Potri.017G050100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G050100.4 pacid=42813981 polypeptide=Potri.017G050100.4.p locus=Potri.017G050100 ID=Potri.017G050100.4.v4.1 annot-version=v4.1
ATGATCACCAGCAATAACCACCACCCCTCCACACGCCACCACCACCACTCCCAACACCGTCAGTCGCAGCATTTCAACCGTCACCGGAACTTCCGAAACC
ACCAAAACCAAAGCCTTTTTGTAGTCAGGCTTTTGTCCAACCACCGTAATAATAACCGCACGCAATCCCCACTCGAAACCCTAATTTCCCAATGTAACCC
TAAACCTGACAAATCCGATACAAACCCCACGTCCGCCGTTGCTGCTCGTTTGTTTTTCCACGACCAGAGCGATGCTATCGCCGCCGTCGTCTTCCTCTGG
GAACGGAGGCTGGCTGGGGATCATGTGTATACTCCGGTGACGGATTTTGATGTAAATGAAGGGGACTTGAATGAGAGGATTAGAGGTTTGTTTAAATTGT
ATGCAGAGAGGGTGTTAGAAGGGGAGGTAGTGAAGAAATTGGAGAGAAAGATTGAAAATTTGGCCGTGGAAATTGGTAAATTTACGAGCTTTTTCAAGCG
GCCGAAAGGGGTTAGGGTTTATAGTGAGAATAAGGTGAAGAAGGAGGCGTTGAGGGTGGAAATGGAGGTGGTTGTGAAGAGGGTGGAGGAGTTTAGGAAG
GGAATGAGGTGTTTGATGGATTGCATTGAAGGGAAGGAGATTGGTGATTTGGGGGTTTTGAGGGTTTATGATGAGGGTAATGGGAGAAAGATGGGAATTT
TTTATTATTGGAGCAGGATTCATTTTCTGATATTGAGAGAGTGTAGAAGGGTAGAAAATGGGTTGCCTGTTTATGGGTTTAGGAGTGAGTTTTTGAAGAT
GCTGCGTTCACAGCAGGTGATGGTCTTGATTGGAGAAACAGGTTCAGGGAAGAGTACGCAGTTGGCACAATTTATTGCTGATTCAGGAGTTGCATCAAGT
GGTTCAATTTTGTGCACTCAACCCCGCAAAATTGCTGCTATATCCCTGGGAAAAAGGGTCGGTGAAGAATGCAATGGATGTTATGAAGATAATTCAATTA
TCTGCTACCCATCTTATTCATCTTCCCAGCAGTTTGGTTCGAAGGTAATATATATGACTGATCACTGTCTACTGCAAAACCTCATGAAAGATAAAAATTT
GTTTGGAGTTTCGTGCATTATTGTAGATGAGGCTCATGAAAGAAGCTTGAATACTGATCTTCTTTTGGGCTTGCTAAAAGAATTACTTCAGGAAAGGCCT
GACTTACAGCTAATTATAATGTCTGCAACAGTTGATGCAAGCAAATTATCCAGTTATTTCTTTGGCTGTGGAACATTTCATGTGTTGGGGAGAAGCTTTC
CTGTTGAAATCAAATATGCTCCTGCTGCATCTAGAGAGTCCCTTGACCCCTTGCCGAGCTCCAATAATGCAGCTCCATATGTTTGTGATGTTGTTAAGAT
GGCAACGGAAATTCATGCAGCAGAGGAGGATGGGGCAATACTTGCTTTTTTGACTTCACAGGCAGAAGTAGAATGGGCATGTGAAAAGTTTCAGTCTCCC
TCTGCAATTGCGCTCCCATTACATGGGAAGCTTTTCCATGAAGAGCAATGTCGTGTTTTTCAGAATTACCCAGGAAAGAGAAAAGTTGTTTTTGCTACAA
ATTTAGCGGAAACATCCATAACAATTCCAGGGGTCAAGTACGTGGTTGATTCTGGTTTAGTTAAAGATAGCAGGTTTGAATCTTCATCTGGTATGAATGT
