Potri.016G129000 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.016G129000.2 pacid=42809885 polypeptide=Potri.016G129000.2.p locus=Potri.016G129000 ID=Potri.016G129000.2.v4.1 annot-version=v4.1
ATGCCGTTGACGAGATACCAGATAAGGAACGAGTATAGTTTAGCCGATCCAGAGCTGTACAAGGCTGCTGATAAAGATGATCCAGAAGCGCTTTTGGAAG
GTGTCGCCATGGCTGGCCTCGTCGGCGTCTTGCGCCAGCTCGGTGACCTCGCCGAGTTCGCTGCTGAGATATTTCATGGACTGCATGAAGAAGTTATGAC
AACTGCAGCAAGAGGCCATGGGCTGATGGCTCGGGTACAACAGCTCGAGGCAGAATTTCCTTCAATCGAGAAGGCATTTCTCTCCCAAACTAATCATTCA
CCATTCTTTTCCAGCTCAGGTGTTGACTGTCATCCTAATTTGCAAATGGAGCAGAATCTAATTGCTCGTGGAGACTTACCTCGCTTTGTAATGGACTCTT
ATGAAGAATGCCGTGGCCCACCACAGTTATTCCTCTTGGACAAGTTTGATGTTGCTGGGGCTGGGGCATGTTTGATGCGTTATACTGATCCATCATTCTT
TAAAGTTGAAACAGCATCTTCTGGAATAGCAACTGTAGAGGTTCAGAGGGAGAAGAAAATTCGAAAAAAGAAGAAAGGATCACGGTATAGAAATGGAGAA
ACTCCTGAAGTTGTACAAACATCACACACCAAATTGCATGAGTTGTTGTTACAGGAACATTTTGAGAATGGTCATTCTGACCCTGCACGGCTAGTGAAAT
TGAAGAGAAGGCAGATAAATGGATCTCCATTTGATCTGAAACCTGGGAAAAGTTACATGGAGAAATTTGTGCTCACTCCTTCACCAGAGCGTAAACAAGT
TTGTGAAGATTCTGTCACTCGATCACCTTTGAAATTTACGTTGGATAATTCTAGTGAATCAGGCTATGAAATACATGAGGTCAGTGTGGTAAGTCCTGCA
AAAAAGTCATTAAATGGAGTCGAAAGCACATCTTCATCACCCAGTGAACAAGAGGCCATGCTTAAACCTGTTAAGGATGAGTTGGATGGGGAGGCTGTTG
ATCGTGGAATTATTAAGGTACTTGACCCAATTGTTGATCGCGGGATGGATGAATTGCCTCCAACCGTCTATAAGATGGCTATTGAAGAGGAATTATTAGT
TGATGCAGATATAAAAAGAGAGGGCACTGTAGATGGGGATCATTCTGATGATATGGCAAGTGAAGTGGACAACTACATGGATGCCCTTACTACCATGGAT
TCTGAAATGGAAACAGACAATGAGTATAAAGCTAAAAATGCTCCGGACTTCATCAATCTGAGGATACAAGGTGCAGATTCTGATGCAAATGAGGAGCAGC
TGGGTTTTCAAGCTAATTCTTCAGATTCTCAATCTATTGGAAACTCTTCTCTGTCTGAGGGTGGGAACAGTTTGTTTAAGAAAGGAACTTCTAGCTCTTC
TTACTCCGAGACTCTCTACAATTTGGTTGAGAATACAGCATCTGATGGTGAAGGATCAGGTAAATGGTTCCCTTCTGCCACTTCTACTGAAAACCATGCA
ACAAATGTTACTGATCTGCCATCAGATCATCCACCTGTATATGCGGAGACTGGCATAACCGAATCTCATCACCTTGTAACATTCAATGATACAAGGGAAG
ACAAAATCCCTGATCCTGTTGAAGCTTCTTGTAGTTCTTGTCCTACAGATTCGAATCCTGTCTTCCTGCATTCAGTTCCTGTTGCACGTTCCATGGTAAG
