Potri.017G046200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G046200.1 pacid=42813516 polypeptide=Potri.017G046200.1.p locus=Potri.017G046200 ID=Potri.017G046200.1.v4.1 annot-version=v4.1
ATGAATTATAGACCATATGAAAAACATGTTCCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTCTTTCTCTCTCTAGTTCTATATATCATAGCTTGCTCA
CTACATATTTTCATACAACAAACTCCTCGTTTAACCTAAGTAGATCTCCTCTCCTATTAGCAACAATGTTGAAGCCACAGCTTCACCAGTCACACCTTTC
AACAAAAATCCCTTTTCTACTACCTAAACCATTCATCCATGGCAGTGGTCATGCTTCTTTTCCTGTCTATTCAAGGTCCTTGTCCACTAAAGCAAACAAA
GAAGTTCGTGTTGGCCATAAACATGGAAGCATTAAATCTATAGCTAGTGTCACACAGCAGTCCACAGATGTTAAGGCTGTGGTGACTGTGAAACAAACAG
TTGTTGATTTTTGGACAGAAATAGGGATAGAACGAGGACTTGATGATTTGAAAGACCTGTTTGGCAAAACACTCCTCTTGGAGCTTGTTAGTGCCGAGCT
TGACTCAAAGACGGGATTAGAGAAGCCCTCAATTCGAAAGTACGCACACAAGATAGATCATGAAGGAGAGGATATCAAGTATGAAGCGGATTTTGTGGTT
CCACCAGATTTTGGAGAGATTGGTGCCATTTTTGTTGAAAATGAGCACCATAAAGAGATGTACCTTCATGATGTTGTGTTTGATGGCTTCCCTACTGGAC
CAGTTCATGTGACCTGTGATTCATGGATCCATTCGAAGTTTGACAATAAAAAGAAGAGACTCTTTTTCACTAACAAGTCATACTTGCCATCAGAAACACC
AAATGGATTGACAAAACTGAGAGAGGAAGAACTTGAAACTTTACGAGGCAACGACAACGGAGAACGAAAGAACGGTGAAAGGATCTATGACTATGACGTG
TACAATGATCTTGGAAATCCTGATAGTGATCCAGAGACAGCAAGACCGGTGCTAGGTGGCAAAGAGCACCCGTACCCTAGGCGTTGCCGTACTGGCCGAC
CACGGACCGAGTCAGATCCACTTACGGAGACAAGGAGTAGCAGCTTCTACGTGCCTCGAGATGAAGAATTCTCAGAAATAAAGATGGGCACGTTTTCTGC
CAAGACTCTCAAATCAGTATTGCATGCCCTAGTTCCTTCTCTATCAACCGCTATTGTTGACAGTGAACTTGGATTCCCATTCTTCAGTTCCATTGACGCA
CTATTCAATGAAGGAATCAATTTGCCTCCCCTTAAGAAACAGGGGTTTTGGAAAGATCTTCTGCCTAACCTTTTCAGGGCCATTACTGATGGAACTAAAG
ATGTCTTGAAATTTGAGACTCCTGATACCATGGAGAGAGACAGATTTTTTTGGTTTAGGGATGAAGAATTTGCTAGACAGACTCTTTCTGGTCTCAACCC
TTGTAGCATCAAAATGGTCACGGAATGGCCATTAAGGAGTAAACTAGACCCTGAAATCTATGGTCCCCAAGAATCAGCAATCACTACAGAAATGGTTGAG
CAAGAAATCAAAGGGTTCATGACATGTGGACAGGCAGTAAAAGATCAGAAGCTTTTCATCCTTGACTACCATGATTTATTCCTACCTTTTGTAAGCAAAA
TAAGAGAGCTGAAAGGCACGACATTATACGGGTCACGGACATTGTTCTTCTTAACCCATGAAGGAACATTGAGACCACTAGCTATTGAGCTCACTCGACC
ACCGATGGATGGTAAGCCGCAATGGAAGCAAGTGTTTAGACCAGCTTGGCATTCCACTGGTGTTTGGCTTTGGAGGCTTGCCAAAGCTCATGTCCTTGCA
CATGAGTCTGGCTATCACCAACTTATTAGTCACTGGCTAAGAACACATTGCTGCACAGAACCATACATTATCGCGGCCAATCGCCAACTTAGTGAAATGC
ATCCAATCTACAGACTCTTGCACCCGCATTTCCGGTACACAATGGAGATCAATGCTCTAGCTCGACAATATCTAATTAATGCAAAAGGGATTATAGAGAC
