PKL.1 (Potri.018G021100) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | PKL.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G021100.2 pacid=42801776 polypeptide=Potri.018G021100.2.p locus=Potri.018G021100 ID=Potri.018G021100.2.v4.1 annot-version=v4.1
ATGACTACCCCTTTTAAAACAGAACAATGTAATAAGATGAATAGTAAGGAAATGAGTAGCTTAGTGGAGAGGCTACGTGTTAGATCAGAAAGGAGGCCAA
TTTATAACCTGGATGAATCAGATGATGATGCTGATTTTGTTTCTGGGAAAGCTAAAAAACCCCAGGAGAAAATTGAAAGATTTGTCAGGGATGATGCGAA
GGAGGATTCCTGTCAAGCTTGCGGAGAGAGTGAGAATCTTCTGAATTGTGAAACTTGCACTTATGCTTACCATCCCAAGTGTCTGCTTCCACCACTAAAA
GCTCCTTTTCCTAGCAATTGGAGATGCCCTGAATGTGTTAGCCCTCTGAATGATATTGATAAGTTATTGGACACTGAAATGCGCCCTACCGTAGCTGATG
ATAGTGATGCCTCTAAGCTAGGCTCAAAGCAAATTTTTGTAAAACAGTATCTTGTGAAGTGGAAAGGCTTATCTTATCTACATTGCACTTGGGTACCTGA
GAGAGAGTTTCTAAAAGCTTTCAAGTCAAATCCTCGTCTGAAGACTAAAGTAAACAATTTCAATCGTCAAATGGCGTCGAACAATAACTCTGAGGATGAC
TTTGTTGCTATCCGACCTGAATGGACAACTGTTGATCGAATTCTTGCTTGCAGGGGAGTTGAGGGTGAGAAGGAATATCTTGTGAAGTACAAGGAACTTC
CTTATGATGAATGTTATTGGGAATTTGAATCAGATGTATCTACATTCCAGCCTGAAATAGAAAGATTTAATAGAATTCAGTCTAGGTCTCACAAACCGAG
TAAGCAAAAGAGCAGCCTTCAAGATGCTACTGATTCAAAGAAGAAATCAAAAGAATTTCAACAGTATGAACACAGTCCTGAATTTCTTTCTGGAGGCTCA
TTGCATCCATATCAGCTAGAAGGGCTGAACTTCTTACGTTTTTCTTGGTCCAAACAAACACATGTAATACTTGCTGATGAGATGGGCCTTGGCAAAACTA
TACAGAGTATTGCATTTTTAGCATCTCTTTTTGAAGAAGGCATCTCTCATCATTTGGTAGTAGCTCCACTGTCAACATTGCGAAACTGGGAACGTGAATT
TGCAACATGGGCGCCTCAAATGAATGTTGTTATGTATGTTGGCTCTGCACAAGCTCGTGCTGTCATTAGGGAATATGAATTTTACTACCCCAAGAAGCAC
AAGAAGATCAAGAAAAAGAAATCTGGTCAAGTTGTCACTGAAAGAAAGCAAGATAGAATAAAATTTGATGTGCTTCTAACGTCTTATGAGATGATTAACT
TAGACACAACATCGCTGAAACCAATAAAGTGGGAGTGCATGATTGTTGATGAAGGTCATCGTCTAAAAAATAAGGACTCAAAATTGTTTCTCTCCATGAA
GCAGTACTATAGTAACCATCGTGTGCTTTTGACTGGAACCCCTCTTCAGAACAATCTGGATGAACTTTTCATGCTCATGCATTTCCTTGATGCTGGGAAG
TTTGCGAGTTTAGAAGAGTTCCAAGAGGAGTTTAAGGATATAAATCAAGAGGAGCAGATATCGAGACTTCATAAAATGTTGGCTCCCCATCTCCTTAGAA
GGGTGAAGAAGGATGTCATGAAGGAGCTACCTCCAAAGAAGGAACTCATTCTACGTGTTGAACTAAGTAGTAAGCAGAAGGAATATTACAAGGCCATTCT
GACTCGTAACTATCAAATACTGACTCGACGTGGCGGTGCACAGATTTCTCTTATCAATGTGGTTATGGAATTGCGCAAACTCTGTTGTCATCCTTATATG
TTAGAAGGAGTGGAACCAGATATAGAAGACACCAATGAATCTTTCAAGCAGCTCGTGGAAACTTCTGGGAAATTGCAACTGTTGCACAAAATGATGGTGA