CCCCTTGTCAGGACCTGAATTAGTCGAGGCATCCTCTGGCAGTACTGAACTTGGCTCCAAATCACCGCACTGTGAAAGAAATGGGTTATACCCGACTGAT
TCTTTCATTGCTTTAACTGACATCCCTTCACAGATGGGGCATGATGCTTCACTTCCAGATTCTTCTAAAAGTCATTCTGTGGATGTGTTAGATCATGAAG
ATCCAGACATGTTAACTGATGCTGTTGTGCATGTGTCGAACATGTCAGATCTGGCTTCTGAAAAGAAAGTCAGTGATGATTCTGTGAATGAAGTGCTTCT
AACAGATTGTGCAGCTGAACATTCAACTTTGACACCAGCAGAGGAGCAATTTCCTCACTCAGCTTTGCCTGTAGTGGAGTTGGATGCTGGTGTCCCATCT
CTGCCTGATAATTCAAATGTTGTGAAGCCTGATGGTTTGGTTTCTAAAGCAGATGATGAAATTTTAACGAGGGAAGGGAGCACAGAAATTTCAACACCAG
TGGTGGATACTTCAGAGTCTGAGTGCATTAACGAGCATCAATTCTCTGATGTAACAGTAGATGCTTCACAAGAGGAACTTGACTCAACAAAATTAAGACT
CCCATGTTCTGAAGAGAATGTGAAACTTGAAGAAATTTCTGAGGGTCCAGATGCTGAGGAAAAGAATGCTTCCACGAAGAAAGTGGATATTACAAGAGGA
GATGCTACTTATTTCGAACATGAATCATGTAGTTCAGATAAACCAACTCCTGAAGATCATGTAAATTTGGCTGATGATGTAACTGAAACTGTTAAAGCAG
AAGATATGGCTGTATCAACTGCTGCTACAAGTGGTGTAGATGCTGAGGAAAAGAATGCCTTCACGAAGAAAGTGGATATTACAAGAGGAGATGCTACTTA
TTTCGAACATGAATCATGTAGTTCAGATAAATCAACTCCTGAAGATCATGTAAATTTGGCTGATGATGTAACTGAAACTGTTCAAGCAGAAGAGATGGCT
GTATCAACTGCTGCTACAAGTGGTGTAGATGCTGAGGAAAAGAATGCATTCACGAAGAAAGTGGATATTACAAGAGGAGATGCTACTTCTTTCGAACATG
AATCATGTAGTTCAGATAAACCAACTCCTGAAGATCATGTAAATTTGGCTGATGATGTAACTGAAACTGTTAAAGCAGAAGATATGGCTGTATCAACTGC
TGCTACAAGTGGTGTAGATGCCGAGGAAAAGAATGCCTTCACGAAGAAAGTGGATATTACAAGAGGAGATGCTACTTCTTTCGAAGATGAATCATGTTCA
GATAAACCAACTCCTGAAGATCATGTAAATTTGGCTGATGCTGTAACTGAAACTGTTAAAGCAGAAGATATGGCTGTATCAACTGCTGCTACAAGTGGTG
TAGATGCCGAGGAAAAGAATGCATTCACGAAGAAAGTGGATATTACAAGAGGAGATGCTACTTCTTTCGAAGATGAATCATGTTCAGATAAACCAACTCC
TGAAGATCATGTAAATTTGGCTGATGCTGTAACTGAAACTGTTAAAGCAGAAGATATGGCTGTATCAACTGCTGCTACAAGTGGTGTAGATGCCGAGGAA
AAGAATGCATTCACAAAGAAAGTGGATAGTACAAGAGGAGATGCTGCTTCTTTCGAAGATGAATCATGTTCAGATAAACCAGCTCCTGAAGATCATGTAA
ATTTGGCTGATGATGTAACTGAAACTGTTAAAGCAGAAGATATGGCTGTATCAATTGCTGCTACAAGTGGTGTGAACAATGAAGATGTCAGCAATGTGAT
CTGTCCGTCTTCAGAGTTAGTTTGTTCACCACCTAGGAATTCTACGGAGATGGTAGAATCTCTTTCAATCTCCGAGGATCCAAATCAGACTACTTTGAAC
