISWCCPVFKSFSLLCVPRTPLPYPVRHDLHSSASLSAKDCSLILPYLLKSCQVLPVGKELLSCLAFFKDLGSCNEG
QSACVTTLHHINTSIEEHESGKGQERNGNYNLDDIEWRKHPPLLSCWIRLLESVDSKDDASICALEAVTTLSIGALCFCLDSKCNLNLNGVAAIKKLFGI
HDDMDGTDSSPENIGFILEMITLLSSKLNDDDYLATDMRESLYQASDSAKSLLLLLQKPTGSVTIDDIMSSEGIQSLPSNELLVHSRINQMADGTAEKFD
GYLYLGGLGDKFLWECPETLPDRLSQNPSMKRKLASLDGSGKRVKGETSVAEATVQNAFSRGMGSSTAPSGPTRRDTFRQRKPNTSRPPSMHVDDYVARE
RSVDGVSNSNVIAVQRVGSTGGRPPSIHVDEFMARQRERQNPMVAVVGEPSAKVKNATPANDVDKEKDNKSKQLKTVLDDDLQGIDIVFDGEESESDDKL
PFPQPDDNLEQLAPVIGDQSSPHSIVEETESDVNGNNQFSHSHTPLASHVDENTQSEFSSRMSVSRPEMPLTREPSVSSDKKFFEQPDDAKNTIKTSAGF
DSISAASTSGFPHQIPVDSRMPPQNFYMKNSLQHSSGSRGLYDSKIPLNQPPLPPMPPPAMSSMIPQNHDPGPTQSSPYVNSGTEVQPPLPAAFQVQSDY
LSAFGSNPSIQMPDSKYSRASISSPSGSAGPHPPLPPTPPPFSSSPYNLPSLNPSTSQSSVYTVGTNELPQTSTSPPIDPRLGNLSVSGAGLTSYMPPPL
MPPMVFSRPATIPVTPYGSIPTQQQGESPNVLQNLSIPQPSVQSIHQLQPLQPPLRRPPQPPQHLWSLAQSSQQLEQGGSLQSSIQMQGHQLQMLQQQQL
PSVHAHYQAQQQELSQSRQQLVEHAQPHVIHQQGDVSSQQQQDLGMSLQEYFKDPKAITSLLSNKEELCRLLEQNPKLMQMLQERLGQQ
|
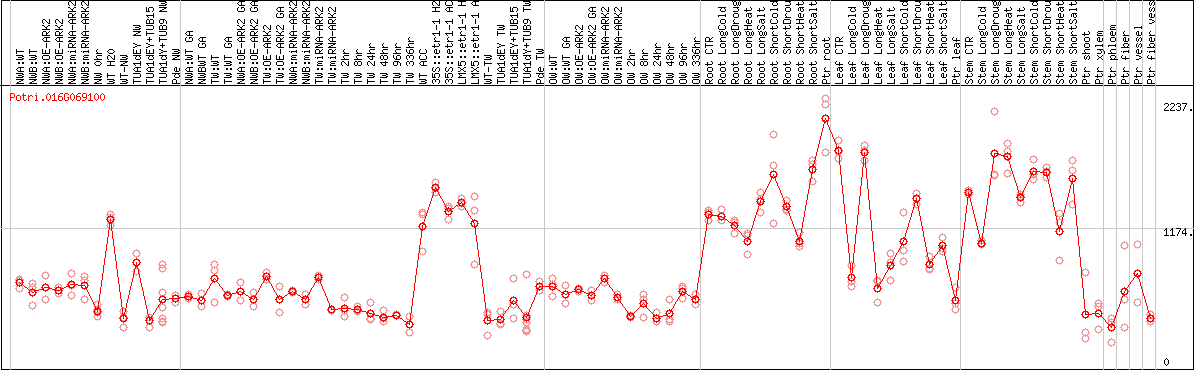
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G069100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
