External link
Symbol
GF14.5
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G02520 468 / 4e-169
GENERALREGULATORYFACTOR7, GF14NU, GRF7
general regulatory factor 7 (.1)
AT5G38480 464 / 8e-168
RCI1, GRF3
general regulatory factor 3 (.1.2)
AT5G16050 444 / 1e-159
GF14UPSILON, GRF5
general regulatory factor 5 (.1)
AT1G78300 434 / 1e-155
14-3-3OMEGA, GF14OMEGA, GRF2
14-3-3 PROTEIN G-BOX FACTOR14 OMEGA, general regulatory factor 2 (.1)
AT1G35160 432 / 5e-155
14-3-3PHI, GF14PHI, GRF4 ,GF14 PHI
GENERAL REGULATORY FACTOR 4, 14-3-3 PROTEIN G-BOX FACTOR14 PHI, GF14 protein phi chain (.1.2)
AT4G09000 431 / 2e-154
GF14CHI, GRF1
GENERAL REGULATORY FACTOR1-G-BOX FACTOR 14-3-3 HOMOLOG ISOFORM CHI, general regulatory factor 1 (.1.2)
AT5G65430 389 / 2e-138
14-3-3KAPPA, GF14KAPPA, GRF8
14-3-3 PROTEIN G-BOX FACTOR14 KAPPA, general regulatory factor 8 (.1.2.3)
AT5G10450 387 / 1e-137
14-3-3lambda, AFT1, GF14LAMBDA, GRF6
14-3-3 PROTEIN G-BOX FACTOR14 LAMBDA, G-box regulating factor 6 (.1.2.3.4)
AT1G26480 358 / 6e-126
GF14IOTA, GRF12
general regulatory factor 12 (.1)
AT2G42590 349 / 2e-122
GENERALREGULATORYFACTOR9, GF14MU, GRF9
general regulatory factor 9 (.1.2.3)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033545
491 / 2e-178
AT3G02520 470 / 4e-170
general regulatory factor 7 (.1)
Lus10017584
487 / 7e-177
AT3G02520 471 / 1e-170
general regulatory factor 7 (.1)
Lus10036481
446 / 2e-160
AT1G78300 481 / 1e-174
14-3-3 PROTEIN G-BOX FACTOR14 OMEGA, general regulatory factor 2 (.1)
Lus10020940
439 / 5e-158
AT1G78300 483 / 2e-175
14-3-3 PROTEIN G-BOX FACTOR14 OMEGA, general regulatory factor 2 (.1)
Lus10008715
425 / 6e-152
AT1G78300 459 / 2e-165
14-3-3 PROTEIN G-BOX FACTOR14 OMEGA, general regulatory factor 2 (.1)
Lus10010349
419 / 7e-150
AT1G78300 454 / 5e-164
14-3-3 PROTEIN G-BOX FACTOR14 OMEGA, general regulatory factor 2 (.1)
Lus10035934
384 / 2e-136
AT5G65430 450 / 1e-162
14-3-3 PROTEIN G-BOX FACTOR14 KAPPA, general regulatory factor 8 (.1.2.3)
