External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G33580 339 / 9e-117
ATBCA5
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
AT1G58180 259 / 2e-86
ATBCA6
A. THALIANA BETA CARBONIC ANHYDRASE 6, beta carbonic anhydrase 6 (.1.2.3.4)
AT1G70410 236 / 3e-77
ATBCA4
beta carbonic anhydrase 4 (.1.2.3)
AT1G23730 225 / 1e-72
ATBCA3
BETA CARBONIC ANHYDRASE 3, beta carbonic anhydrase 3 (.1)
AT5G14740 224 / 3e-72
BETACA2, CA18, CA2
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
AT3G01500 221 / 4e-71
SABP3, ATBCA1, CA1, ATSABP3
ARABIDOPSIS THALIANA SALICYLIC ACID-BINDING PROTEIN 3, BETA CARBONIC ANHYDRASE 1, carbonic anhydrase 1 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G114600
535 / 0
AT4G33580 335 / 4e-115
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
Potri.005G156600
350 / 7e-121
AT4G33580 303 / 1e-102
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
Potri.010G041100
229 / 6e-74
AT5G14740 392 / 4e-138
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Potri.008G189800
227 / 1e-73
AT5G14740 375 / 5e-133
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Potri.001G348900
218 / 1e-68
AT3G01500 429 / 2e-151
ARABIDOPSIS THALIANA SALICYLIC ACID-BINDING PROTEIN 3, BETA CARBONIC ANHYDRASE 1, carbonic anhydrase 1 (.1.2.3)
Potri.015G076000
197 / 1e-61
AT5G14740 303 / 4e-104
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Potri.015G075902
196 / 4e-61
AT5G14740 304 / 1e-104
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015833
355 / 5e-122
AT4G33580 306 / 2e-103
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
Lus10020415
343 / 1e-114
AT1G15820 379 / 9e-130
light harvesting complex photosystem II subunit 6 (.1)
Lus10009274
288 / 8e-97
AT4G33580 281 / 2e-94
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
Lus10015892
223 / 1e-71
AT4G33580 205 / 3e-65
A. THALIANA BETA CARBONIC ANHYDRASE 5, beta carbonic anhydrase 5 (.1.2)
Lus10022235
225 / 2e-71
AT3G01500 460 / 3e-163
ARABIDOPSIS THALIANA SALICYLIC ACID-BINDING PROTEIN 3, BETA CARBONIC ANHYDRASE 1, carbonic anhydrase 1 (.1.2.3)
Lus10030614
220 / 2e-70
AT1G70410 351 / 6e-123
beta carbonic anhydrase 4 (.1.2.3)
Lus10030874
201 / 3e-63
AT5G14740 319 / 8e-111
