Potri.017G140501 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.017G140501.1 pacid=42814264 polypeptide=Potri.017G140501.1.p locus=Potri.017G140501 ID=Potri.017G140501.1.v4.1 annot-version=v4.1
ATGATGGAGAAGACTACAGCGAAAACTGAAGAAGAAGTTGCTGGTAGAAGCTTATTAGATTTGGTGTTCTCTTGGTCTATCACGGATGTACTTAACAGAG
ATCTTTACAAAAATCAGGTGAAGAAGATACCAGAGACATTCACGTCAACGTCACATTACATGAAGTTATTCATTCCTGCTCTAATCGAGGAAACTCGTGC
AGATTTGTGCTCAAACATGATGAAGGTCTCCCAAGCACCTACAAGGGAGATCTTTTCAATTGAAAGATCAAAAGAGTATAAACCTCCCAAAGACGTATTT
TATAAGATATGGTTGAATAGAATGAGTAAAACTGGAAATGGGAAAGGAATATACGAGCCCGAGGTTGGACATCTCATTGCTTTGACAGATGCAAGACCGA
AAGACATTGCTGATTTGAACAGTCCTGGGATAAACTATCTTCTTGCTTATGTTCATGAGGTATCTAATGGGCTGGATGATGACAATAATCATGAAACGTT
ATCAATTCTCACATCCAAACCCATTCAATTTGAACTAGAAAACAAGCAGAATAAAAGGGAGTCTGTCATTGCTGGACAAGAGATTCAAAAGAAAAGCAGA
GCCACTTTCTTCGTTGTTTACCTAGCAAACATGACGACGAATGCCCGCATATGGAGATCATTGAACTCAGATCTGCAAGGAGGGAACACGAACGTTATCC
AGAATGTGCTTGAAACTAGTTCACCTGATAGCCAAGATTGTTCTCACTGCTTATCTGAAGTGAATAGAAGTGCTGCACTTTCTGGTATGGAAGAAACCAT
AATCAGCTCTTATAATTTAAATGAATCACAGGAAAATGCAATTGTAAGCTGCATCGGTCTGAGTGAATGCCAGCATCAAAGCACTGTCAAACTAATTTGG
GGTCCTCCAGGGACTGGAAAGACAAAGACAGTTGGTTTATTACTGTTTTCTCTTCTTAAATTGAAGTGCAGGACACTTACATGTTCTCCTACCAATATTG
CTGTGTTGCAAGTGACATCGGGACTCCTAAAGCTAGTTACAGACTCTCTTGAATATGACACCTATGGACTTGGAGATGTAGTTCTCTTTGGGAATGGGGG
GCGACTGAAGATTTCTGAGAATGATGATCTTGAAGATATATTTCTTGACCATCGTGTTGAAGTGCTTTATCTTTGCTTTGCCCCGTCAACCGGGTGGAAG
CACACTGTAGATTCAATGATAAATTTGCTTGAAGATCCTGAGCATCAGTACCGCAGGTACTTGGAAAACATGAAAAAAGAGAACGAGGGAGGGGATCGGG
ATGATGAAATGATTGAATTTCAAGAGATGAACAGCAACAAAGAGAAGGATGAAGTGGTCAGTGAACAAAACCAGAAGGGCAGGAACAGCAGGAAAGTTCT
GAAGAAAATTCTTTTACAGGCATTGAAAGACAACAAGAAAACTGAAAAAAAGAAACAGAAGGTTTCTTATCATCAAGATAAGCTTCCAAGGTGTTTGGGA
AAGGGAGATCAATATGGAAAGGAAAACAAGGAAGATAATATTCTGCCGTTCGAGGAATTTGTAAAGAAAAGGTTCAATATCCTTAGTGAGAAGCTGGACT
