Potri.016G113800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.016G113800.5 pacid=42808958 polypeptide=Potri.016G113800.5.p locus=Potri.016G113800 ID=Potri.016G113800.5.v4.1 annot-version=v4.1
ATGACCCAACCACCAAATATCGTTCGGAAACTACTGGTAGAAGTAGTCGACGCACGGGATCTTCTCCCCAAAGACGGTCAGGGAAGCTCCAGTGCCTGTG
TCATCGCCGACTTCGACGGTCAAAGAAAACGAACCACCACGAAATATCGTGACCTCAACCCCGTGTGGAAAGAGACGTTGGAGTTCATCGTTTCTGACCC
AAACAACATGGAGTTCGAAGAGCTTGAAGTCGAGGTTCTTAATGACAAGAAATTCGGCAATGGTAGCGGCCGTAAAAACCATTTCTTGGGGAGGGTTAAA
GTGTATGGCTCTCAGTTTTCTAAAAGAGGTGAAGAAGGGATTGTTTACTTCCCTCTTGAGAAGAAAAGTGTGTTCAGCTGTATAAGGGGAGAAATCGGTC
TGAGAATTTGTTTTTACGATGAGTTAGTTGAAGAGGATCAGCAACAAGCTCCAGCTCCATCGGAGGAGGATGCAGATACGTTGCAAGATCAAAAGCCGCT
AAAATCCCCGGCAGTTATTGAGGAGGAAGGCAGAGTTTTTGAAGTTCTCGCACGACCGGAGATTAACTGCCACGACTATCACCACCCCCACCACCACCAT
TTTCATCATAATGGGACTCACTCGCCTCCTTTTGTGGTTATCGAGGAGTCTCCACCGCCGGTGGTGCAGGTAAACTCGGAACCATCACTAGGTTCGCAAC
AAGTGCCTCTTCCAGAGGAGCCGCATTACGTAGAAACCCACACTCAGTACCATCCAGAAGTGAGGAGGATGCAGACTACGAGAGTCGCGTCCAGTGGTGA
TAACAGAGTGAAGACATTGAGGCCGCCAATCGGAGATTTCTCTCCTAAAGTAATTTCTGGCAGATTCAAGTCGGAATCGACGGAGAGGATCCATCCTTAC
GATCTTGTCGAACCAATGCAGTATTTGTTCATCAGCATTGTCAAGGCTCGTGGTTTATCGCAAAACGAGAGTCCTATCGTCAAGTTAAGGACCTCAACTC
ATTGTGTGAGGTCTAAACCAGCGAGTTACCGACCTGGTGCCTCACCGGATTCGCCTGAGTGGCACCAGGTTTTTGCATTGGGTCATAATAACAAGACTGA
CGGACAGCTCCCCAATGCAGCTGGCAACATAGAGATCTCAGTTTGGGACGCACGTTCGGAGCAGTTTCTCGGCGGTGTTTGCTTTGATATCTCTGAAGTT
CCGGTGAGGGATCCGCCGGACAGTCCGCTTGCTCCTCAGTGGTACCGATTGGAAAGTGACGCTGCCGCTGGTCAGATATGTAATAGGGTTTCTGGTGATA
TCCAGCTCTCTGTATGGATTGGGACTCAAGCTGATGATGCATTTGCTGAAGCGTGGAGCTCAGACGCCCCTTACGTCTCTCACACGCGCTCAAAGGTTTA
