External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G43820 144 / 1e-40
SGT1, ATSAGT1, GT, UGT74F2
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
AT1G05675 141 / 1e-39
UDP-Glycosyltransferase superfamily protein (.1)
AT1G05680 139 / 1e-38
UGT74E2
Uridine diphosphate glycosyltransferase 74E2 (.1)
AT1G24100 138 / 1e-38
UGT74B1
UDP-glucosyl transferase 74B1 (.1)
AT2G43840 129 / 3e-35
UGT74F1
UDP-glycosyltransferase 74 F1 (.1.2)
AT2G31790 114 / 1e-29
UDP-Glycosyltransferase superfamily protein (.1)
AT2G31750 107 / 4e-27
UGT74D1
UDP-glucosyl transferase 74D1 (.1)
AT4G15480 104 / 7e-26
UGT84A1
UDP-Glycosyltransferase superfamily protein (.1)
AT4G15490 102 / 3e-25
UGT84A3
UDP-Glycosyltransferase superfamily protein (.1)
AT3G21560 98 / 1e-23
UGT84A2
UDP-Glycosyltransferase superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024117
166 / 9e-49
AT1G05675 385 / 4e-130
UDP-Glycosyltransferase superfamily protein (.1)
Lus10009412
160 / 9e-47
AT2G43820 474 / 1e-165
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
Lus10008742
155 / 9e-45
AT2G43840 478 / 6e-167
UDP-glycosyltransferase 74 F1 (.1.2)
Lus10020559
154 / 2e-44
AT2G43820 436 / 3e-150
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
Lus10020556
154 / 2e-44
AT2G43820 472 / 1e-164
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
Lus10006353
154 / 3e-44
AT2G43820 398 / 1e-135
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
Lus10006352
151 / 3e-43
AT2G43820 414 / 6e-142
UDP-glucose:salicylic acid glucosyltransferase 1, Arabidopsis thaliana salicylic acid glucosyltransferase 1, UDP-glucosyltransferase 74F2 (.1)
Lus10039041
145 / 6e-41
AT1G05675 343 / 3e-114
UDP-Glycosyltransferase superfamily protein (.1)
Lus10024118
145 / 1e-40
AT1G05675 375 / 2e-126
UDP-Glycosyltransferase superfamily protein (.1)
Lus10024486
144 / 2e-40
AT1G05680 389 / 4e-132
Uridine diphosphate glycosyltransferase 74E2 (.1)
PFAM info
Representative CDS sequence
>Potri.017G032766.1 pacid=42813393 polypeptide=Potri.017G032766.1.p locus=Potri.017G032766 ID=Potri.017G032766.1.v4.1 annot-version=v4.1
ATGGAGAAGCAAGAAAGAATCTGTCATGTAGTGGTCATACCTTATCCTGCTCAAGGTCACATTAACCCCATGATCCAGTTCTCCAAGCGCCTAGCCTCCA
AAGGACTCCAAGTAACACTGGTCATCTTCTCAAGCCAGACGCTCTCGACGCCTGCCTCGCTTGGCTCAGTCAAAGTGGTGACCATTTCTGATAGTTCTGA
TACAGGCAGCAGTTCCATAGGTGACCTTCTAAAACAATTCCAAGCTACAGTGGCACCAAAACTGCCACAGCTCGTTGTAGAACTAGGTATCTCTTCTGGG
CATCCAGTAAGTTGCCTAGTGTATGACTCATTCATGCCTTGGGTTCTAGAGATAGCTAGACAGCTTGGCCTCATTGGGGCTTCCTTCTTCACACAGTCAT
GTGCTGTTAACTCTGTCTACTACCAAATCCATGAGGGGCAGCTAAAGATTCCACTAGAGAAGTTTCCTGTATCAGTTCCAGGGCTGCCACCTCTAGACGT
TGATGAACTGCCATCATTCGTGCACGACATGGAATCAGAATACTCATCTATACTTACCCTAGTAGTCAATCAGTTTTTGAATTTCAGGGGACCTGATTGG
GTCTTTGTGAACAGTTTCAACTCCCTGGAGGAGGAGGTACCCTTGAAGAAGTTACATAAAGAAAAGTAG
AA sequence
>Potri.017G032766.1 pacid=42813393 polypeptide=Potri.017G032766.1.p locus=Potri.017G032766 ID=Potri.017G032766.1.v4.1 annot-version=v4.1
MEKQERICHVVVIPYPAQGHINPMIQFSKRLASKGLQVTLVIFSSQTLSTPASLGSVKVVTISDSSDTGSSSIGDLLKQFQATVAPKLPQLVVELGISSG
HPVSCLVYDSFMPWVLEIARQLGLIGASFFTQSCAVNSVYYQIHEGQLKIPLEKFPVSVPGLPPLDVDELPSFVHDMESEYSSILTLVVNQFLNFRGPDW
VFVNSFNSLEEEVPLKKLHKEK
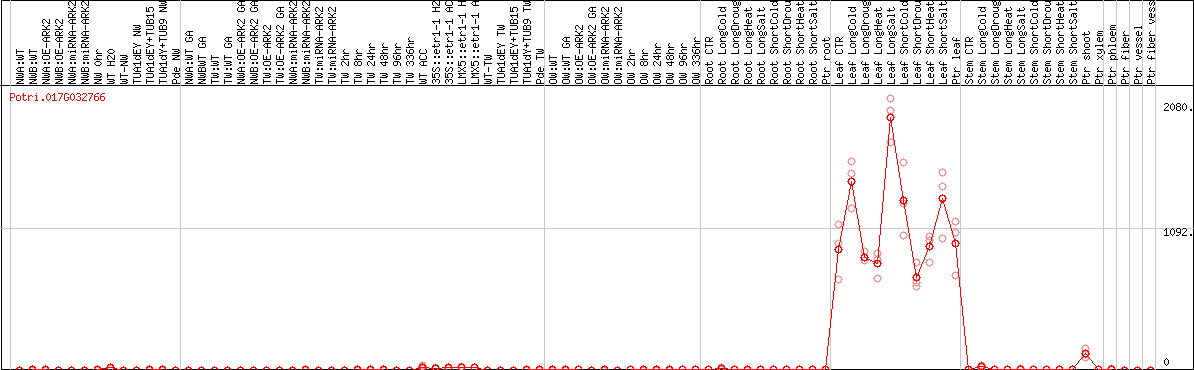
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G032766 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.