CTTGATGAGGTAACTTCTGCCAAATGTCTGTCAGAATCACAGGTAAAGATGGAAGTGACTTCTACAGATTGGGATTCTAATTCATACAAACCAGTTTCTG
AAGACTATCGAAATCAAGAGGTAATTGAAGTTCATAACCCTTCTTCGGAAGTGAGCAACCAGGAGTCTGAATCTAAAGATAACCATCAGAGTCATTGTGG
GGAGGTTGGTGACAATACTGTTTGTTCGCCTGTCTGTTATCCACCAGAGTCAGGAAATGGTTTAGAACAGTCAATTGAGGTGCAAGCTGATCAAATTAGT
TCAGAGTCCATGCATGCAGACGATGCAAGTTCTTTGTTGAGTAGTCAAACTTCATCTGCTGGCTACCTTTTAGGGCCAGGAATTCCTCTGGATCACACAT
CAGAATTGCAATCTGATCAACTAGACAGGAGATGCTTGAAGTCAGGCGAAGCAAGTTCTAGATCAGCAGATGTTAAGTCAGAACAGATACAGAATCTGCA
TAACATAACCGAGGAAAGATGCCCTGACCCTTCCAGTTTGAAAGATCTCTCCAGTCAAGAATTTCTACTGCAATCAGCCTGTCAGGGACATAATGTTACT
GATCAAGCAACAAATCCATTTGATTCTGCTTTTCCCAGCTTTGGTGTACTCCCTGTTCCTGAGACAAGCCAGGTCAATCCAGAGGCAATGCCACCATTGC
CACCTCTACCTCCTATGCAGTGGAGGTTGGGGAAAATTCAACCTGGTCCACTAGATGCAGACAGAGACATGATGGATCATAGTCAACGCACTTCTCAACC
AATTGAAACATTTATTCTTGATCAGAAGGTCCAGTTTGATTTCCCAGCATTAGATAGAGAGATTGTGCATCCTTCAAACCCATTTTTGTCTCTCCCTGTT
GAAGACAGTCAGAGGTCTCAACACTTGACAACAGAGTTAATGGGAAATTCCTTGCTGCCAACTCGACTCTTGTCAGAAATGCCAACCATTGACAATGATG
CACAATATCAACAGGATGACCTCCTGTCAGACAGAACACAATCTGTGAACTCATCTTTGGCTTTATCAGAAATGCCTGATGAGAGGCATGAGCATGGTTT
CCTTCAACTAGGAGGAGAATCCACGCAATTCAGTTCCAATCCATTCTCACTAGAGCTGGGCATAAACGATACAGCTGCTTTAAATGATCCAATGCTCACA
CAGGGACTACCAATCCGTCTGTTTAACCAATCAGCACCCGAAACAGGATTGGAGGTGAAATTTCCCGGACAGAGTTCACAAAATGCAGAAGGGGAACAAG
GGAATTCTTCTGGTAAATCTGCAGTACCACTAAATACAGAAGAAGAGCAGCATCACCATGATTTTGTAACTTCACATGGACTGCCAATATGGCCGCCAAC
TACATTAGGTATGACACCACCAACATATGAAGTTGGAAAGACAAATGGAAAGAAGATTCCTCGCCCTCGAAATCCTCTCATTGATGCTGTTGCTGCACTT
GATAAAAGCAAGTTGAGAAAGGTGGCTGAACGAGTTCGACCGCAGCTGGGACCCAAGGTAGAAGAAAGAGATTCACTGCTGGAACAGATACGAACCAAGT
CCTTTAACTTGAAGCCTGCAACCGCGACAAGGCCTAGCATGCAGGGTGTTCAGGGTCCAAAAACCAACTTGAAGGTTGCTGCCATCTTAGAAAAAGCAAA
TGCAATTCGCCAGGCATTGACTGGAAGCGACGAAGATGACGATTCAGACAGCTGGAGTGATTCTTAA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.016G129000.2 pacid=42809885 polypeptide=Potri.016G129000.2.p locus=Potri.016G129000 ID=Potri.016G129000.2.v4.1 annot-version=v4.1
