External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G52560 281 / 5e-99
MMZ4 ,UEV1D ,UEV1D-4
MMS2 ZWEI HOMOLOGUE 4, ubiquitin E2 variant 1D-4 (.1.2.3.4)
AT2G36060 270 / 9e-95
MMZ3 ,UEV1C
UBIQUITIN E2 VARIANT 1C, MMS ZWEI homologue 3 (.1.2.3)
AT1G23260 219 / 1e-74
MMZ1 ,UEV1A
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
AT1G70660 214 / 8e-73
MMZ2 ,UEV1B
UBIQUITIN E2 VARIANT 1B, MMS ZWEI homologue 2 (.1.2)
AT1G70650 211 / 3e-66
Ran BP2/NZF zinc finger-like superfamily protein (.1.2)
AT5G25760 53 / 2e-09
PEX4, UBC21
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
AT2G02760 50 / 4e-08
ATUBC2
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
AT1G14400 49 / 5e-08
ATUBC1, UBC1
UBIQUITIN CONJUGATING ENZYME 1, ubiquitin carrier protein 1 (.1.2)
AT3G13550 49 / 1e-07
EMB144, COP10, CIN4, FUS9
FUSCA 9, EMBRYO DEFECTIVE 144, CONSTITUTIVE PHOTOMORPHOGENIC 10, CYTOKININ-INSENSITIVE 4, Ubiquitin-conjugating enzyme family protein (.1.2)
AT1G50490 48 / 2e-07
UBC20
ubiquitin-conjugating enzyme 20 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029415
281 / 5e-99
AT3G52560 290 / 7e-103
MMS2 ZWEI HOMOLOGUE 4, ubiquitin E2 variant 1D-4 (.1.2.3.4)
Lus10004211
263 / 1e-91
AT2G36060 273 / 8e-96
UBIQUITIN E2 VARIANT 1C, MMS ZWEI homologue 3 (.1.2.3)
Lus10021797
225 / 2e-76
AT1G23260 268 / 3e-93
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
Lus10034611
224 / 6e-76
AT1G23260 266 / 1e-92
UBIQUITIN E2 VARIANT 1A, MMS ZWEI homologue 1 (.1)
Lus10009495
52 / 2e-08
AT5G25760 273 / 1e-93
ubiquitin-conjugating enzyme 21, peroxin4 (.1.2)
Lus10036727
49 / 7e-08
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10037203
49 / 7e-08
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10016670
49 / 1e-07
AT5G56150 149 / 1e-46
ubiquitin-conjugating enzyme 30 (.1.2)
Lus10007126
49 / 1e-07
AT3G08690 148 / 4e-46
ubiquitin-conjugating enzyme 11 (.1.2)
Lus10013263
48 / 1e-07
AT2G02760 304 / 6e-108
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.016G073100.1 pacid=42809012 polypeptide=Potri.016G073100.1.p locus=Potri.016G073100 ID=Potri.016G073100.1.v4.1 annot-version=v4.1
ATGACGTTTGGCTCAGGAGGATCCAGTGTCGTGGTCCCTCGGAACTTCAGATTGCTGGAGGAACTTGAACGTGGAGAAAAAGGCATTGGGGATGGCACAG
TAAGCTATGGAATGGATGATGGAGATGACATTTACATGCGCTCTTGGACTGGCACTATAATTGGTCCTCACAATACTGTACATGAAGGTCGAATTTATCA
GTTAAAGCTCTTCTGTGATAAAGATTACCCGGAGAAACCACCAAGTGTTCGCTTTCATTCACGGATCAATATGACTTGTGTTAACCATGAAACCGGAGTG
GTGGAACCAAAAAAGTTCGGACTTCTTGCAAATTGGCAGCGAGAGTACACCATGGAGGACATACTGACACAGCTGAAGAAGGAGATGGCAGTACCACACA
ACCGTAAGCTGGTCCAGCCTCCTGAGGGTACCTGCTTCTAA
AA sequence
>Potri.016G073100.1 pacid=42809012 polypeptide=Potri.016G073100.1.p locus=Potri.016G073100 ID=Potri.016G073100.1.v4.1 annot-version=v4.1
MTFGSGGSSVVVPRNFRLLEELERGEKGIGDGTVSYGMDDGDDIYMRSWTGTIIGPHNTVHEGRIYQLKLFCDKDYPEKPPSVRFHSRINMTCVNHETGV
VEPKKFGLLANWQREYTMEDILTQLKKEMAVPHNRKLVQPPEGTCF
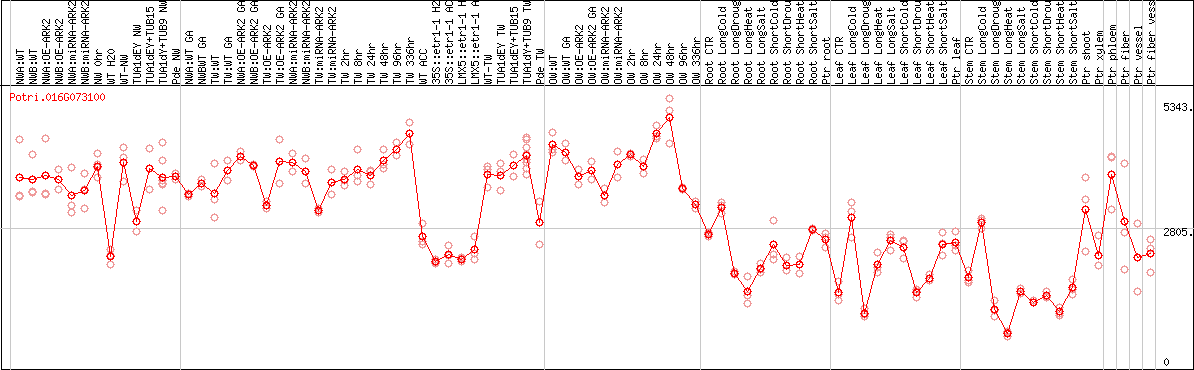
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G073100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.