Potri.017G044000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G044000.1 pacid=42813127 polypeptide=Potri.017G044000.1.p locus=Potri.017G044000 ID=Potri.017G044000.1.v4.1 annot-version=v4.1
ATGACTCGTTCGACTCGGAAGTTTAGGGTGTTCGAGTTCGACGAAGAAGAAGAAGATAGAGTCGAGAAGGAGTCGGCGAAGTTCGTCGGCAAGTTTCGAA
TTCAAAAGAGAAAACGAAACGACAACAAGAAGGACGACGACGCCTCTTCTCCTCTCACTAAGTATAATTTCCTCCAGTGCTTTGGAGGATGTACTGGGAC
TGTGAAAATAGAAAGAAGCAATGAACCTATTGATATAGATGATGGACCAATTGATGTTGACACTGGAGGAGAGATGGCTACTTTACGCAAGGGAAAGAGC
GATGAAGTAGTATATATTGATACAACCATTATAGACAATCAATGCCAATATTCTGTTTCAGTTTCTGCTCGCATGCCACAAGAAGATTGTGCTGACAAAG
AGGAGATTTCTCAGATGGAAACCCTTATTACAGATGATGGGCCAATTGATGTTGACGCTGGAGTTGCAGGAGAGATCAATACTTTATGCAAGGGAAAGAG
TGATGAAGTAGTTGATATTGACACAACCATTTTGGACGATCAGTGTCAATGTTCTGTTTCAGTTCCTGCTCAAATGCCACAAGAAGATTGTGCTGTCAAG
GAGGAGATTTCTCAGCTGGATACCCTTGGGATTGATATAGATGATGGACCAATTGATGTTGCCATTGGAGTTGCTGGAGAGACCGATACTTTACGCAAGG
GAAATAGTGATGGAGTAGTAGATATTGATGCAACCATTATAGATGATCAATGCCAATATTCTGTTTCAGTTCCCGCTTGCATGCCACAAGAAGATTGTGC
TGACAACGAGATTTCTCAGCTGGATACCCTTAGGTTATCTAGTTTTTCAAATTATGAGAATGAGTCAGTTGGTATGATTTCAGATAATGATGTCAGCATT
GAAATGAGCTCGTCAACTTCTGTCTCTACTCCCTCAGAAGATGAAGTTCCATCAGGAAACCAAGTGCTAGAGTGTGCTTCTCTTGGACATAAAATTGATT
ACACAAATTATACAGTTGCTGTTTTTCCTGACTATATTCTATGTGGCGACATATATGGTACAGAGTCTTGCTTAACTTTCTCAGGAAGCAGCATCAGAAT
GGAGGGTTCTACTGCAAATGGGGTCAAGGGAATATTTAATGCTGAGTGGAATCTTGACGATCTTATTAGTATTGAGTCAGAATGGTGTGAGATGGTTACA
ACTGCTATGGTTTATCTCTGCCTTAAATCAAAGGTTTCTGAAGGAGCTGGAAATACAAATGATGCTTCAGATGTGGACAAGTTGAAGTTTTCAGTTTATG
ATCCTCATTGGCATGAAGGAGAAGAAGCAATTAAATCATTGAATGTCAGATACAAGGATATCTGGAATGTTACTTCTGAATCTGATTTGGAGAAGGATGG
GAATGCCTCTTTTGGGCACAATGGCATGTTCACCTCAAAGCCTTATTTTCCTTTTATTCATGAAACATTCGAAGAGGTTATTTATCCTAAAGGTGATCCT
GATGCTGTTTCAATTAGTAAGCGAGATGTAGAGCTTCTACATCCTGAGACATTCATCAATGATACCATCATTGACTTTTATATCCAATATTTGAAGAATA
AAATTCAGCCAGATGACAGGCAAAGGTTCCACTTCTTCAATAGTTTTTTCTTTCAGAAGCTTGCTGATCTAGACAAGGGCCCATCTAATGCTTGTGAAGG
CAGGATAGCATTTCAACGTGTTCGTAAATGGACAAGAAAGTTGAATATTTTCGAGAAGGATTACATTTTTATTCCTGTAAATTACAGTCTTCACTGGAGT
TTGATTGTCATTTGTCATCCTGGTGAAGTGGTTCATTCCAGAGAGGATGAAAGTGGAAATTCAAGGAAAGTACCATGCATTTTGCACATGGATTCCATTA
GAGGAAGTCATAAGGGCCTCAAGAATCTTATTCAAAGTTATCTCTATGAAGAGTGGAGAGAAAGGCATAATGAGATTGTGGATGACACATTGTCAAAGTT
CTTACATTTACGGTTTGTCGCACTTGAGCTACCGCAGCAGGAAAATTTGTATGACTGTGGCCTGTTCTTACTCCATTATGTGGAGCTTTTTCTTGAAGAA