MPLTRYQIRNEYSLADPELYKAADKDDPEALLEGVAMAGLVGVLRQLGDLAEFAAEIFHGLHEEVMTTAARGHGLMARVQQLEAEFPSIEKAFLSQTNHS
PFFSSSGVDCHPNLQMEQNLIARGDLPRFVMDSYEECRGPPQLFLLDKFDVAGAGACLMRYTDPSFFKVETASSGIATVEVQREKKIRKKKKGSRYRNGE
TPEVVQTSHTKLHELLLQEHFENGHSDPARLVKLKRRQINGSPFDLKPGKSYMEKFVLTPSPERKQVCEDSVTRSPLKFTLDNSSESGYEIHEVSVVSPA
KKSLNGVESTSSSPSEQEAMLKPVKDELDGEAVDRGIIKVLDPIVDRGMDELPPTVYKMAIEEELLVDADIKREGTVDGDHSDDMASEVDNYMDALTTMD
SEMETDNEYKAKNAPDFINLRIQGADSDANEEQLGFQANSSDSQSIGNSSLSEGGNSLFKKGTSSSSYSETLYNLVENTASDGEGSGKWFPSATSTENHA
TNVTDLPSDHPPVYAETGITESHHLVTFNDTREDKIPDPVEASCSSCPTDSNPVFLHSVPVARSMVSPLSGPELVEASSGSTELGSKSPHCERNGLYPTD
SFIALTDIPSQMGHDASLPDSSKSHSVDVLDHEDPDMLTDAVVHVSNMSDLASEKKVSDDSVNEVLLTDCAAEHSTLTPAEEQFPHSALPVVELDAGVPS
LPDNSNVVKPDGLVSKADDEILTREGSTEISTPVVDTSESECINEHQFSDVTVDASQEELDSTKLRLPCSEENVKLEEISEGPDAEEKNASTKKVDITRG
DATYFEHESCSSDKPTPEDHVNLADDVTETVKAEDMAVSTAATSGVDAEEKNAFTKKVDITRGDATYFEHESCSSDKSTPEDHVNLADDVTETVQAEEMA
VSTAATSGVDAEEKNAFTKKVDITRGDATSFEHESCSSDKPTPEDHVNLADDVTETVKAEDMAVSTAATSGVDAEEKNAFTKKVDITRGDATSFEDESCS
DKPTPEDHVNLADAVTETVKAEDMAVSTAATSGVDAEEKNAFTKKVDITRGDATSFEDESCSDKPTPEDHVNLADAVTETVKAEDMAVSTAATSGVDAEE
KNAFTKKVDSTRGDAASFEDESCSDKPAPEDHVNLADDVTETVKAEDMAVSIAATSGVNNEDVSNVICPSSELVCSPPRNSTEMVESLSISEDPNQTTLN
LDEVTSAKCLSESQVKMEVTSTDWDSNSYKPVSEDYRNQEVIEVHNPSSEVSNQESESKDNHQSHCGEVGDNTVCSPVCYPPESGNGLEQSIEVQADQIS
SESMHADDASSLLSSQTSSAGYLLGPGIPLDHTSELQSDQLDRRCLKSGEASSRSADVKSEQIQNLHNITEERCPDPSSLKDLSSQEFLLQSACQGHNVT
DQATNPFDSAFPSFGVLPVPETSQVNPEAMPPLPPLPPMQWRLGKIQPGPLDADRDMMDHSQRTSQPIETFILDQKVQFDFPALDREIVHPSNPFLSLPV
EDSQRSQHLTTELMGNSLLPTRLLSEMPTIDNDAQYQQDDLLSDRTQSVNSSLALSEMPDERHEHGFLQLGGESTQFSSNPFSLELGINDTAALNDPMLT
QGLPIRLFNQSAPETGLEVKFPGQSSQNAEGEQGNSSGKSAVPLNTEEEQHHHDFVTSHGLPIWPPTTLGMTPPTYEVGKTNGKKIPRPRNPLIDAVAAL
DKSKLRKVAERVRPQLGPKVEERDSLLEQIRTKSFNLKPATATRPSMQGVQGPKTNLKVAAILEKANAIRQALTGSDEDDDSDSWSDS
|
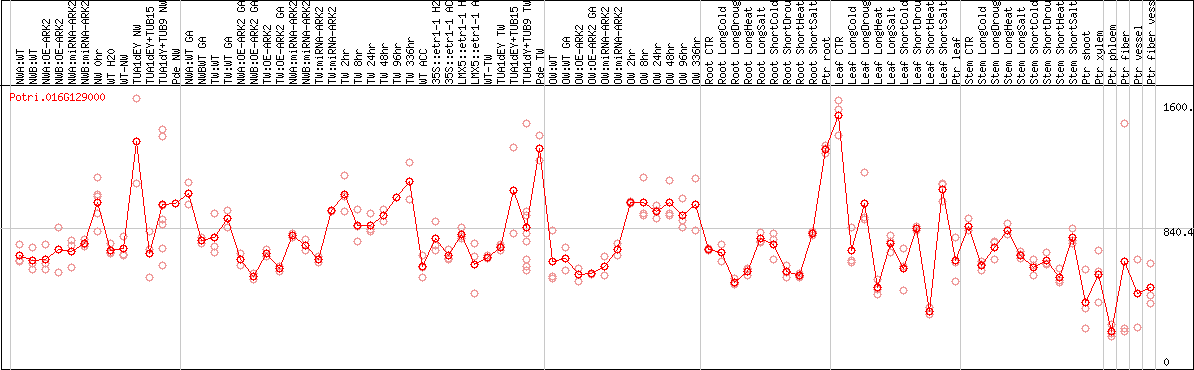
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G129000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
