Potri.016G090800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.016G090800.4 pacid=42809602 polypeptide=Potri.016G090800.4.p locus=Potri.016G090800 ID=Potri.016G090800.4.v4.1 annot-version=v4.1
ATGGACACTGAATTTGCAAATGCTAACCATCCACTGCATTTTCTTCATCGGCTGCTTCCTGCTGTTGGACCTGGACTTCTGATTGCAATTGGATATGTTG
ATCCTGGAAAGTGGGCAGCAACTGTTGAAGGAGGTGCTCGTTTTGGGTTTGATCTGGTACTACCGATGCTCCTTTTCAATTTTGTTGCCATTTTATGCCA
GTATCTCTCTGCTCGAATTGGTGTGATCACTAGAAAAGATCTTGCCCAGATTTGCAATGATGAGTATGACAAGTGGACATGCATGTTTCTAGGAGTTCAA
GCAGCACTTTCCGTGATTGCATTGGACCTCACAATGATTCTCGGCATTGCACATGGTCTTAATCTTCTCTTCGGGATGGACTTATCCACTTGTGTTTCTT
TAGCTGCTGCTGAGGCCATTTTATTTCCATTTTTTGCTACCCTCATGGAGAGATGCAAGGCAAGTTTCCTTTGCACATGCATTGCAGGCTTCATATTACT
ATTGTATTTCTTCGGGGTTCTCATCAGCCAACCAGGAATCCCTCTTTCCATAAACGGGACGCGAACAAAATTAAGCGAGGAGAGTGTGTTTGCCCTGATG
AGTCTACTTGGAGCAAGTATCATGCCTCACAATTTTTTCCTCCATTCTGCTATTGTGCTGCAACATCAGGGACCACCAAATATTTCCAGGGATGCCTTGT
GTCTTAACCATTTTTTTGCCATTTTATGCATCTTCAGTGGCATCTATCTGGTAAATTTTGTGCTTATGAACTCCGCTGCAAATGTGTTCCACAGCACTGG
CCTTGTTTTACTTACTTTTCCTGATGCGATGTCACTAATGGAACAGGTGTTCAGGAGTCCAGTAGCACCCTTTGGCTTTTCACTCATCTTGTTTTTTGCG
AATCAAATCACAGCATTTTCCTGGAATCTTGGTGGGCAAGTAGTTTTACACAATTTTCTCAGATTGGATATTCCTAATTGGCTTCAACGTGCGACATTCA
GAATTATTGCAGTTGTTCCAGCCCTTTATTGTGTGTGGACTTCAGGAGTTGAAGGAATATACCAGTTGCTTATACTTACACAGGTTATGGTTGCTTTACT
TCTTCCATCTTCCGTGATCCCTCTTTTCCATATTGCTTCATCAAGACAAGTGATGGGTGTCTACAAAATTTCTCCATTCTTGGAGTTCGTAGCGCTCATT
TCATTCATGGGGATGCTTGGCATAAAGATTATTTTTGTGGTGGAAATGGTATTTGGGGATAGTGATTGGGTAGGTACTTTGAGATGGAGTACAGTAAGTG
GTTCGTCTACCTCTTACATTGTTCTTCTCATTACTGCCTGTTCATCATTTTGTTTGATGCTCTGGCTAGCAGCTACCCCATTGAAATCTGCTACTCGCTT
AGATGCTCAGGTATGCAACTGGGATGTACAAAATGCTGTATCCGAACCATCCACGCTGATAGAGGAAGAATTTTTGACTGAAAATATATGTACTGGAGAG
GAGCTTATCGAGAGGCAGGAACAATTACCAGAACCAGGGAAATCTTTTGAGAGTTACTCAAATATAACTGTTGCAAATGCTGATCCTGATTTGCCTGAGA
CAATTATGGAGTCTGATCAGGAACTTCATTTGACTACTATCAAAGAAAAACATTCTGAAGTCGCATTTTCTAGTCCCCAAACATTCTATGAGGAAACATC