GTCATTCTTTCCAGGCAAGTATTCTATGGAATTAAGCTCTGTTGTCTATGATCAAGAATGGCGGTTTGACTACGAGGCGCTGCCCAAGGATCTAATAAAC
AGGGGAATGGCTGTGGAAGATCCATCTGCTCCACATGGATTGAAATTGATGGTGGAAGACTATCCATATGCCAATGATGGTTTGGTTTTATGGGATATCA
TCAAAGAGTGGGTTAGTGACTATGTCAATCACTACTATCCAGACTCAAGTCTCATTGTGTCTGATAATGAGCTCCAAGCATGGTGGACAGAAGTTCGAAC
GGAAGGTCACGCTGACAAGAAAGATGAACCCTGGTGGCCTGTGCTTAAAACACCTCAGGACCTTATTGAAACCATGACAACTATCATATGGATAGCCTCC
GGTCATCATGCCGCAGTGAATTTCGGACAATATACATATGCAGGATACTTTCCAAATAGGCCTACAACTGCTAGGATGAATATGCCAACCGAAGACCCGA
ACGATGAGTTGCTGAAACTATTTTGGGAGAAACCTGAAGTGATACTATTGACAACATTCCCTTCACAAATTCAGGCAACAACAGTGATGGCTATATTGGA
TGTGTTATCAAATCATTCTCCCGATGAGGAGTATCTTGGGCAGCAAATAGAGCCATCATGGACAGAGGAACCGGCTATAAATGCGGCTTTCGTAAAGTTC
AATGGGAGGTTGAAGGAGTTTGAAGGAATTATTGATGAAAGGAATGCTGACATTAAATTGAAGAATAGAAATGGAGTTGGCGTTGTGCCTTATGAGCTTT
TGAAGCCATTTTCTGACCCTGGAGTTACAGGAAAAGGAGTTCCTTGCAGCATCTCTATCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G046200.1 pacid=42813516 polypeptide=Potri.017G046200.1.p locus=Potri.017G046200 ID=Potri.017G046200.1.v4.1 annot-version=v4.1
MNYRPYEKHVPLSLSLSLSLSLSLSSSIYHSLLTTYFHTTNSSFNLSRSPLLLATMLKPQLHQSHLSTKIPFLLPKPFIHGSGHASFPVYSRSLSTKANK
EVRVGHKHGSIKSIASVTQQSTDVKAVVTVKQTVVDFWTEIGIERGLDDLKDLFGKTLLLELVSAELDSKTGLEKPSIRKYAHKIDHEGEDIKYEADFVV
PPDFGEIGAIFVENEHHKEMYLHDVVFDGFPTGPVHVTCDSWIHSKFDNKKKRLFFTNKSYLPSETPNGLTKLREEELETLRGNDNGERKNGERIYDYDV
YNDLGNPDSDPETARPVLGGKEHPYPRRCRTGRPRTESDPLTETRSSSFYVPRDEEFSEIKMGTFSAKTLKSVLHALVPSLSTAIVDSELGFPFFSSIDA
LFNEGINLPPLKKQGFWKDLLPNLFRAITDGTKDVLKFETPDTMERDRFFWFRDEEFARQTLSGLNPCSIKMVTEWPLRSKLDPEIYGPQESAITTEMVE
QEIKGFMTCGQAVKDQKLFILDYHDLFLPFVSKIRELKGTTLYGSRTLFFLTHEGTLRPLAIELTRPPMDGKPQWKQVFRPAWHSTGVWLWRLAKAHVLA
HESGYHQLISHWLRTHCCTEPYIIAANRQLSEMHPIYRLLHPHFRYTMEINALARQYLINAKGIIETSFFPGKYSMELSSVVYDQEWRFDYEALPKDLIN
RGMAVEDPSAPHGLKLMVEDYPYANDGLVLWDIIKEWVSDYVNHYYPDSSLIVSDNELQAWWTEVRTEGHADKKDEPWWPVLKTPQDLIETMTTIIWIAS
GHHAAVNFGQYTYAGYFPNRPTTARMNMPTEDPNDELLKLFWEKPEVILLTTFPSQIQATTVMAILDVLSNHSPDEEYLGQQIEPSWTEEPAINAAFVKF
NGRLKEFEGIIDERNADIKLKNRNGVGVVPYELLKPFSDPGVTGKGVPCSISI
|
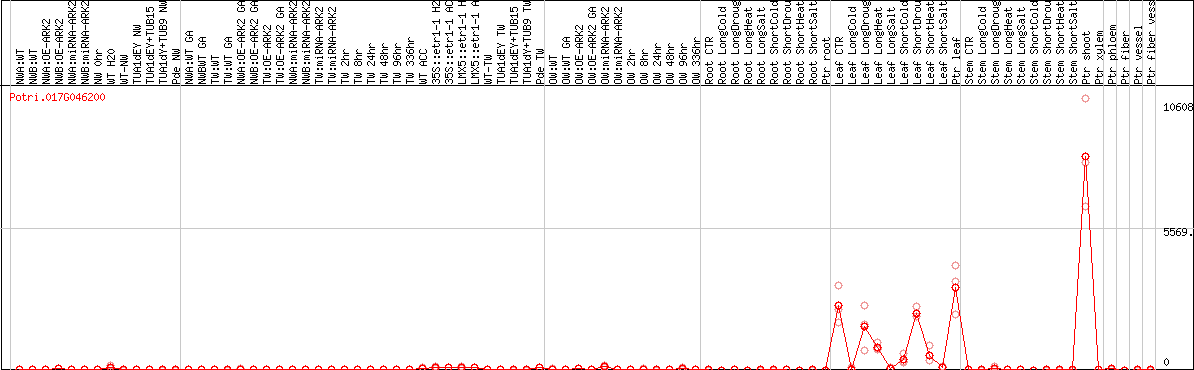
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G046200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