Lus10025727
380 / 2e-134
AT5G65430 454 / 3e-164
14-3-3 PROTEIN G-BOX FACTOR14 KAPPA, general regulatory factor 8 (.1.2.3)
Lus10040534
371 / 1e-130
AT5G65430 439 / 1e-157
14-3-3 PROTEIN G-BOX FACTOR14 KAPPA, general regulatory factor 8 (.1.2.3)
Lus10014899
369 / 6e-130
AT5G65430 438 / 2e-157
14-3-3 PROTEIN G-BOX FACTOR14 KAPPA, general regulatory factor 8 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00244
14-3-3
14-3-3 protein
Representative CDS sequence
>Potri.017G113300.2 pacid=42814559 polypeptide=Potri.017G113300.2.p locus=Potri.017G113300 ID=Potri.017G113300.2.v4.1 annot-version=v4.1
ATGATGTCTCCAACTGAACCATCACGTGAAGAGAGTGTTTACATGGCCAAGTTGGCTGAACAGGCCGAACGTTATGAAGAAATGGTGGAGTTTATGGAGA
AAGTTGCAAAGACAGTCGATAATGAGGAGCTAACCATGGAGGAAAGGAACTTGCTCTCCGTGGCCTACAAAAATGTGATTGGAGCTAGGAGGGCTTCATG
GAGGATCATCTCTTCCATTGAGCAGAAGGAAGAGAGTAGGGGAAATGAAGATCATGTCACAATCATCAAGGAGTACAGGGGAAAGATCGAAGCTGAGCTC
AGCAAGATCTGTGACGGAATCTTGAGCCTCCTTGAGACACATCTTGTTCCCTCCGCCTCAGCTGCTGAGTCCAAGGTATTTTACCTCAAGATGAAGGGTG
ATTATCACAGGTATCTTGCCGAGTTTAAGACCGGGGCTGAGAGGAAGGAAGCTGCTGAGAGCACTTTGTTGTCTTACAAGTCTGCTCAGGATATTGCTCT
TTCTGAACTGGCTCCTACCCACCCAATAAGGCTGGGGCTTGCACTTAACTTCTCTGTCTTCTACTATGAGATCCTTAACTCTCCTGATCGTGCTTGCAGT
CTTGCTAAGCAGGCTTTTGATGAGGCTATTTCCGAGCTGGATACTTTGGGTGAGGAGTCTTACAAGGACAGCACATTGATCATGCAACTTCTCCGTGACA
ATCTGACACTCTGGACTTCTGATATCACGGATGAGGCTGGGGATGAGATCAAGGACGCATCAAAACGTGAATCAGGCGATGGGCCGCAGTGA
AA sequence
>Potri.017G113300.2 pacid=42814559 polypeptide=Potri.017G113300.2.p locus=Potri.017G113300 ID=Potri.017G113300.2.v4.1 annot-version=v4.1
MMSPTEPSREESVYMAKLAEQAERYEEMVEFMEKVAKTVDNEELTMEERNLLSVAYKNVIGARRASWRIISSIEQKEESRGNEDHVTIIKEYRGKIEAEL
SKICDGILSLLETHLVPSASAAESKVFYLKMKGDYHRYLAEFKTGAERKEAAESTLLSYKSAQDIALSELAPTHPIRLGLALNFSVFYYEILNSPDRACS
LAKQAFDEAISELDTLGEESYKDSTLIMQLLRDNLTLWTSDITDEAGDEIKDASKRESGDGPQ
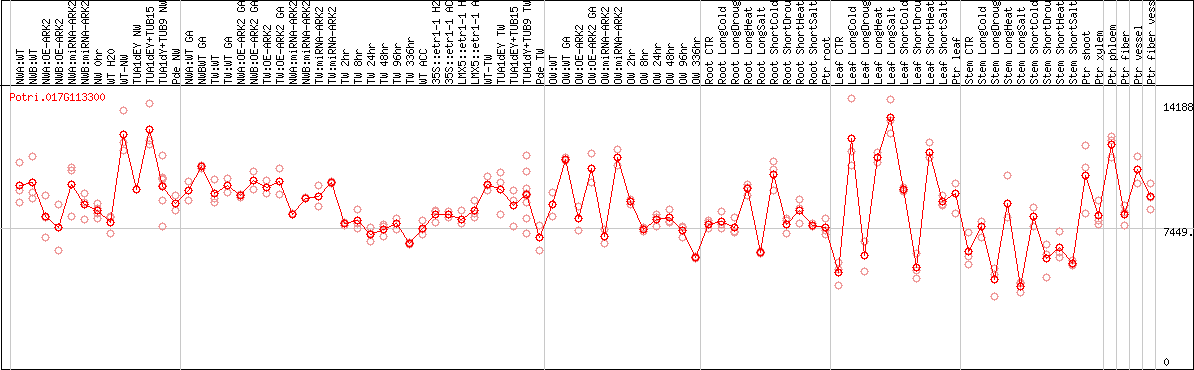
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G113300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.