GACTTAAAGAACAAGGACATAGAGTTCTCATTTACTCACAATTTCAGCACATGCTGGACCTACTGGAAGATTACTGCACTCATAAGAAATGGACATATGA
ACGGATAGATGGAAAGGTTGGTGGAGCTGAGAGACAAATACGCATAGATCGATTTAATGCCAAAAATTCTTCAAGATTTTGCTTTCTTCTCTCAACTAGA
GCTGGAGGACTGGGAATAAATCTTGCAACTGCTGATACAGTTATTATATATGATAGTGATTGGAATCCTCATGCTGATCTTCAAGCAATGGCTAGGGCAC
ATCGACTTGGGCAAACCAACAAGGTAATGATTTATAGGCTCATAACCCGAGGAACTATTGAGGAACGGATGATGCAGATGACTAAGAAGAAAATGGTTTT
AGAGCACCTTGTTGTTGGGAGGCTGAAGGCACAAAATATTAACCAAGAAGAGCTAGATGACATCATAAGATATGGTTCAAAGGAGCTTTTTGCTGATGAG
AATGATGAAGCAGGAAAATCACGTCAAATTCACTATGACGATGCTGCTATACAGCGATTACTTGACCGCGAACAAATTGGAGATGAAGAAACGTCATTGG
ATGATGAAGAGGAAGATGGATTTTTAAAGGCATTTAAGGTTGCAAATTTTGAGTATATTGACGAAGCAGAGGCTGCAGCAGAGAAGGAAGCTCAAAAGGC
AGCAATGGAAACTAAAACTACCATAAGCAATTCCGAAAAAACAAACTATTGGGAGGACTTGCTAAAAGACAGTTATGAAGTGCACAAAATCGAGGAGTCT
AATGCTTTGGGCAAAGGAAAGAGAAGCCGTAAGCAGATGGTCTCTGTTGAAGAAGATGACCTTGCTGGTCTAGAAGATGTAAGCTCTGATGGCGAGGATG
ATAACTATGAAGCTGAGCTGACTGATGGTGAAACAACTTCATCTGGAATTCAGACATCTGGAATTCAGACATTAAAAAGGCCTTATAAGAAGAAGGGTCG
AGTGGATAATATGGAGCCAATTCCTCTGATGGAAGGTGAAGGTAGATCATTTAGAGTATTGGGTTTTAATCAAAATCAGAGGGCTGCATTTGTACAAATT
TTGATGAGGTATGGGGTCGGAGACTATGACTGGAAGGAGTTTGCCCCTCGTTTGAAACAGAAGACATATGAAGAAGTAGAAAATTATGGAAGATTATTCT
TGACACATATAGCAGAAGATCTATCTGACTCCCCTAATTTTTCAGATGGTGTTCCTAAAGAGGGTCTTAGAATACAAGATGTACTTATTAGGATTGCTGT
CCTGCTTTTGATAAGGGATAAGGCAAGGTTTGCATCAGAGAACCCAGGCAGCCTCCTTTACACAGATGACATTATGGTCCGCTACCCAGGCCTGAAGAGC
GGAAAATTTTGGAAACAAGAGCATGATTCGTTGTTACTTCATGCGGTATTGAAGCATGGGTATGGAAGATGGCAAGCTATTGTTGATGACAAGGACTTAA
AGGTTCAAGAAATCATTTGTAAAGAGTTGAATCTTCCTTTTATAAGACTACCTGTCCTTGGACAAGCTGCTTCTCAAGCACAAAACGGTTCTACTTCTAA
CATGGATAATGCAGAAGCTCCTAGCACCCAGACACAAGCAAATGGTACTGGAAATGTAGCAGCAGCTGATGTTGCTCATGGAACAACTGATGTTGCAAAT
CAAGCACAGTTATATCAAGATTCATCTATCTTATTTCATTTTAGAGATATGCAGAGACGACAGGTAGAGTTCATTAAGAAAAGAGTGCTTCTTTTGGAGA
GAGGACTTTACGCAGAGTACCAGAAAGAATACTTTGGTGGTGATATAAAAGCAAATGAGATTACAAGTGAAGAGGCTGATTGTGAAACAATGGCGGCTGA
CAGATCAAGTTTAGGCTCCATAGAGATTAGTGCTCAAATGATTGACCAGTTGCCTCGAATGGAATCAATTGCATTAGAAGAAATTTCAGCAGCTGCTTGT
GATGACAATCCCGATAGATTGGCATTACCTCAGCTTTATAACAAGATGTGCACGGTGTTGGAGCAAAACATTCATGAATCAATTCAGATATCCTTGACCA
ACCAACCTGCAAGTCTCAAGTTGAGACAGGATCTTCAGCCATTAGAGACGGTTTATGAACAGATAAATCAATTTCTCTCTCCTTCACAACAGAAATCTTC