CCAGTCCCCTAAGCTGTGGTATTTGAGAGTGACGGTTATTGAAGCTCAAGATCTTCACCTTTCGTCCAATCTGCCACCTTTGACAGTTCCGGATATCAGA
ATCAAAGCTCAATTGGGGTTCCAATCGGCGAGGACGAGGCGAGGGTCCATGAGCAATCATAGTACGTCGTTCCGATGGATCGATGACCTAATTTTCGTCG
CCGGCGAGCCGCTGGAAGAGTCATTGATTCTGCTAGTGGAGGACCGTACCACCAAGGAAGCGGTACTCCTCGGGCACATCATAATTCCCGTGAGCTCAAT
CGAGCAACGGTATGATGAGCGCCACGTGGCATCCAAATGGTTCGCGTTGGAAGGAGGCGGCGGGGACACTGGGGGAGCAGGCTGTGCTACTGGCGGGTCC
TACCGTGGGAGAATCCATTTGCGTCTCTGTTTGGAAGGAGGGTATCACGTGCTTGATGAAGCGGCGCACGTGTGCAGTGATTTTAGACCCACGGCTAAGC
AGCTATGGAAGCCAGCTATTGGGGTTTTGGAGCTGGGGATTTTAGGGGCTCGTGGGTTGCTGCCCATGAAAACCAAAGGAGGGGGCAAAGGTTCGACCGA
TGCTTATTGTGTGGCTAAGTATGGCAAAAAATGGGTCCGAACAAGAACTATCACGGATAGCTTTGAGCCACGTTGGAACGAGAAGTATACCTGGCAGGTC
TATGATCCCAGCACTGTTCTCACCATTGGGGTTTTCGACAATTGGCACATGTTTGGTGAAATGTCTGATGATAAACCTGATTGCCGTATCGGGAAAATAC
GTATACGAGTATCTACATTGGAGAGCAACAAGGTGTACATGAATTCGTATCCCTTGTTGGTTTTATTAAGAACCGGGTTGAAGAAAATGGGTGAGATCGA
ATTAGCGGTTCGGTTTGCTTGCCCGAGTTTATTACCGGATACCTGTGCAGTTTATGGGCAGCCATTGCTTCCTAAAATGCACTATCTTCGGCCGCTCGGG
GTAGCCCAACAAGAGGCATTGCGTGGTGCAGCCACCAAAATGGTTTCTTTGTGGCTTGCGCGGTCCGAGCCACCATTGGGACCAGAGGTGGTTAGGTACA
TGCTGGATGCGGATTCTCATGCATGGAGCATGAGAAAGAGTAAAGCCAATTGGTTTCGAATCGTAGCGGTTCTTGCATGGGCTGTCGGGTTGGCTAAATG
GTTGGATGATATTCGAAGATGGAGAAACTCGGTCACAACTGTATTGGTTCATATTTTGTATTTGGTGCTTGTTTGGTATCCGGAATTGGTAGTCCCAACA
GGGTTCTTATATGTGTTCTTGATTGGTGTTTGGTACTATAGGTTCAGGCCTAAGATACCGGCAGGCATGGATATCCGGTTATCACAAGCAGAAACCGTTG
ACTCGGATGAACTAGATGAGGAATTCGACACGGTACCAAGCATGAGACCACCCGAAATTATAAGGGCCCGTTATGACAGGTTAAGGATGTTGGCAGCCCG
AGTCCAAACAGTTTTGGGCGATTTTGCAACCCAAGGGGAGAGGGTTCAAGCTTTGGTGAGTTGGAGAGATCCGAGAGCCACAAAGTTGTTCATAGCGGTA