GCTCCAATTGATTTCAGTCCTTTTAAGATAACAGAGTTTTCAAACTTTCTTAGTAGAAATTGGTTTATACCTGGGGAGGCCTCACTGAAAAGGACACACA
TCCAGAAATTGATCTGTGAAATCATTGAAGATCAGTCTTGTACTCAATCTTCTGATCCAAATGAGCAAGAAACTGGAGTTGAGTTCCTTGAGGAGGTATG
CAGTGCAGTATCTGGTCCTGATACTGATATGGAGATTCAGGTTTCACTCACAGCAAAATCTCCAATCAGTGGTGCACAGCGACGAAGACTTGAAGAACTA
GGATTGAATTCCACGGATTTGCTCAAACCAGGAACTAGTGCAAGGTTTTTTTCCAATGGAAATTGCTGGCAGACAGGAACACTTCATTGGAGGACTTGCA
TGTCACCTGTGGAGGAAGCTGAGGAAACCGGCGAACGAATCTCTGATTCATTGTCATACACAGAAGATGATCAGCTACCTGCTGGCTTGGCAGTGGAATT
TCCCTCAACCTCACTTTCTCTCAAAGACCTTAGATCTCTCGGTTCATCTAGGAATAAAAAATACATGCAGCTTGAGGAACCTTATGACGACTCCAGTCCA
GAAGCATCTATCAGTAGATCCCCGAAATCATCAGAGATAGGGGTGGTAGGGGTTGATGAAGATCGTTTGCATTCACAGTTTGAGGGATTAGACCATCAAA
AACACACAGATTGCCATGAACTTTCTTCAAAATCAACCGAGCCTGAGGAATTTGTTGAGGGCTCTCGAGAGGGAAACTGCATGCACAATGACCATGTGAC
TAATGATTCGCCATCATCCTTTCATAACACTGGCAATTGCAATAAACTTGGCGCATCAGTTGACGCGATGAACACTGCTGAAAACTTTTTGAGAGGAGTC
CAAAAACCTGTTATCGATTTGACATCTGATGAACATACTGCAGAGAGGACAAAGCATACATCTGACGGCATTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G044000.1 pacid=42813127 polypeptide=Potri.017G044000.1.p locus=Potri.017G044000 ID=Potri.017G044000.1.v4.1 annot-version=v4.1
MTRSTRKFRVFEFDEEEEDRVEKESAKFVGKFRIQKRKRNDNKKDDDASSPLTKYNFLQCFGGCTGTVKIERSNEPIDIDDGPIDVDTGGEMATLRKGKS
DEVVYIDTTIIDNQCQYSVSVSARMPQEDCADKEEISQMETLITDDGPIDVDAGVAGEINTLCKGKSDEVVDIDTTILDDQCQCSVSVPAQMPQEDCAVK
EEISQLDTLGIDIDDGPIDVAIGVAGETDTLRKGNSDGVVDIDATIIDDQCQYSVSVPACMPQEDCADNEISQLDTLRLSSFSNYENESVGMISDNDVSI
EMSSSTSVSTPSEDEVPSGNQVLECASLGHKIDYTNYTVAVFPDYILCGDIYGTESCLTFSGSSIRMEGSTANGVKGIFNAEWNLDDLISIESEWCEMVT
TAMVYLCLKSKVSEGAGNTNDASDVDKLKFSVYDPHWHEGEEAIKSLNVRYKDIWNVTSESDLEKDGNASFGHNGMFTSKPYFPFIHETFEEVIYPKGDP
DAVSISKRDVELLHPETFINDTIIDFYIQYLKNKIQPDDRQRFHFFNSFFFQKLADLDKGPSNACEGRIAFQRVRKWTRKLNIFEKDYIFIPVNYSLHWS
LIVICHPGEVVHSREDESGNSRKVPCILHMDSIRGSHKGLKNLIQSYLYEEWRERHNEIVDDTLSKFLHLRFVALELPQQENLYDCGLFLLHYVELFLEE
APIDFSPFKITEFSNFLSRNWFIPGEASLKRTHIQKLICEIIEDQSCTQSSDPNEQETGVEFLEEVCSAVSGPDTDMEIQVSLTAKSPISGAQRRRLEEL
GLNSTDLLKPGTSARFFSNGNCWQTGTLHWRTCMSPVEEAEETGERISDSLSYTEDDQLPAGLAVEFPSTSLSLKDLRSLGSSRNKKYMQLEEPYDDSSP
EASISRSPKSSEIGVVGVDEDRLHSQFEGLDHQKHTDCHELSSKSTEPEEFVEGSREGNCMHNDHVTNDSPSSFHNTGNCNKLGASVDAMNTAENFLRGV
QKPVIDLTSDEHTAERTKHTSDGI
|
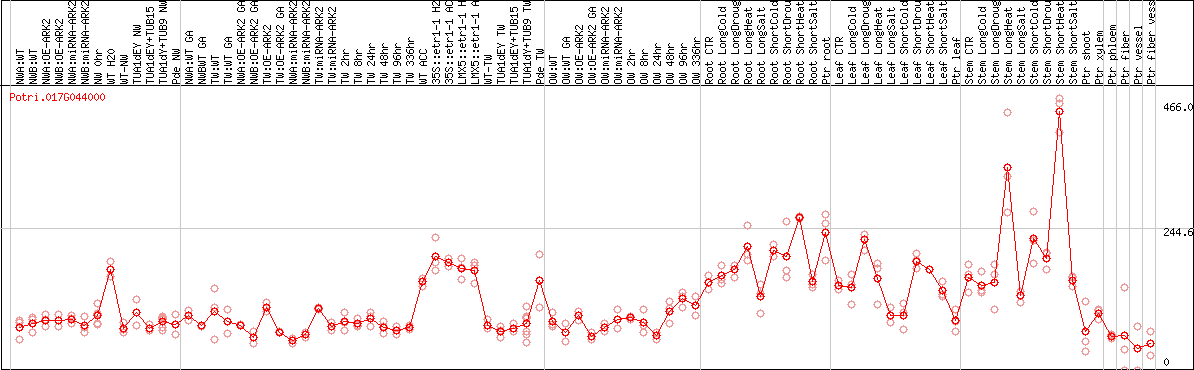
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G044000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
