External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40435 155 / 2e-49
unknown protein
AT3G56220 132 / 4e-40
bHLH
transcription regulators (.1)
AT1G29270 56 / 3e-10
unknown protein
AT1G12860 57 / 5e-10
bHLH
SCRM2, ICE2, bHLH033
SCREAM 2, INDUCER OF CBF EXPRESSION 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G26744 54 / 3e-09
bHLH
SCRM, ATICE1, ICE1, bHLH116
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
AT5G10570 50 / 6e-08
bHLH
bHLH061
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G65640 44 / 9e-06
bHLH
bHLH093
beta HLH protein 93 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G106900
259 / 1e-90
AT2G40435 150 / 4e-47
unknown protein
Potri.013G154100
141 / 7e-44
AT2G40435 126 / 9e-38
unknown protein
Potri.019G126800
140 / 2e-43
AT2G40435 144 / 1e-44
unknown protein
Potri.004G060800
78 / 5e-19
AT1G29270 69 / 4e-15
unknown protein
Potri.011G070000
70 / 1e-15
AT1G29270 75 / 2e-17
unknown protein
Potri.015G105200
52 / 3e-08
AT3G26744 323 / 1e-104
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
Potri.012G106000
50 / 1e-07
AT3G26744 339 / 9e-111
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
Potri.002G108400
43 / 3e-05
AT5G65640 233 / 2e-74
beta HLH protein 93 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025257
145 / 5e-45
AT2G40435 134 / 2e-40
unknown protein
Lus10017415
143 / 2e-44
AT2G40435 146 / 3e-45
unknown protein
Lus10010217
142 / 6e-44
AT2G40435 152 / 1e-47
unknown protein
Lus10009089
136 / 2e-41
AT2G40435 129 / 2e-38
unknown protein
Lus10029251
87 / 3e-22
AT2G40435 91 / 5e-24
unknown protein
Lus10007304
86 / 6e-22
AT1G29270 97 / 4e-26
unknown protein
Lus10043199
52 / 3e-08
AT3G26744 387 / 1e-129
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
Lus10032542
52 / 3e-08
AT3G26744 386 / 6e-129
SCREAM, A. THALIANA INDUCER OF CBP EXPRESSION 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.4)
Lus10039631
45 / 4e-06
AT5G65640 230 / 3e-74
beta HLH protein 93 (.1.2)
Lus10002160
45 / 7e-06
AT5G65640 233 / 5e-75
beta HLH protein 93 (.1.2)
PFAM info
Representative CDS sequence
>Potri.016G132600.1 pacid=42810393 polypeptide=Potri.016G132600.1.p locus=Potri.016G132600 ID=Potri.016G132600.1.v4.1 annot-version=v4.1
ATGTCTTCCAGGAAGAAAAAGAAGGCAGCTCTTTACGAGAAGTTACGTGCTGCTACCAATTCTAATGCTATGAACAAAACCTCAATCATAGTAGACGCGT
CAAAATATATTGGAGAGTTGAAGAACAAGGTGGATAGGCTAAAAAAAGAAATCGGAACATCTTCAACCCCCCAAAATTCATTACCTGCGCAGGTCACAGT
GGAAAACCTAGAAAAGGGTTTCCTTATTAATGTATTTTCAGGAAAGAATTGCCCTGGATTACTTGTCTCCATACTTGAAGCCTTCGAGGAACTAGGCCTT
GATGTGCTTGATGCTAGGGTTTCTTGTGAAGACAATTTCCAACTTGAAGCGATTGGTGGAGACCAAAACCAAGGCCATGATGCTCAAGTAGTGAAACAGG
CAGTGCTGCAAGCTATCCATAACTGGAATGAAGGCAGCTAG
AA sequence
>Potri.016G132600.1 pacid=42810393 polypeptide=Potri.016G132600.1.p locus=Potri.016G132600 ID=Potri.016G132600.1.v4.1 annot-version=v4.1
MSSRKKKKAALYEKLRAATNSNAMNKTSIIVDASKYIGELKNKVDRLKKEIGTSSTPQNSLPAQVTVENLEKGFLINVFSGKNCPGLLVSILEAFEELGL
DVLDARVSCEDNFQLEAIGGDQNQGHDAQVVKQAVLQAIHNWNEGS
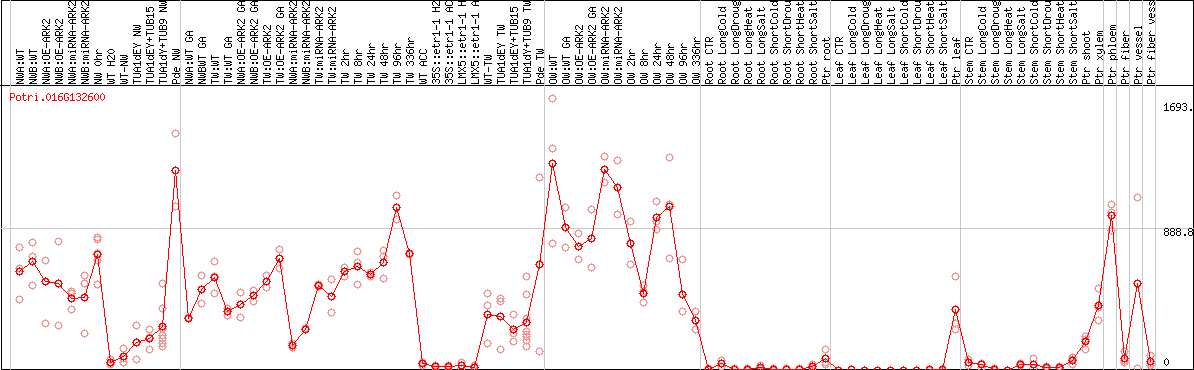
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.016G132600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.