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Lus10016443
200 / 4e-62
AT5G14740 298 / 4e-101
CARBONIC ANHYDRASE 18, BETA CARBONIC ANHYDRASE 2, carbonic anhydrase 2 (.1.2.3.4.5)
Lus10008780
150 / 9e-43
AT3G01500 357 / 1e-122
ARABIDOPSIS THALIANA SALICYLIC ACID-BINDING PROTEIN 3, BETA CARBONIC ANHYDRASE 1, carbonic anhydrase 1 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00484
Pro_CA
Carbonic anhydrase
Representative CDS sequence
>Potri.017G044700.1 pacid=42813093 polypeptide=Potri.017G044700.1.p locus=Potri.017G044700 ID=Potri.017G044700.1.v4.1 annot-version=v4.1
ATGGCTATCCCATCACCACCTTTCTCTCTTTCAAAACACTCCTTATCCAATTCCCCTTCATTTCATGCTTCCAACCCTTCATTGGATCCCTCTAAAGCTT
CAGCCTTGGGAACCCAAAATGTCTTTGGTTCTAAGGCTAATTTAGGAGGAGTTGAGCAGACCCATTTGAGATTGCGGAATAATTTGAAGACAAATTCAGA
TTTGAGATTACAGGCTTCCAGGGAGCCTCCTGGACTGACTAAGGAACTTAAAAGTGACAAATCCGAGCGCATGGAAAGAATCGAACATGGTTCTGATTTA
TTTGATGAGATGAAACAACGGTTCCTGAGCTTCAAAAAGCATAAATACATGCAAAACTTGGAACTCTACGAAAAGCTTGCCAAAGGGCAAGCACCAAAGT
TTATGGTGATTGCTTGTGCAGACTCAAGGGTTTGCCCTTCGTCCATCTTAGGATTCCAGCCTGGTGAAGCTTTTGTCGTTCGCAATGTTGCAAATATGGT
GCCACCCTATGAGAATGGACCATCTGAAACAAATGCAGGCTTAGAGTTTGCTGTAAATTCTCTGAAAGTTGAAAACATTTTAGTAATTGGTCACAGTCAA
TGCGGAGGCATTCGCGCTCTAATGAGTATGCATGATGACGTAGAAACAAGTAGCCTCATTGGAAGTTGGGTTTCTGTGGGGATGAATGCAAGAGTAAGAA
CTAAGGCAGCCACGAAACTTCTGAACTTTGACCAGCAGTGCAAACATTGTGAAAAGGAATCAGTCAACTGTTCATTGGCAAACCTCCTCACTTATCCATG
GGTGGAAGAAAAAGTGAGGAATGGGGAACTTGCTATTCACGGTGCCTATTATGACTTCGTCGACTGTGCATTTGAGAAATGGACTCTGGATTACAAGGAA
AGCAATCTGAAGGACAAAGGTGGGAGAGTTGCAGTAAAAGACCGAGCATTTTGGTTCTGA
AA sequence
>Potri.017G044700.1 pacid=42813093 polypeptide=Potri.017G044700.1.p locus=Potri.017G044700 ID=Potri.017G044700.1.v4.1 annot-version=v4.1
MAIPSPPFSLSKHSLSNSPSFHASNPSLDPSKASALGTQNVFGSKANLGGVEQTHLRLRNNLKTNSDLRLQASREPPGLTKELKSDKSERMERIEHGSDL
FDEMKQRFLSFKKHKYMQNLELYEKLAKGQAPKFMVIACADSRVCPSSILGFQPGEAFVVRNVANMVPPYENGPSETNAGLEFAVNSLKVENILVIGHSQ
CGGIRALMSMHDDVETSSLIGSWVSVGMNARVRTKAATKLLNFDQQCKHCEKESVNCSLANLLTYPWVEEKVRNGELAIHGAYYDFVDCAFEKWTLDYKE
SNLKDKGGRVAVKDRAFWF
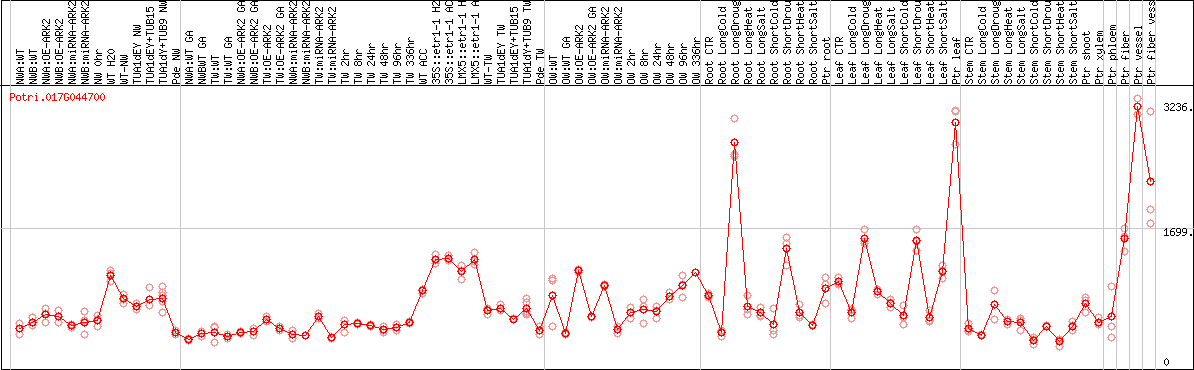
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G044700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.