External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G06350 59 / 2e-12
EMB3004, MEE32
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G029800
74 / 8e-18
AT3G06350 630 / 0.0
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Potri.010G019000
61 / 6e-13
AT3G06350 736 / 0.0
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Potri.014G135500
45 / 2e-07
AT3G06350 444 / 8e-151
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Potri.005G043400
42 / 3e-06
AT3G06350 508 / 1e-175
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Potri.013G029900
39 / 4e-05
AT3G06350 516 / 4e-179
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012797
66 / 1e-14
AT3G06350 748 / 0.0
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Lus10033981
63 / 9e-14
AT3G06350 752 / 0.0
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Lus10005928
52 / 1e-09
AT3G06350 180 / 9e-52
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
Lus10040908
46 / 8e-08
AT3G06350 291 / 2e-92
MATERNAL EFFECT EMBRYO ARREST 32, EMBRYO DEFECTIVE 3004, dehydroquinate dehydratase, putative / shikimate dehydrogenase, putative (.1)
PFAM info
Representative CDS sequence
>Potri.017G103401.1 pacid=42814407 polypeptide=Potri.017G103401.1.p locus=Potri.017G103401 ID=Potri.017G103401.1.v4.1 annot-version=v4.1
ATGATCCTTGCAAATACCACATCTGCTGGAATGGAACCAAGGATAGCTTCTGAACACTATTCATTAGTCTTCAATACAATTTACACTCCAAAATTAGCTT
CTCTCTTAAGAGAAGCACAGGAGGCTGGAAGCACCATTGTTTAA
AA sequence
>Potri.017G103401.1 pacid=42814407 polypeptide=Potri.017G103401.1.p locus=Potri.017G103401 ID=Potri.017G103401.1.v4.1 annot-version=v4.1
MILANTTSAGMEPRIASEHYSLVFNTIYTPKLASLLREAQEAGSTIV
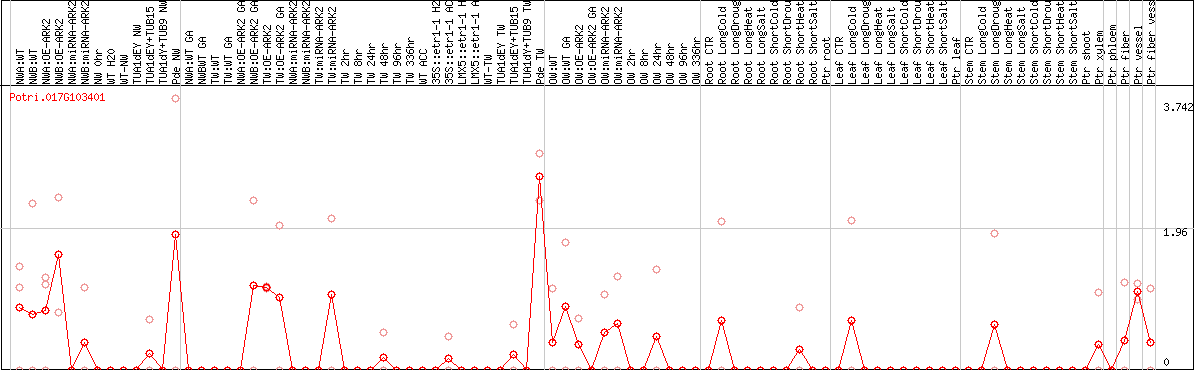
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G103401 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.