GCTCAGAGTTTCCAAGATCAGCCAAAGTTCTGCAAATCAACGAGCTGGCCGTGCTGGTAGAACAGATCCTGGCAAGTGTTACCGGTTGTACTCGGTTTCT
GATTATCAGTCAATGGATTTGCATCAGGAACCTGAAATTTGTAAAGTTCATCTTGGCATTGCAGTTTTGAGGATTCTTGCCTCTGGTATTAAGAATGTGC
TAGAGTTTGATTTTATTGATGCTCCAAGTGTGGATGCAATCAACAAGGCAATCAGGAACCTTGTTCAGTTAGGTGCTGTTGCCTGGAAACATGATGCTTT
TGTATTAACTGCTGATGGACACTACTTAGTTAAATTGGGTATGGAGCCTCGACTTGGGAAAATTATTCTTGAAAGTCTCCGTTATGGTTTGCGCAAGGAG
GGAGTTGTCCTTGCTGCAGCTATGGCAAATGCCAGTAACATTTTTTGCAGAGTGGGTACTTATGATGAGAAACTGAAGTCTGATTGCCTTAAGGTCCGGT
TCTGCCATCACGATGGAGATCTCTTCACTTTGCTCTCCGTCTACAGGGAATGGGAGAGTTTACGGCAAGAGAATAGAAATAAATGGTGTTGGGAAAATAG
AATCAATGCAAAAACTATGCGAAGATGCAGAGATACAGTATTGGAGTTGGAAAACTGTTTGAAAAATGAACTCAATATCATTATTCCAACTTACTGGCTT
TGGGATCCTCTTGTTGCCTCTGTGCATGATGAAAACATGAAGAAGATTATACTTTCTTCACTAGCAGATAATGTAGCTATGTATTCTGGATATGATCGCC
TTGGTTATGAGGTTGTTTTAAGTGGTGAATACTTCCAACTTCACCCTTCATGTTCATTACAAGTGTATAACCAGAAGCCACATTGGGTTGTCTTTGCTGA
ACTCTTGTCCATATCAAGTCAATATTTGGTTTGTGTTACAGCAATTGACTTTGACAGTTTATCTACCTTCATACATCCTTTATTTGATGTCTCAAAGATG
GAATCTAGAAAACTGCAGTTGAGGGTGATAAAAGGATTTGGAGGTGTTGCTTTGAAACGGTTTTGTGGAAAGTCAAATAGTAGCTTGATAGCACTTGTTT
CTCGCATGCGTGCTATTTACATGGATGAACGCATTGGAATTGAAATCAATGTGGGTGACAACGAGATTCAATTGTTTGCATCGTCCAAAGATATTGAAAA
GATTTATGAATATGTCAATAATGCGTTAAGATATGAAACAAAGTGGTTAAGGAATGAATGCTTGGAGAAGTGCTTATATCATGAGGTACGGGCTGGTGCT
TCCCCCCCTGTTGCACTTGTTGGAGCTGGTGCTGAAATTAAGCATCTGGAGTTGGGAAATAGATGTTTAACAGTGGATGTGCACCTTTCCAATGTGAATG
TGGTAGATGACAAGGAAGTTTTGACATTCTTAGAGAAATCTGTATCTGGAATTTGTGGTTATAACAAATTCACGGGCATTGGACAGCATGGTGGTGATGC
AGAAAGGTGGGGCAGGGTGTCATTCCTTACTCCTGAAGCAGCTAGAAAAGCTCTATATTTTAATGGATCCGAGTTATGTGGATGTGTACTTAAGTTGTCC
CTGTCTCGGTCATCTGTTGGAGGTATTCGAAAGTCATCATTTGCTGCCGTAAAAGCAAAAATCAGTTGGCCACGCCGGTATAGCAAAGGGTATGCAATTG
TCAGATGTGAAAGGAATGATGCCCAGTTCATTGTTGATGATTGCTTTAATGTTCTGATTGGAGGCAGGTTTGTTCAGTGTCAGACCAGCACAAGAGACAT