TTTTAATTTTTGGTTTGTATACGCATTTGCCAACGTCTGTCATTTCATTCGAAGTTGTGAAGAACATGATAAAAGCTCTTGATTCACTCAGCTGCCTCAA
AACATTGTTGAATGGTGTTAGCCTAGGAGATGGAGGTTTAGAGCTAGACATCAATGATTTTGAAAATGAAGAAAGCAGTGCCTGTCAATATTCAAGGTTG
GCCACTAAAAGAAAGGATTGCATTCAGATACTGAATTCACTTCCTCGATCATTTGACGTGCCCAATATTTTTGAAAGTTATCAAGTGAGAAACTTTTGCT
TGGAAAATGCATGTCTGATTTTCTGTACCGCTTCAAGCTCTGCCATGTTGCACACAGAAGGAATGAAACCGATAAAATTGTTGGTTGTTGATGAAGCTGC
GCAGCTTAAAGAATGTGAATCCACCATTCCATTGCAGCTTTCTGGTCTTCGCCATGCCGTTCTTATAGGTGACGAGCGCCAACTTCCTGCCATGGTTCAG
AGTCAGATTTCCGAGAAAGCTGAATTCGGGAGGAGCTTGTTCGAGAGATTGGTCATACTGGGACACGAGAAACACCTACTGAATATGCAGTACAGGATGC
ATCCATCTATAAGCTTATTCCCAAATAAAGAGTTCTATGGTGGGCTAATTCAGGATGCTTCAACTGTGAAAGAAAGAAACTATCAGAAACTATTTCTTCA
AGGAAATATGTATGGGCCTTACTCATTTATTAATGTAGCCAGTGGGAAAGAAGAATTCAATAACGGTGGCAGCAAGAAAAATTTGGTGGAGGTTGCTGTA
GTTTCAGAGCTAGTTGCAAGCCTTTTTAAAGAATTTACAAGGGCAAGAAAGAGGATGAGCGTAGGAGTCATATCACCGTACAACGCTCAAGTTTATGCAA
TTCAAGAAAAAATTGGAAAGACTTACAGTGCACATAGTGATTTTGCCGTAAATATTCGGTCTGTCGATGGATTTCAAGGAGGCGAGGAGGATGTTATAAT
TATCTCTACTGTGAGATGTAATGCAAATGGAAAAATAGGTTTCCTTGCCAATCGCCAAAGGGTGAATGTGGCATTGACCCGTGCAAGGCACTGCCTCTGG
ATATTGGGAAATGGAGCAACTCTTGTCAACAGCGATTCTATTTGGAAGAAATTAGTTACTGATGCCAAGGAACGGGGGTGTTTCTATAATGCCGAAGAGG
GTAAAAGCTTGTCTAAGGCTATTACGGATGACTTTTTAGAATCGGACCAACTTGATGCTCTGCTGAATGTGAATTCTCCACTGTTCAGAAATGCAAGATG
GAAGTTTTGCTTCAGTAACGATTTTCGGAAATCCATTCTGAAAGTGAGAAATGAGGCTCGTCAGGAAGTGTTTTCTTTGTTGTCAAAGCTTTCGAGTGGC
TGGCGGGAATCTCCTGAAGAGAGAATCATCGTTGTCCGACATGGAACTTCTTCTGAACTGTTAGAACAATACAGAGTCAATGATCAGCTTAAACTGATTT
GGACAGTGGATATAATCAAGGAAAACTCAAACCACACACAAATTCTCAAGGTGTGGGATGTTTTACCATCTCCTGATTTACCAAAACTAGCAAGGCATCT
CGATGATGTCTTTGGGAATTATACAGTTGATAAGATGAATCGCTGCAAGCACAAATTTATAGAAGGGAATTTGGTTGTTCCAATGAGATGGCCACTGTAT