TGTTTGGCTATAACATTGATACTGTATGTGGTTCCACCAAAAATGGTAGCTGTGGCCTTAGGATTCTATTTTTTACGACACCCCATGTTTAGAGACCCAA
TGCCTCCGGCTAGCTTGAACTTTTTCCGCCGGCTTCCGAGCTTATCAGATCGGTTAATGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.016G113800.5 pacid=42808958 polypeptide=Potri.016G113800.5.p locus=Potri.016G113800 ID=Potri.016G113800.5.v4.1 annot-version=v4.1
MTQPPNIVRKLLVEVVDARDLLPKDGQGSSSACVIADFDGQRKRTTTKYRDLNPVWKETLEFIVSDPNNMEFEELEVEVLNDKKFGNGSGRKNHFLGRVK
VYGSQFSKRGEEGIVYFPLEKKSVFSCIRGEIGLRICFYDELVEEDQQQAPAPSEEDADTLQDQKPLKSPAVIEEEGRVFEVLARPEINCHDYHHPHHHH
FHHNGTHSPPFVVIEESPPPVVQVNSEPSLGSQQVPLPEEPHYVETHTQYHPEVRRMQTTRVASSGDNRVKTLRPPIGDFSPKVISGRFKSESTERIHPY
DLVEPMQYLFISIVKARGLSQNESPIVKLRTSTHCVRSKPASYRPGASPDSPEWHQVFALGHNNKTDGQLPNAAGNIEISVWDARSEQFLGGVCFDISEV
PVRDPPDSPLAPQWYRLESDAAAGQICNRVSGDIQLSVWIGTQADDAFAEAWSSDAPYVSHTRSKVYQSPKLWYLRVTVIEAQDLHLSSNLPPLTVPDIR
IKAQLGFQSARTRRGSMSNHSTSFRWIDDLIFVAGEPLEESLILLVEDRTTKEAVLLGHIIIPVSSIEQRYDERHVASKWFALEGGGGDTGGAGCATGGS
YRGRIHLRLCLEGGYHVLDEAAHVCSDFRPTAKQLWKPAIGVLELGILGARGLLPMKTKGGGKGSTDAYCVAKYGKKWVRTRTITDSFEPRWNEKYTWQV
YDPSTVLTIGVFDNWHMFGEMSDDKPDCRIGKIRIRVSTLESNKVYMNSYPLLVLLRTGLKKMGEIELAVRFACPSLLPDTCAVYGQPLLPKMHYLRPLG
VAQQEALRGAATKMVSLWLARSEPPLGPEVVRYMLDADSHAWSMRKSKANWFRIVAVLAWAVGLAKWLDDIRRWRNSVTTVLVHILYLVLVWYPELVVPT
GFLYVFLIGVWYYRFRPKIPAGMDIRLSQAETVDSDELDEEFDTVPSMRPPEIIRARYDRLRMLAARVQTVLGDFATQGERVQALVSWRDPRATKLFIAV
CLAITLILYVVPPKMVAVALGFYFLRHPMFRDPMPPASLNFFRRLPSLSDRLM
|
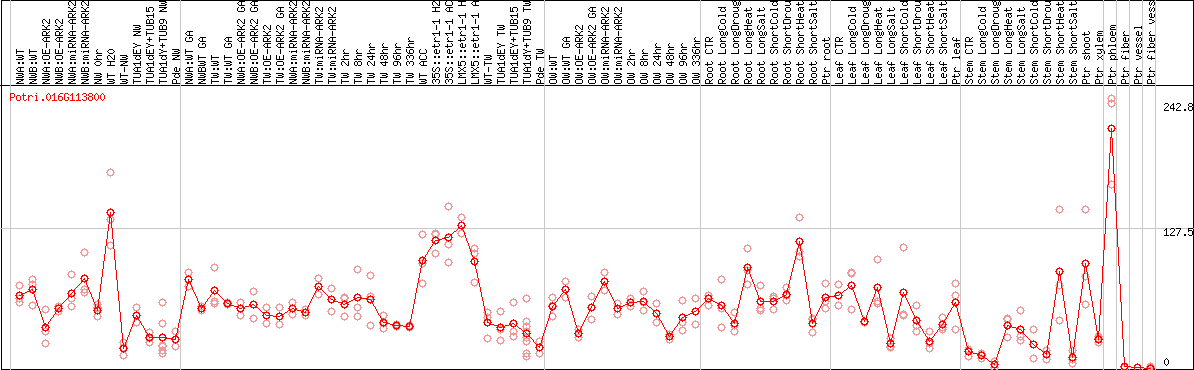
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G113800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