GAACTCTGTAGTTATCAGAGGACTGGATAAAGAAACTTCTGAAGCAGAAATTTTAGAAGTTCTGCACAAGACAACAAATAGGAGAATTTTGGATGTATTC
CTAATCAGAGGAGATGAAGCAAATAATCATTCTGTTGATGCTTTTGAGCAAGCAATTCTCAAGGAAATTGCTCCTTTCATGCCTAGCCAGGGCCCTTTGA
GCAATTACTGCCATGTTCAGGTCTTTGCCCCTGAGCCAAAGGATAGTTTCATGAAGGCTTGGATCACTTTTGATGGAAAATTACATTTGGAGGCAGCTAA
AGCTCTGCAACATATGCAAGGAAAAGCACTTGCTGGTTGTTTTTCATGGCAGAAGATGCAATGCCAGCAGGTGTTCCATAGCTCTGCATCATGTTCTGCT
TCTGTTTATGCATTCATTGAGAGGCAGCTGAATATTTTACTGAAAAGCTTCAAGTTTCGACCTGGTGTGTGTTGCAATTTAGAAAGGAATGAGAATGGCT
CTTATAGAGTGAAGATATCTGCTAATGCAACAAAGACAGTAGCTGAGTTGAGAAGGCCGCTGGAGCAACTGATGAATGGAAAGAAGGTAAATCATTGTAG
CCTCACCCCATTGGTTCTGCAGCTTCTCTTTTCCAAAGATGGCATCATGCTTATGAAGTCACTGCAGCAAGAGATGGGAACATATATTCTCTTTGATAGG
CAGAACCTCACTGTTAGGATCTTTGGCCCCGAGAAGAAAGTTGCTCTTACTGAACAGAAGTTGATAGCATCTCTTCTAGCCCTCCATGATAAGGAACAAA
CAGATATCCGCCTTCGAGGTGGAGCAATGCCGTATGATCTTATGAAAAAGGTGGTGGAGAAGTTTGGTCCTGACTTGCATGTGCTCAAAGAGACGTTCCC
TGAGGCTGAATTTATGCTTAATACAAGGCGACATGTTATTTCTTTTTCTGGAAAAAAGGATCTGAGACTCCAAGTGGAGCAGATGATTCGTGACTTTGTG
CGGTCTGTCGGTGTTAATGGATCCATCAAACGATATGAAGATGACAATATTGCCTGCCCTATTTGCCTTTGTGAGGTTGAGGACTGTTACCAGCTTGAGG
CTTGTGGGCATAAATTCTGCCAGTCATGCTTGGTTGAACAGTTAGAGTCAGCAATGAGAGGACGTGATGGCTTCCCTGTAGGTTGTGCTCATGAAGGTTG
TGGCATGCACATTTGGCTTACAGATCTGAAGTCTCTTTTACCATGTGAGAAGTTAGAGGATCTCTTCAGGGCCTCCTTGTCAGCCTTTGTGGCATCTAGT
GGTGGAACTTACAGATTTTGCCCATCTCCTGATTGCCCTTCAGTATATCACGTAGCAAGTGGGATGGTTGGTGATCTATTTGTTTGTGGAGCATGCTATG
CCGAGACATGTACAAGGTGCCATGTGGAGTATCATCCATTCGTATCATGTGAAAAATACAAGGAGCTCAAGGAGGATCCTGACATGTCCTTGAAGGAATG
GTGCAAGGGAAAAGAGCATGTTAGAAACTGCCCAGTATGTGGATACACAATTGAGAAGGTAGATGGCTGCAACCACATTGAGTGCAGATGTGGGAAACAC
ATCTGCTGGGTCTGCTTGGAGGTCTTTATGAGCGGTGATGACTGCTATGCGCACTTGAGGTCTGTACATCCTGCTACATTATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G050100.4 pacid=42813981 polypeptide=Potri.017G050100.4.p locus=Potri.017G050100 ID=Potri.017G050100.4.v4.1 annot-version=v4.1
