External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G04820 153 / 2e-45
OFP
ATOFP13, OFP13
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
AT2G36050 109 / 1e-28
OFP
ATOFP15, OFP15
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 15, ovate family protein 15 (.1)
AT3G52540 99 / 2e-24
OFP
ATOFP18, OFP18
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 18, ovate family protein 18 (.1)
AT1G79960 66 / 2e-12
OFP
ATOFP14, OFP14
ovate family protein 14 (.1)
AT4G14860 64 / 3e-12
OFP
ATOFP11
ovate family protein 11 (.1)
AT1G05420 62 / 2e-11
OFP
ATOFP12, OFP12
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
AT3G52525 57 / 4e-10
OFP
ATOFP6, OFP6
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
AT5G22240 58 / 5e-10
OFP
ATOFP10, OFP10
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 10, Ovate family protein (.1)
AT2G18500 59 / 7e-10
OFP
ATOFP7, OFP7
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 7, ovate family protein 7 (.1)
AT5G19650 54 / 2e-08
OFP
ATOFP8, OFP8
ovate family protein 8 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021303
153 / 2e-44
AT5G04820 103 / 8e-26
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10016979
147 / 8e-43
AT5G04820 105 / 3e-27
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10043433
117 / 2e-31
AT5G04820 146 / 2e-42
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10034150
108 / 3e-28
AT5G04820 134 / 8e-38
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10043406
87 / 4e-21
AT5G04820 64 / 1e-12
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10010554
81 / 7e-18
AT1G05420 104 / 2e-27
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10006120
78 / 2e-16
AT1G05420 103 / 6e-26
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10006912
71 / 1e-14
AT1G05420 112 / 2e-30
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10014677
70 / 5e-14
AT1G05420 109 / 2e-29
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10035891
61 / 1e-10
AT1G79960 166 / 2e-48
ovate family protein 14 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04844
Ovate
Transcriptional repressor, ovate
Representative CDS sequence
>Potri.016G072900.1 pacid=42808992 polypeptide=Potri.016G072900.1.p locus=Potri.016G072900 ID=Potri.016G072900.1.v4.1 annot-version=v4.1
ATGAAGCTCCCTTTTCTCTCCAAGAATAATGCAAATACGGATCACTCATCAAGAGCTTTATGGCCTTGGCCTGCCTATTGTCAGCAACCAAGAACCCTCT
CTTTTAGTTTTAGAACAAGTGGTGGAATGCTCAAGACCATTAACCCAGGTTTCTTAGATGCTACTAACACTGATGTGGTGGACAGTACTTCACCTGAGTC
ATGGTTCACCAACTCCTGTGAGTCAGCTAGCTTCTCAACAGCATCAGATGATCAGTCAGGAGCTGGTGAGTCTATAGAGACAGTGATCAAAGGCTTGAGG
TCAGAGAGGCTGTTCTTTAAACCAGGGGAGACAAACTCAATCTTGGAAGAAGCTAAGGCAGGTGGTGAGTTTCCATTTAAAGAAAGTGTGGTATTGTCAA
TGGATTCTCGAGACCCTTATTTGGATTTCAAGAAGTCAATGGAGGAGATGGTTGAGGCTCATGGATTAACAGACTGGGAAGGTCTTGAAGAGCTTTTGTC
TTGCTACTTGAAGGTGAATGGAAAGAGCAATCATGGGTACATCATTGGTGCTTTTGTAGATTTGCTTGTTGGTCTTGCTATTGCTTCTTCTTCTTCTTCT
TCAAGTACTATCACTACTGCTAGTACTACGCAACATCATCATGATTCTCCTTCTTCTCCTTTGTCATTTTACACTTCTTCTGCATCTTCTGATGATTCGT
CTTCTACTCCTTGTTGTGTATCCTCGTTAGGAAATGAAGTTGATATAATCAGTCCTTGTTTAACTTCATTAGAAGCTGAAAATGAGATTAAAAAGTATTA
A
AA sequence
>Potri.016G072900.1 pacid=42808992 polypeptide=Potri.016G072900.1.p locus=Potri.016G072900 ID=Potri.016G072900.1.v4.1 annot-version=v4.1
MKLPFLSKNNANTDHSSRALWPWPAYCQQPRTLSFSFRTSGGMLKTINPGFLDATNTDVVDSTSPESWFTNSCESASFSTASDDQSGAGESIETVIKGLR
SERLFFKPGETNSILEEAKAGGEFPFKESVVLSMDSRDPYLDFKKSMEEMVEAHGLTDWEGLEELLSCYLKVNGKSNHGYIIGAFVDLLVGLAIASSSSS
SSTITTASTTQHHHDSPSSPLSFYTSSASSDDSSSTPCCVSSLGNEVDIISPCLTSLEAENEIKKY
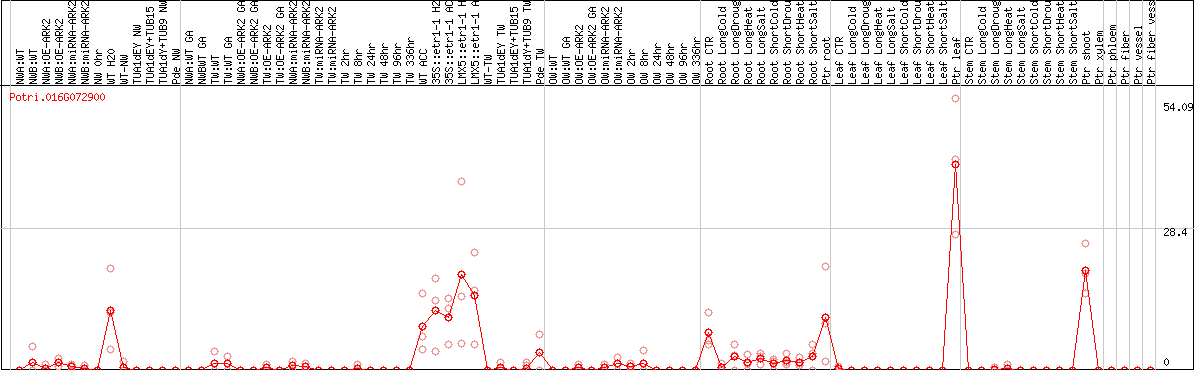
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G072900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.