TTCGATGGTGCTGCCGAGCGTAGTATCCCTGAAATTGATCCTGTAGAACTTCTGTCGCAACCTTCGGCTTCACTTATGGCGCGGACACCAAAAGCGAAGA
CACCAAATCCACATCGCAGATTACCTAAGTATATGAATAAAAACAACAAAAAATATCGCAGCACATCACTTTTTGGATGGGATAATGAACCCATATGGGT
CTATTTCATCATTTTTATTTTGATTATTTTAATTATTTATGTTCTTCAATTTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.017G140501.1 pacid=42814264 polypeptide=Potri.017G140501.1.p locus=Potri.017G140501 ID=Potri.017G140501.1.v4.1 annot-version=v4.1
MMEKTTAKTEEEVAGRSLLDLVFSWSITDVLNRDLYKNQVKKIPETFTSTSHYMKLFIPALIEETRADLCSNMMKVSQAPTREIFSIERSKEYKPPKDVF
YKIWLNRMSKTGNGKGIYEPEVGHLIALTDARPKDIADLNSPGINYLLAYVHEVSNGLDDDNNHETLSILTSKPIQFELENKQNKRESVIAGQEIQKKSR
ATFFVVYLANMTTNARIWRSLNSDLQGGNTNVIQNVLETSSPDSQDCSHCLSEVNRSAALSGMEETIISSYNLNESQENAIVSCIGLSECQHQSTVKLIW
GPPGTGKTKTVGLLLFSLLKLKCRTLTCSPTNIAVLQVTSGLLKLVTDSLEYDTYGLGDVVLFGNGGRLKISENDDLEDIFLDHRVEVLYLCFAPSTGWK
HTVDSMINLLEDPEHQYRRYLENMKKENEGGDRDDEMIEFQEMNSNKEKDEVVSEQNQKGRNSRKVLKKILLQALKDNKKTEKKKQKVSYHQDKLPRCLG
KGDQYGKENKEDNILPFEEFVKKRFNILSEKLDFLIFGLYTHLPTSVISFEVVKNMIKALDSLSCLKTLLNGVSLGDGGLELDINDFENEESSACQYSRL
ATKRKDCIQILNSLPRSFDVPNIFESYQVRNFCLENACLIFCTASSSAMLHTEGMKPIKLLVVDEAAQLKECESTIPLQLSGLRHAVLIGDERQLPAMVQ
SQISEKAEFGRSLFERLVILGHEKHLLNMQYRMHPSISLFPNKEFYGGLIQDASTVKERNYQKLFLQGNMYGPYSFINVASGKEEFNNGGSKKNLVEVAV
VSELVASLFKEFTRARKRMSVGVISPYNAQVYAIQEKIGKTYSAHSDFAVNIRSVDGFQGGEEDVIIISTVRCNANGKIGFLANRQRVNVALTRARHCLW
ILGNGATLVNSDSIWKKLVTDAKERGCFYNAEEGKSLSKAITDDFLESDQLDALLNVNSPLFRNARWKFCFSNDFRKSILKVRNEARQEVFSLLSKLSSG
WRESPEERIIVVRHGTSSELLEQYRVNDQLKLIWTVDIIKENSNHTQILKVWDVLPSPDLPKLARHLDDVFGNYTVDKMNRCKHKFIEGNLVVPMRWPLY
FDGAAERSIPEIDPVELLSQPSASLMARTPKAKTPNPHRRLPKYMNKNNKKYRSTSLFGWDNEPIWVYFIIFILIILIIYVLQF
|
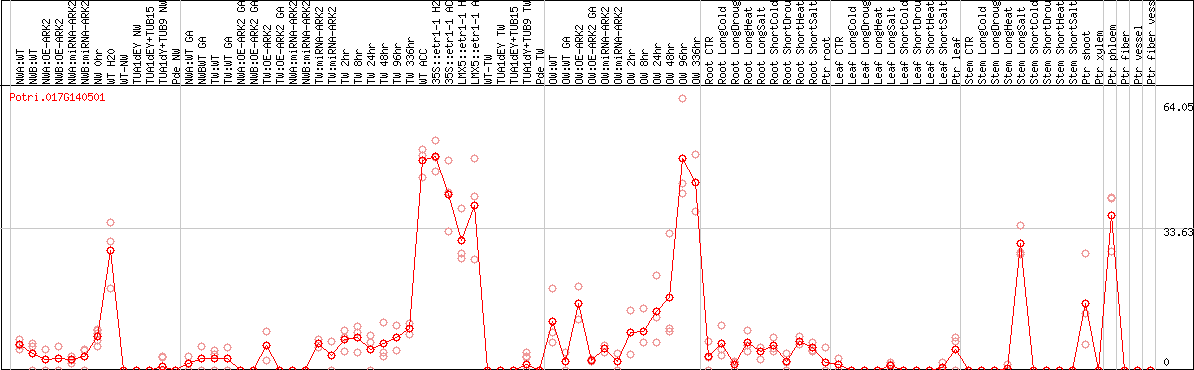
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.017G140501 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