ACCTACAACAGAGTCAGCATCTCTGTCAGCTTCTGTCAACTTGGTTCCTGATGCTGAATTGCTTGTTGCCAAGAAGGCTAAGATTGAATCAATGGACCCA
GTTGAAAAGACTCTGGATATAGAGGGTGAGTTGCATACTGAAAAGGAGGATGATGAAGGGGATAACTGGGAGCCTGAAGATTCATCCAAAGGGGTTCCTG
GAAGCACTTTGTCTTTGACATCTGATGGTCCTGGATCATTTAGGAGTCTTAGTGGTAAAAGTGATGCAGGTGGGAATGGTGCTGGAAGTCTTTCAAGATT
AGCAGGGTTGGGGCGCGCTGCAAGACGTCAATTAGCTGCAGTTCTTGATGAGTTTTGGGGACAATTGTATGACTTCCATGGGCAAATAACTCAAGAAGCA
AAGACCAAGAAACTGGATGCCTTGGGGGTTGATTTGAAACTTGCTTCTTCCCAGCTGAAAGTGGACACTGCTGGGAAGGAATCAAGCGGATATTTCTCAT
TGGTAGGTGGCAGAGCATCTGATTCATTGATCAATTCAAGTTTATGCGACTCCCCCAAGCAATTGCGGGTGCAAAGCAATATTGACTCATCTTATGGAGT
TCAGAGGGGACCATCCTCATTGTGGTCCAACCACATGCAGTTGCTGGATGCATATGTACAAGGTCCCAGCCAGAGTATTGCTGACTCAAGTGAGAGACGG
TATTCTGGTGTACGCACGCCACCATCTTCTGATGGCTGGGATAATCAGCCAGCTACAGTTCATGGTTATCAGATTGCATCTATTGCCAATCGAATTGCCA
AGGACAGGGGTTTCAGTAGCTTAAATGGCCAAATGGAGTCACCAGCACCAATATCCCCATCCTTAGGACCTAGAAATTATAGGGACCCGCTCACTGTCTC
TATGGGGAAAAATTTGCAAAATGGACTAAGCTCTTCACAAGCGTCAGGATTTCAGAATCTTGCAGTAACTAGAAATAGCCCATTACAGTCTGAAAGACCT
TATCATGATGTATACTCTGGATCTGCTGATGATACAGGAATGTCAGCTAATACAAAGAAGTACCATAGCTTGCCTGACATTTCAGGGCTTGCTGGTCCCT
ATAGGGACCTGTATATGTCTGAAAAGAATGCTCAGTGGGACAAGTCTGCAGGATTTGGTTCCTCTGTTGGTAGATCAGCTTATGAACAATCTTATTATTC
AAATACCGGATCAGGAGCAGGAGGTCCTTTGTCTTTTAATGGACTATCAAAAGGGCATGGGGATGCTTTCTCGCTTCACATGACTCCAGACCCTGGATCC
CTCTGGTCCAAGCAACCATTTGAACAGTTTGGTGTTGCTGACAAAATTCGTGCTGTTGGGAGTGGATTAGGAAATCGGTCTAATTCAATTAATCGCGAAG
TAACTTCTCCTGTGGATTCAGAGGCCCAACTTCTTCGGTCTTTCAGGCATTGTATTGTGAAGCTTTTGAAATTGGAAGGATCTGACTGGCTGTTTAGGCA
AAACGATGGAGCTGATGAGGATCTAATTGATTGTGTGGCTGCAAGGGAGAGGTATTTGTATGAAGCTGAAACAAGAGAAATGAACCATGTAGATCACATG
GGTGGATCTACATATTTATATTCTGATAGGAAATCTGGTTCTGCTTTGAGGAATGATGATGCCAGTATTACCAACATTATGGTTTCCTCAGTTCCTCATT
GCGGGGAGGGCTGTGTTTGGAGATCGGATTTGATAATAAGCTTTGGGGTCTGGTGTATTCACCGGATTCTTGATTTGTCACTAATGGAAAGCCGTCCAGA
GTTGTGGGGGAAATACACTTATGTGTTGAATCGTCTTCAGGGCATCATAGAATTGGCATTTTCAAAGCCCCGTACTCCCATGTCCCCATGCTTCTGCCTT