MITSNNHHPSTRHHHHSQHRQSQHFNRHRNFRNHQNQSLFVVRLLSNHRNNNRTQSPLETLISQCNPKPDKSDTNPTSAVAARLFFHDQSDAIAAVVFLW
ERRLAGDHVYTPVTDFDVNEGDLNERIRGLFKLYAERVLEGEVVKKLERKIENLAVEIGKFTSFFKRPKGVRVYSENKVKKEALRVEMEVVVKRVEEFRK
GMRCLMDCIEGKEIGDLGVLRVYDEGNGRKMGIFYYWSRIHFLILRECRRVENGLPVYGFRSEFLKMLRSQQVMVLIGETGSGKSTQLAQFIADSGVASS
GSILCTQPRKIAAISLGKRVGEECNGCYEDNSIICYPSYSSSQQFGSKVIYMTDHCLLQNLMKDKNLFGVSCIIVDEAHERSLNTDLLLGLLKELLQERP
DLQLIIMSATVDASKLSSYFFGCGTFHVLGRSFPVEIKYAPAASRESLDPLPSSNNAAPYVCDVVKMATEIHAAEEDGAILAFLTSQAEVEWACEKFQSP
SAIALPLHGKLFHEEQCRVFQNYPGKRKVVFATNLAETSITIPGVKYVVDSGLVKDSRFESSSGMNVLRVSKISQSSANQRAGRAGRTDPGKCYRLYSVS
DYQSMDLHQEPEICKVHLGIAVLRILASGIKNVLEFDFIDAPSVDAINKAIRNLVQLGAVAWKHDAFVLTADGHYLVKLGMEPRLGKIILESLRYGLRKE
GVVLAAAMANASNIFCRVGTYDEKLKSDCLKVRFCHHDGDLFTLLSVYREWESLRQENRNKWCWENRINAKTMRRCRDTVLELENCLKNELNIIIPTYWL
WDPLVASVHDENMKKIILSSLADNVAMYSGYDRLGYEVVLSGEYFQLHPSCSLQVYNQKPHWVVFAELLSISSQYLVCVTAIDFDSLSTFIHPLFDVSKM
ESRKLQLRVIKGFGGVALKRFCGKSNSSLIALVSRMRAIYMDERIGIEINVGDNEIQLFASSKDIEKIYEYVNNALRYETKWLRNECLEKCLYHEVRAGA
SPPVALVGAGAEIKHLELGNRCLTVDVHLSNVNVVDDKEVLTFLEKSVSGICGYNKFTGIGQHGGDAERWGRVSFLTPEAARKALYFNGSELCGCVLKLS
LSRSSVGGIRKSSFAAVKAKISWPRRYSKGYAIVRCERNDAQFIVDDCFNVLIGGRFVQCQTSTRDMNSVVIRGLDKETSEAEILEVLHKTTNRRILDVF
LIRGDEANNHSVDAFEQAILKEIAPFMPSQGPLSNYCHVQVFAPEPKDSFMKAWITFDGKLHLEAAKALQHMQGKALAGCFSWQKMQCQQVFHSSASCSA
SVYAFIERQLNILLKSFKFRPGVCCNLERNENGSYRVKISANATKTVAELRRPLEQLMNGKKVNHCSLTPLVLQLLFSKDGIMLMKSLQQEMGTYILFDR
QNLTVRIFGPEKKVALTEQKLIASLLALHDKEQTDIRLRGGAMPYDLMKKVVEKFGPDLHVLKETFPEAEFMLNTRRHVISFSGKKDLRLQVEQMIRDFV
RSVGVNGSIKRYEDDNIACPICLCEVEDCYQLEACGHKFCQSCLVEQLESAMRGRDGFPVGCAHEGCGMHIWLTDLKSLLPCEKLEDLFRASLSAFVASS
GGTYRFCPSPDCPSVYHVASGMVGDLFVCGACYAETCTRCHVEYHPFVSCEKYKELKEDPDMSLKEWCKGKEHVRNCPVCGYTIEKVDGCNHIECRCGKH
ICWVCLEVFMSGDDCYAHLRSVHPATL
|
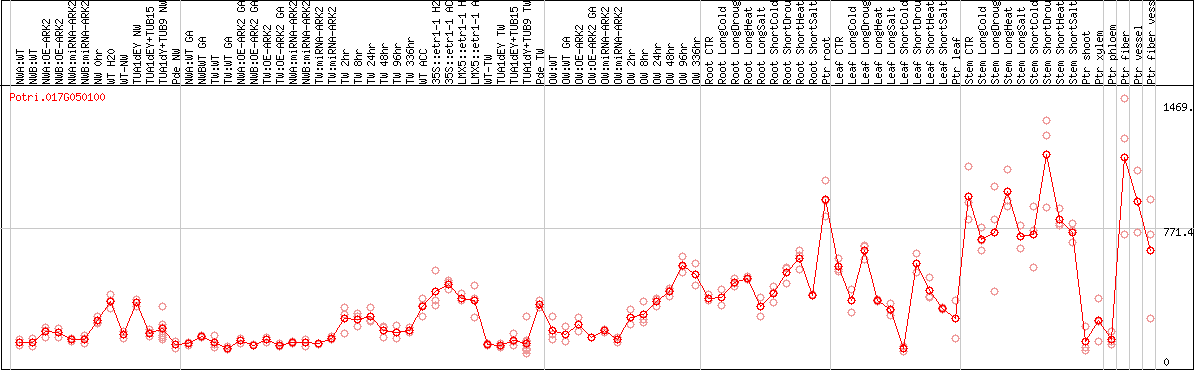
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G050100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
