External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G01750 128 / 4e-37
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
AT2G14560 115 / 2e-32
LURP1
LATE UPREGULATED IN RESPONSE TO HYALOPERONOSPORA PARASITICA, Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
AT3G11740 114 / 8e-32
Protein of unknown function (DUF567) (.1)
AT1G33840 103 / 3e-27
Protein of unknown function (DUF567) (.1)
AT3G16900 92 / 2e-23
Protein of unknown function (DUF567) (.1)
AT3G56180 85 / 2e-20
Protein of unknown function (DUF567) (.1)
AT3G10986 69 / 1e-14
Protein of unknown function (DUF567) (.1)
AT3G15810 66 / 3e-13
Protein of unknown function (DUF567) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G131400
315 / 1e-110
AT5G01750 147 / 3e-44
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G131300
313 / 6e-110
AT5G01750 143 / 7e-43
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G131100
303 / 3e-106
AT5G01750 154 / 3e-47
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G130800
302 / 9e-106
AT5G01750 162 / 4e-50
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G130500
279 / 2e-96
AT5G01750 218 / 6e-72
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G131850
276 / 3e-95
AT5G01750 213 / 3e-70
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G130700
273 / 8e-94
AT5G01750 209 / 2e-68
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G131500
253 / 4e-86
AT5G01750 139 / 2e-41
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Potri.016G131800
241 / 2e-81
AT5G01750 213 / 3e-70
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014021
186 / 1e-59
AT5G01750 202 / 6e-66
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10002477
185 / 3e-59
AT5G01750 218 / 5e-72
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10001255
185 / 3e-59
AT5G01750 218 / 5e-72
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10025253
171 / 9e-54
AT5G01750 169 / 4e-53
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10022754
159 / 6e-49
AT5G01750 214 / 3e-70
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10014154
151 / 7e-46
AT5G01750 206 / 2e-67
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10009092
148 / 1e-44
AT5G01750 161 / 7e-50
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10025254
147 / 1e-44
AT5G01750 166 / 1e-51
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10019896
142 / 2e-42
AT5G01750 169 / 3e-53
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
Lus10009094
135 / 6e-40
AT5G01750 150 / 1e-45
Protein of unknown function (DUF567) (.1), Protein of unknown function (DUF567) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0395
Tubby_C
PF04525
LOR
LURP-one-related
Representative CDS sequence
>Potri.016G131600.1 pacid=42810341 polypeptide=Potri.016G131600.1.p locus=Potri.016G131600 ID=Potri.016G131600.1.v4.1 annot-version=v4.1
ATGGCTACTGGGCAAGCACCACGCAACCCCATTCCAGCTATGAGAACATACCCACCGGTGGAGCATCCAGTGGTGGTAATAGGGCCACAGTACCTGGCAC
AGTACCCTGTTGAGCTCGCCGTCAACGATTTTAAGGTGTCTGACATTAATGGCACCCTCGTCTTCAAGGTCAAGTTTAAACTATCAAGCATTAATCGTCT
TTTTCTGAATGATGCAGCCGGAAACACCCTTGTCAATCTCAGGAAGAAGACAATGACCATGCATGGGAGGTGGGAGGCTTTTAGAGGAGAAAGCAAGGAG
GAGAATGATTTGCTTTTCACAGCCAAGAAATCCAAGACTGAGGTAGACGTATTCTTGGGTAATAACAAAGGAGAGGTCCCTGATTTCAAGGTCAAAGAAG
GCTACAGCAAGAGTTCCCGCTCTATACTTCTTCGAGATTCCAATACCATGATTGCACAGGTGCATGGACGAGACACTCTCGCCATTATGCCTAATGTTGA
TTATGCCTTCATAGTGGCTCTTCTGGTGGTGATTCTGGAGGGGATCAATCCGGATGATCATAGGGAAGGCGCGGCCCTTAAAGGCATTAACGGTGTTATC
GCGTAA
AA sequence
>Potri.016G131600.1 pacid=42810341 polypeptide=Potri.016G131600.1.p locus=Potri.016G131600 ID=Potri.016G131600.1.v4.1 annot-version=v4.1
MATGQAPRNPIPAMRTYPPVEHPVVVIGPQYLAQYPVELAVNDFKVSDINGTLVFKVKFKLSSINRLFLNDAAGNTLVNLRKKTMTMHGRWEAFRGESKE
ENDLLFTAKKSKTEVDVFLGNNKGEVPDFKVKEGYSKSSRSILLRDSNTMIAQVHGRDTLAIMPNVDYAFIVALLVVILEGINPDDHREGAALKGINGVI
A
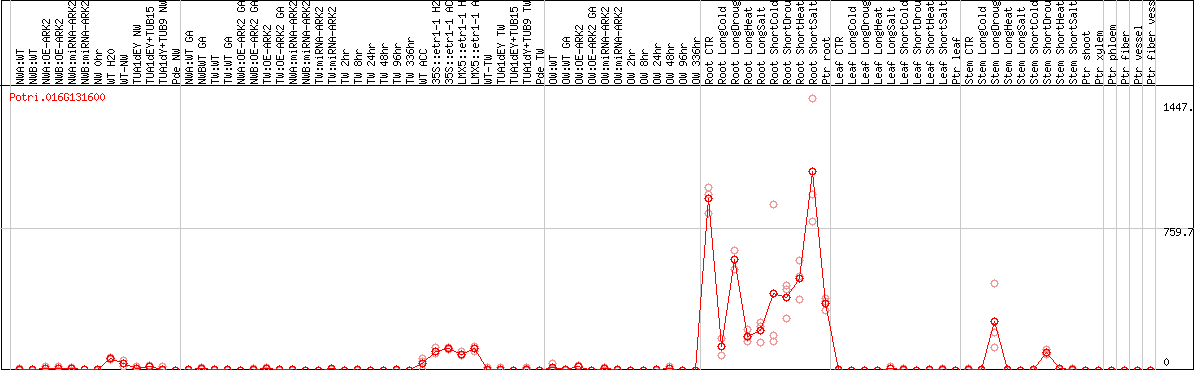
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G131600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.