TACTTCAGAACAGGCTACTCTTGGTTCAAGCAAACATGTCCAAGCTGAATCACAGAGCAGTCAGGCAGATTTCCATTCACCATCTGACCAGCTGAAGGAG
AATGATGACACCACTGCTGCTACCGAAGTTGTTGAGATGAAAGATGCAACAACAGAGCCAAAATTGCAAGGAACCATTGCATTATCAAACGAAGAACTAG
TTAAAGAAACAAGTAAATCTCCTAGTGATTCTCCACCTTCTGCATGCCCAGTTACTCCTCCAAAAGAACCAACTTGTTCACCAGGGACAACTGAAAAGGA
TGTGGGAATGGTCGATATGAATAATGAAAATGACACCCAACACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G021100.2 pacid=42801776 polypeptide=Potri.018G021100.2.p locus=Potri.018G021100 ID=Potri.018G021100.2.v4.1 annot-version=v4.1
MTTPFKTEQCNKMNSKEMSSLVERLRVRSERRPIYNLDESDDDADFVSGKAKKPQEKIERFVRDDAKEDSCQACGESENLLNCETCTYAYHPKCLLPPLK
APFPSNWRCPECVSPLNDIDKLLDTEMRPTVADDSDASKLGSKQIFVKQYLVKWKGLSYLHCTWVPEREFLKAFKSNPRLKTKVNNFNRQMASNNNSEDD
FVAIRPEWTTVDRILACRGVEGEKEYLVKYKELPYDECYWEFESDVSTFQPEIERFNRIQSRSHKPSKQKSSLQDATDSKKKSKEFQQYEHSPEFLSGGS
LHPYQLEGLNFLRFSWSKQTHVILADEMGLGKTIQSIAFLASLFEEGISHHLVVAPLSTLRNWEREFATWAPQMNVVMYVGSAQARAVIREYEFYYPKKH
KKIKKKKSGQVVTERKQDRIKFDVLLTSYEMINLDTTSLKPIKWECMIVDEGHRLKNKDSKLFLSMKQYYSNHRVLLTGTPLQNNLDELFMLMHFLDAGK
FASLEEFQEEFKDINQEEQISRLHKMLAPHLLRRVKKDVMKELPPKKELILRVELSSKQKEYYKAILTRNYQILTRRGGAQISLINVVMELRKLCCHPYM
LEGVEPDIEDTNESFKQLVETSGKLQLLHKMMVRLKEQGHRVLIYSQFQHMLDLLEDYCTHKKWTYERIDGKVGGAERQIRIDRFNAKNSSRFCFLLSTR
AGGLGINLATADTVIIYDSDWNPHADLQAMARAHRLGQTNKVMIYRLITRGTIEERMMQMTKKKMVLEHLVVGRLKAQNINQEELDDIIRYGSKELFADE
NDEAGKSRQIHYDDAAIQRLLDREQIGDEETSLDDEEEDGFLKAFKVANFEYIDEAEAAAEKEAQKAAMETKTTISNSEKTNYWEDLLKDSYEVHKIEES
NALGKGKRSRKQMVSVEEDDLAGLEDVSSDGEDDNYEAELTDGETTSSGIQTSGIQTLKRPYKKKGRVDNMEPIPLMEGEGRSFRVLGFNQNQRAAFVQI
LMRYGVGDYDWKEFAPRLKQKTYEEVENYGRLFLTHIAEDLSDSPNFSDGVPKEGLRIQDVLIRIAVLLLIRDKARFASENPGSLLYTDDIMVRYPGLKS
GKFWKQEHDSLLLHAVLKHGYGRWQAIVDDKDLKVQEIICKELNLPFIRLPVLGQAASQAQNGSTSNMDNAEAPSTQTQANGTGNVAAADVAHGTTDVAN
QAQLYQDSSILFHFRDMQRRQVEFIKKRVLLLERGLYAEYQKEYFGGDIKANEITSEEADCETMAADRSSLGSIEISAQMIDQLPRMESIALEEISAAAC
DDNPDRLALPQLYNKMCTVLEQNIHESIQISLTNQPASLKLRQDLQPLETVYEQINQFLSPSQQKSSTSEQATLGSSKHVQAESQSSQADFHSPSDQLKE
NDDTTAATEVVEMKDATTEPKLQGTIALSNEELVKETSKSPSDSPPSACPVTPPKEPTCSPGTTEKDVGMVDMNNENDTQH
|
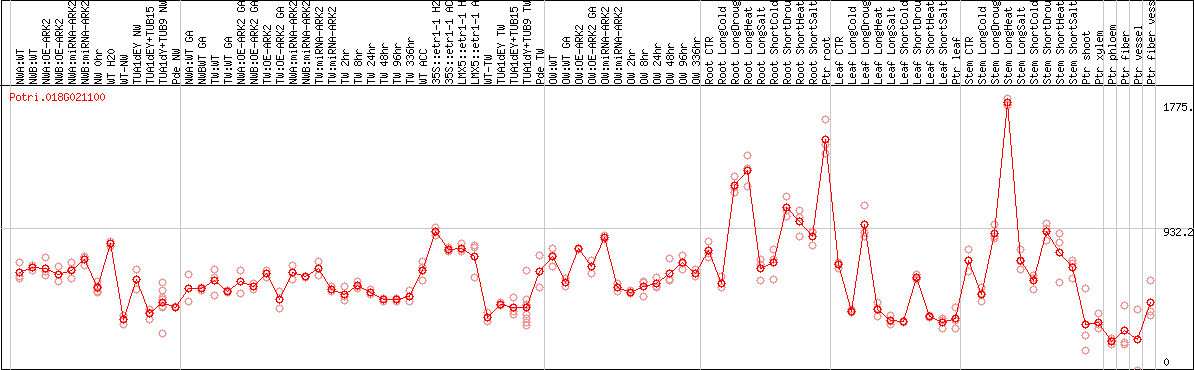
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G021100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
