External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G10860 340 / 2e-120
CBSX3
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
AT1G47271 186 / 8e-60
Cystathionine beta-synthase (CBS) family protein (.1)
AT3G52950 57 / 2e-09
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1), CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.2)
AT2G36500 55 / 7e-09
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
AT5G63490 55 / 8e-09
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
AT5G50530 52 / 6e-08
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
AT5G50640 52 / 6e-08
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
AT1G69800 40 / 0.0008
Cystathionine beta-synthase (CBS) protein (.1), Cystathionine beta-synthase (CBS) protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G161500
376 / 1e-134
AT5G10860 343 / 8e-122
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
Potri.005G228300
227 / 4e-76
AT1G47271 251 / 1e-85
Cystathionine beta-synthase (CBS) family protein (.1)
Potri.002G034800
97 / 5e-26
AT1G47271 102 / 4e-28
Cystathionine beta-synthase (CBS) family protein (.1)
Potri.005G070500
56 / 3e-09
AT3G52950 750 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1), CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.2)
Potri.007G098400
50 / 2e-07
AT3G52950 728 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1), CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.2)
Potri.012G099600
49 / 7e-07
AT5G50530 639 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
Potri.015G098100
49 / 1e-06
AT5G50530 628 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
Potri.001G303900
44 / 3e-05
AT4G34120 267 / 1e-90
LOSS OF THE TIMING OF ET AND JA BIOSYNTHESIS 1, CYSTATHIONE [BETA]-SYNTHASE DOMAIN-CONTAINING PROTEIN 1, CBS domain containing protein 2, Cystathionine beta-synthase (CBS) family protein (.1)
Potri.009G099200
41 / 0.0002
AT4G34120 266 / 1e-90
LOSS OF THE TIMING OF ET AND JA BIOSYNTHESIS 1, CYSTATHIONE [BETA]-SYNTHASE DOMAIN-CONTAINING PROTEIN 1, CBS domain containing protein 2, Cystathionine beta-synthase (CBS) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019117
335 / 2e-118
AT5G10860 347 / 2e-123
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
Lus10034441
338 / 7e-116
AT5G10860 350 / 4e-120
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
Lus10035014
317 / 2e-111
AT5G10860 341 / 5e-121
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
Lus10021675
317 / 3e-111
AT5G10860 338 / 7e-120
CBS domain containing protein 3, Cystathionine beta-synthase (CBS) family protein (.1)
Lus10001964
187 / 9e-60
AT1G47271 202 / 5e-66
Cystathionine beta-synthase (CBS) family protein (.1)
Lus10035965
50 / 4e-07
AT3G52950 714 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1), CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.2)
Lus10043123
49 / 7e-07
AT5G50530 714 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
Lus10025698
49 / 7e-07
AT3G52950 728 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1), CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.2)
Lus10032625
49 / 7e-07
AT5G50530 721 / 0.0
CBS / octicosapeptide/Phox/Bemp1 (PB1) domains-containing protein (.1)
Lus10002834
44 / 2e-05
AT4G34120 260 / 2e-87
LOSS OF THE TIMING OF ET AND JA BIOSYNTHESIS 1, CYSTATHIONE [BETA]-SYNTHASE DOMAIN-CONTAINING PROTEIN 1, CBS domain containing protein 2, Cystathionine beta-synthase (CBS) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00571
CBS
CBS domain
Representative CDS sequence
>Potri.017G000100.1 pacid=42812892 polypeptide=Potri.017G000100.1.p locus=Potri.017G000100 ID=Potri.017G000100.1.v4.1 annot-version=v4.1
ATGCAAGGCGCACTTCGCACGTTTTTAACCCATGGAAATGTTGTCAAGGATTCACTCTTCCAACATCTCCGTGTTTCGAGTCCAGTCTTGCGGCCCGTTG
TGTTTTCTCGTTTCGAATCAGTCTCCTCTGCTCCTATTGAGGAGACTGGCTTCGAGAGCACCACAATTGCTGACATCCTCAAAGAAAAAGGTAAAAATGC
GGATGGTTCATGGCTTTGGTGCACTACGGATGACACTGTTTATGATGCTGTCAAGTCGATGACACATCACAACGTTGGGGCCTTGGTGGTTGTCAAACAT
GGTGAGCAGGAATCAATTGCAGGAATCATCACAGAGAGGGACTATCTGCGGAAGATAATTGTGCAGGGAAGATCGTCCAAGTCAACCAAGGTTGGGGATA
TCATGACTGAAGAGAACAAGCTTATTACGGTCACGCATGATACCAAAGTTCTGAAGGCAATGCAATTAATGACAGATAGACGCATTAGGCACATTCCTGT
GATTGATGATAAGGGAATGATTGGCATGGTATCTATTGGAGATGTTGTTCGCGCCGTGGTGAGTGAGTACAGGGAGGAGCTGGACCGTCTGAATGCATAC
ATACAAGGAGGTTATTGA
AA sequence
>Potri.017G000100.1 pacid=42812892 polypeptide=Potri.017G000100.1.p locus=Potri.017G000100 ID=Potri.017G000100.1.v4.1 annot-version=v4.1
MQGALRTFLTHGNVVKDSLFQHLRVSSPVLRPVVFSRFESVSSAPIEETGFESTTIADILKEKGKNADGSWLWCTTDDTVYDAVKSMTHHNVGALVVVKH
GEQESIAGIITERDYLRKIIVQGRSSKSTKVGDIMTEENKLITVTHDTKVLKAMQLMTDRRIRHIPVIDDKGMIGMVSIGDVVRAVVSEYREELDRLNAY
IQGGY
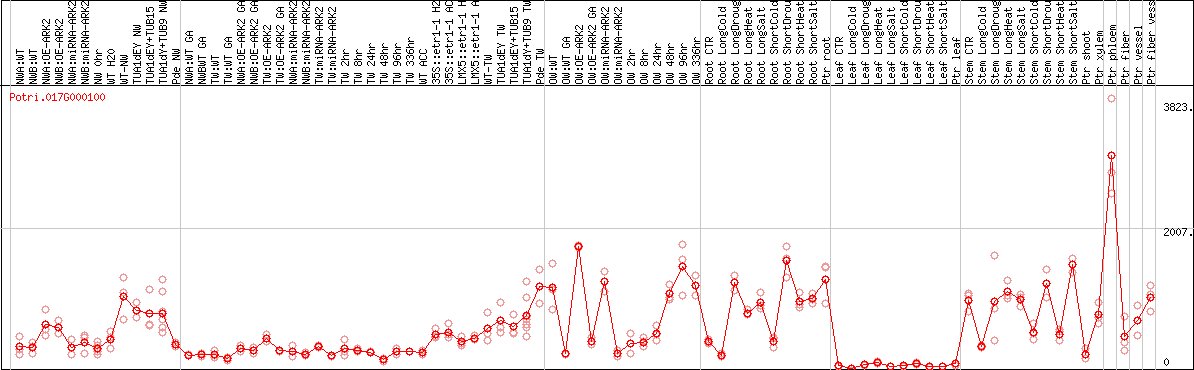
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G000100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.