CAAATCCCTGCATCTCACCAGCATAGATCAAGTCCACCTGCTTCGAATGGAATGTTGCCCCCTGCTTCAAAACCAGGTAGGGGGAAATGCACAACTGCAG
CGACGCTTCTGGACTTGATTAAGGATGTGGAGATTGCCATATCTTGCCGAAAGGGTCGATCGGGAACTGCTGCTGGTGATGTTGCTTTTCCGAAGGGAAA
AGAGAATCTGGCTTCTGTGCTTAAGCGCTACAAGCGCCGGTTATCCAACAAACTAATCGGTAGTAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.016G090800.4 pacid=42809602 polypeptide=Potri.016G090800.4.p locus=Potri.016G090800 ID=Potri.016G090800.4.v4.1 annot-version=v4.1
MDTEFANANHPLHFLHRLLPAVGPGLLIAIGYVDPGKWAATVEGGARFGFDLVLPMLLFNFVAILCQYLSARIGVITRKDLAQICNDEYDKWTCMFLGVQ
AALSVIALDLTMILGIAHGLNLLFGMDLSTCVSLAAAEAILFPFFATLMERCKASFLCTCIAGFILLLYFFGVLISQPGIPLSINGTRTKLSEESVFALM
SLLGASIMPHNFFLHSAIVLQHQGPPNISRDALCLNHFFAILCIFSGIYLVNFVLMNSAANVFHSTGLVLLTFPDAMSLMEQVFRSPVAPFGFSLILFFA
NQITAFSWNLGGQVVLHNFLRLDIPNWLQRATFRIIAVVPALYCVWTSGVEGIYQLLILTQVMVALLLPSSVIPLFHIASSRQVMGVYKISPFLEFVALI
SFMGMLGIKIIFVVEMVFGDSDWVGTLRWSTVSGSSTSYIVLLITACSSFCLMLWLAATPLKSATRLDAQVCNWDVQNAVSEPSTLIEEEFLTENICTGE
ELIERQEQLPEPGKSFESYSNITVANADPDLPETIMESDQELHLTTIKEKHSEVAFSSPQTFYEETSPTTESASLSASVNLVPDAELLVAKKAKIESMDP
VEKTLDIEGELHTEKEDDEGDNWEPEDSSKGVPGSTLSLTSDGPGSFRSLSGKSDAGGNGAGSLSRLAGLGRAARRQLAAVLDEFWGQLYDFHGQITQEA
KTKKLDALGVDLKLASSQLKVDTAGKESSGYFSLVGGRASDSLINSSLCDSPKQLRVQSNIDSSYGVQRGPSSLWSNHMQLLDAYVQGPSQSIADSSERR
YSGVRTPPSSDGWDNQPATVHGYQIASIANRIAKDRGFSSLNGQMESPAPISPSLGPRNYRDPLTVSMGKNLQNGLSSSQASGFQNLAVTRNSPLQSERP
YHDVYSGSADDTGMSANTKKYHSLPDISGLAGPYRDLYMSEKNAQWDKSAGFGSSVGRSAYEQSYYSNTGSGAGGPLSFNGLSKGHGDAFSLHMTPDPGS
LWSKQPFEQFGVADKIRAVGSGLGNRSNSINREVTSPVDSEAQLLRSFRHCIVKLLKLEGSDWLFRQNDGADEDLIDCVAARERYLYEAETREMNHVDHM
GGSTYLYSDRKSGSALRNDDASITNIMVSSVPHCGEGCVWRSDLIISFGVWCIHRILDLSLMESRPELWGKYTYVLNRLQGIIELAFSKPRTPMSPCFCL
QIPASHQHRSSPPASNGMLPPASKPGRGKCTTAATLLDLIKDVEIAISCRKGRSGTAAGDVAFPKGKENLASVLKRYKRRLSNKLIGSK
|
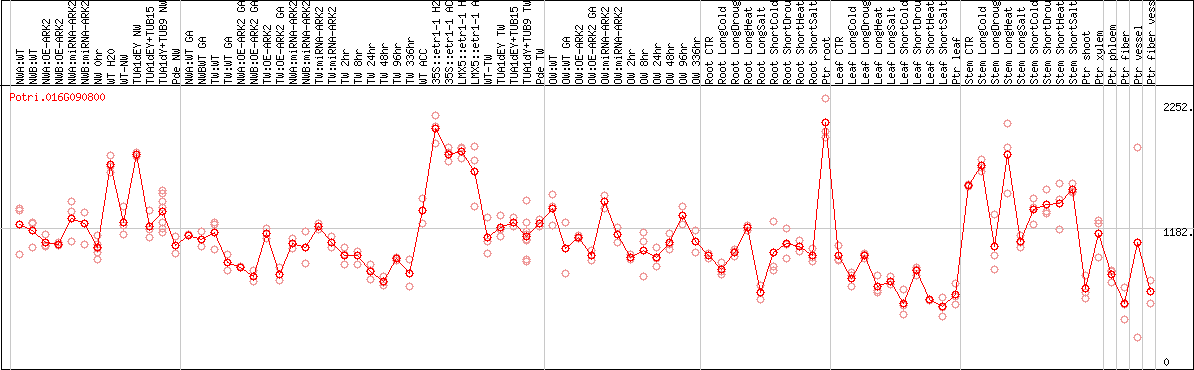
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G090800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
