External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G26660 321 / 2e-110
ATSPX2
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
AT5G20150 313 / 5e-108
ATSPX1
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
AT5G15330 200 / 1e-62
ATSPX4
ARABIDOPSIS THALIANA SPX DOMAIN GENE 4, SPX domain gene 4 (.1)
AT2G45130 177 / 1e-54
ATSPX3
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
AT4G22990 45 / 4e-05
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
AT4G11810 44 / 8e-05
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G253400
467 / 3e-168
AT5G20150 308 / 1e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.018G131500
330 / 9e-114
AT5G20150 356 / 1e-124
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.006G069500
325 / 1e-111
AT5G20150 359 / 8e-126
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.002G143900
209 / 5e-67
AT2G45130 276 / 1e-93
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
Potri.014G061400
207 / 3e-66
AT2G45130 269 / 4e-91
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
Potri.001G193200
175 / 2e-54
AT5G20150 158 / 2e-48
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.017G086600
172 / 1e-51
AT5G15330 335 / 4e-115
ARABIDOPSIS THALIANA SPX DOMAIN GENE 4, SPX domain gene 4 (.1)
Potri.001G135950
88 / 1e-21
AT2G45130 121 / 7e-35
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
Potri.003G120600
44 / 8e-05
AT4G22990 1112 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002018
400 / 9e-142
AT2G26660 308 / 2e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
Lus10002908
398 / 6e-141
AT2G26660 309 / 1e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
Lus10043272
330 / 6e-114
AT5G20150 343 / 8e-120
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10030095
317 / 5e-109
AT5G20150 356 / 8e-125
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10019415
315 / 4e-106
AT5G20150 326 / 2e-110
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10009314
53 / 1e-07
AT4G22990 941 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Lus10015852
45 / 5e-05
AT4G22990 926 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Lus10006340
43 / 0.0002
AT4G22990 1128 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03105
SPX
SPX domain
Representative CDS sequence
>Potri.018G028200.1 pacid=42800807 polypeptide=Potri.018G028200.1.p locus=Potri.018G028200 ID=Potri.018G028200.1.v4.1 annot-version=v4.1
ATGAAGTTCTGGAAGAGCTTAAGTAATTTAATGGAGGAAACGTTGCCGGATTGGAGAGATAAGTTCTTATCTTATAAAGATTTGAAGAAACAATTGAAGT
TAATTTATCCTAAGGAGCGTGATAAGCCACTGAATAAACGGCCTAGATTGGACGATGATCAAATGGACAGTGGTGAGGCGGAGAAGGAGGTAATTGATTT
TGTTAGGGTTTTGGAAGATGAGATGGAGAAGTTTAATTCCTTTATTGTCGAGAAAGAAGAGGATTATGTTATCAAATGGAAGGAGCTACAAGATAGAGCA
GAAAAGGCGAAGGATTCAAATGAAGAGCTGATGAAAGTAGGGAGGGAGATAGTGGATTTTCATGGAGAGATGGTGTTGTTGGAGAATTACAGTGCCCTTA
ACTATACAGGACTAGTGAAGATACTGAAGAAATATGATAAGAGAACTGGAGCTCTGGTCCGCATGCCCTTCATCCAAAGGATTATGCAACAGCCATTCTA
CACAACTCATGTACTTAACAAGCTTATTAAGGAGTGTGAGACGATTCTGGATTATATTTTCTCTAGAAAAGAACCATCAGTCTCACCTCAAATAACTGAT
GAGATAAGCGGCCTTGATACCAAAACTTCAACGGAAAGTTCAGAGAGGTCGCTCAGAGTTCCCAGCGAACTCCCAGAGATAGAGTATATGGAAAGCATGT
ACGTGAAGCTAACACTGTCAGCACTTCGCGTCTTGAAGGATGTTCGGAGTGGAAGCTCAACTGTAAGCGTGTATTCATTGCCCCCTTTGCAAATTAATAC
ACAGGAGGGGGATTGGAAGAAAGTTAATGTCTTGGAACAAGCAGCCAAATAG
AA sequence
>Potri.018G028200.1 pacid=42800807 polypeptide=Potri.018G028200.1.p locus=Potri.018G028200 ID=Potri.018G028200.1.v4.1 annot-version=v4.1
MKFWKSLSNLMEETLPDWRDKFLSYKDLKKQLKLIYPKERDKPLNKRPRLDDDQMDSGEAEKEVIDFVRVLEDEMEKFNSFIVEKEEDYVIKWKELQDRA
EKAKDSNEELMKVGREIVDFHGEMVLLENYSALNYTGLVKILKKYDKRTGALVRMPFIQRIMQQPFYTTHVLNKLIKECETILDYIFSRKEPSVSPQITD
EISGLDTKTSTESSERSLRVPSELPEIEYMESMYVKLTLSALRVLKDVRSGSSTVSVYSLPPLQINTQEGDWKKVNVLEQAAK
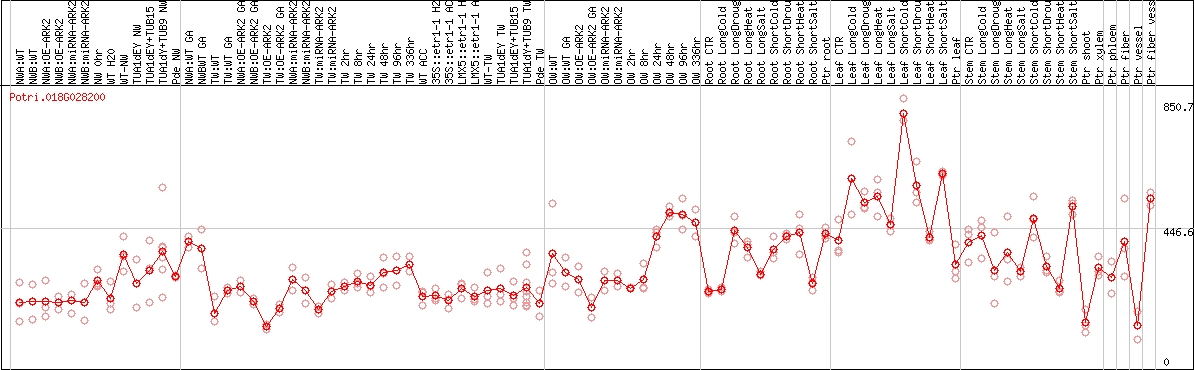
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G028200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.