External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G36930 159 / 3e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G72920 147 / 4e-44
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G72940 146 / 8e-43
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72900 141 / 6e-41
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72930 135 / 7e-41
TIR
toll/interleukin-1 receptor-like (.1.2)
AT1G72890 139 / 1e-39
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
AT5G17680 142 / 2e-39
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT1G27170 141 / 4e-39
transmembrane receptors;ATP binding (.1.2)
AT1G72840 138 / 5e-38
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT1G72910 133 / 6e-38
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G104301
316 / 9e-101
AT5G17680 566 / 1e-178
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.013G098100
282 / 9e-99
AT5G36930 159 / 1e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.013G097050
280 / 4e-98
AT5G36930 158 / 3e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.013G097300
306 / 3e-97
AT5G17680 598 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.013G096849
276 / 4e-96
AT5G36930 167 / 4e-48
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.017G105201
298 / 8e-95
AT4G12010 549 / 4e-174
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Potri.017G104901
293 / 2e-92
AT5G17680 536 / 2e-167
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.017G103701
292 / 4e-92
AT5G17680 544 / 5e-170
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.013G097000
288 / 3e-91
AT5G17680 571 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011104
192 / 4e-57
AT5G17680 574 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10030839
179 / 3e-52
AT4G12010 552 / 7e-176
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10006929
163 / 1e-50
AT1G72890 146 / 1e-39
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
Lus10041060
173 / 3e-50
AT5G17680 641 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10042752
158 / 8e-50
AT4G16990 145 / 5e-41
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
Lus10014671
159 / 5e-49
AT5G36930 144 / 4e-39
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10006928
165 / 1e-48
AT5G36930 165 / 5e-43
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10003749
160 / 5e-46
AT5G17680 517 / 5e-161
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10039850
160 / 1e-45
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10013729
158 / 1e-45
AT5G36930 310 / 2e-92
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF13676
TIR_2
TIR domain
Representative CDS sequence
>Potri.017G105501.1 pacid=42813192 polypeptide=Potri.017G105501.1.p locus=Potri.017G105501 ID=Potri.017G105501.1.v4.1 annot-version=v4.1
ATGGCTTCCACCAGCGTGCAAGGAATAACCTCATCTTCATCTTCTTCTTCTTCTTCTCCTCCACTATATATGTATGATGTGTTCCTTAGTTTTAGAGGCA
AAGACACCCGGAACAACTTTACTAGTCATCTATATTATAATCTGGCACAGAGAGGCATCGATGTGTACATGGATGATAGGGGGCTCGAGAGAGGAAAGAC
CATCGAACCTGCTCTCTGGAAGGCCATCGAAGAATCAAGGTTTTCAGTCATTATATTTTCAAGAGAATACGCTTCTTCACCATGGTGCTTGGATGAACTT
GTCAAGATTGTTCAGTGCATGAAAGAGACGGGGCACACTGTTCTGCCAATTTTTTATGATGTGGATCCCTCAGAGGTAGCCGAGCAGAAAGGGCAATATG
AGAAAGCCTTCGTTGAGCATGAACAAAATTTCAAGGAAAACTTGGAGAAGGTCCGGAACTGGAAAGATTGTCTCTCTACGGTGGCCAATCTATCTGGCTG
GGACGTCCGGAATAGGTAA
AA sequence
>Potri.017G105501.1 pacid=42813192 polypeptide=Potri.017G105501.1.p locus=Potri.017G105501 ID=Potri.017G105501.1.v4.1 annot-version=v4.1
MASTSVQGITSSSSSSSSSPPLYMYDVFLSFRGKDTRNNFTSHLYYNLAQRGIDVYMDDRGLERGKTIEPALWKAIEESRFSVIIFSREYASSPWCLDEL
VKIVQCMKETGHTVLPIFYDVDPSEVAEQKGQYEKAFVEHEQNFKENLEKVRNWKDCLSTVANLSGWDVRNR
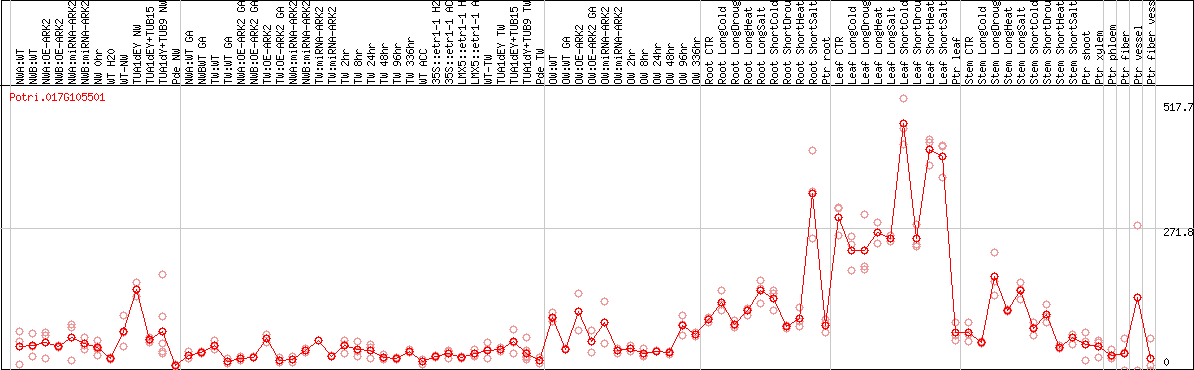
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G105501 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.