Potri.018G031600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G031600.1 pacid=42801897 polypeptide=Potri.018G031600.1.p locus=Potri.018G031600 ID=Potri.018G031600.1.v4.1 annot-version=v4.1
ATGACGAAGAAGAGGGTGTTGTTCTGTGGACAAGGGGGGCTGGAAAATGGAAGCCTTTACGGAAGTGAACAAGGGAATAATGGAATACGTGACAAAGAAC
ATGATGAATTATCAGGAGACGATGCCAGCCATTTTAGCCTTACCGGTGAGATTCTCCCTTCTCTAGGTGCCACCGCTCGTAGCAACCGCCGTGTCATTCT
CCGTCGCTACATTCTCTCTCCCTTCGCCCCTAACTACAGATTATGGGATACTTTCTTGGTGTTTCTAGTTTTCTACACGGCATGGGTGTCTCCGTTTGAG
TTTGGTTACCTTAGCAAACCTAGTGGGGGTCTGGCCATAACAGATAATGTTGTCAATGGATTTTTTGCCATTGATATTGTTTTGACCTTCTTTGTTGCAT
ACCTTGACAAAAGTTCATATCTACTAGTTGATAACCGAAAAAAGATTGCTTGGAGGTATACAAAATCGTGGTTGGTCCTTGATGTTATCTCCACAATCCC
ATCTGAGCTTATCAGGGAAATATTGCCTGATAAACTTCAGTCATACGGGTACTTTAGTATGCTTCGCCTCTGGCGTCTCCGGCGAGTTAGTAAATTCTTT
TCAAGATTGGAAAAGGATAGGAACTACAATTACTTTGTGGTTCGATGCGTGAAGCTCATATGTGTTACTCTATTTGTAGTTCACATGGCCGGTTGCTTCT
ACTATCGCATTGCTGTAAATTACAAGGATCCAAGTAAAACATGGATCGGATCCGTTTGGGAAGATTTTCATGGAGAGAGCCTGTGGATTCGTTATGTCAA
ATCACTTTACTGGTCTACAACCACGCTCTCTACCACTGGATATGGTGATTTGCATGCTGTCAATCCGCAGGAGATGATATTCGTCATGTTCTACATGATG
TTTAATCTTGGACTGACATCGTATTTGATTGGAAACATGACCAATTTGGTTGTCCATGCCACCTTCCGAACAAGGCAATTTAGAGATACTATCCAAGCTG
TCTCAAGTTTTGCACAAAGGAACCGCCTTCCGATTCGGTTACAAGAACAGATGCTTGCTCATTTGTGTTTGAAGTACAGAACCGACTCGGAGGGATTGCA
TCAACAAGAGACCATTGGTTCTCTTCCTAAAGCAATTCGATCGAGCATTTCTAACTATCTCTTTTATTCACTTGTGGATAAAGTGTACTTGTTTCGTGGG
GTATCAAATGACCTGCTCTTTCAGTTGGTCACAGAGATGAAAGCTGAGTACTTTCCACCCAGAGAAGATGTAATTTTGCAAAATGAAGCACCAACAGATC
TGTATATTTTAGTCACTGGCGCGGTGGAACTTATCGTGCACAGGAATGGAATAGAGCAGGTTGTTGGTGAGGCTGCAACTGGAGATGTTATTGGTGAGAT
TGGTCTGCTATGTTACAGACCACAGTTATTCACGGTTCGAACCAAACGGTTGAGTCAGCTCCTACGTTTGAATCGAACTGCGTTCTTAAATAATGTCCAA
TCAAATGTTGGAGATGGGACAGTAATCATGAACAACCTTCTTCAGCATTTGAAAGAGCTGAATGATCCAGAGATGGAAGGAATTTTGCACCACACAGAGC
ACATGCTGAATCAGGACAGAATGGATTTACCTCTTACTTTGTGCATCGCAGCAATGAGGGGTGATGATCTATTGTTGCATCAGTTGCTGAAGCAGGGTTC
AGATCCAAATGAGTCGGACGAAAATGGACGAACTGCGCTGCATATTGCAGCTTCAAATGGGAATGAGCACTGTGTAGTCCTACTTCTGGAGTATGGAGTG
GATCCTAACATCAAAGATTCAGAGGGCAATGTTCCTCTATGGGAAGCGCTACAGGGGAATCATAAATCTGTATTTAAACTTCTTTCAGACAATGGGGCAA
CTATAACTTCTGGTGATGTGGGCCAATTTGCATATACAGCTGCTGAGCAAAATAACTTGGATTTGCTCAAGGAAATTGTCAAATATGGTGGAGATGTAAC
ACTGCCTGCAAGGTGTGGAACTACTGCCATTCATACAGCAATCTCTGAAGGAAACACTGAAATGGTCAAGTTCATCTTAGACCAAGGAGCCGATGTTGAT
AAGCCAGATCTCCATGGATGGACACCAAGGGCCTTGGCTGATCATCAGGGCCAAGAGGAAATTCAAGCTTTGTTCGAAAATAGGATGCAGACGAATAAAA
AAACGGTTTCCACAATCCCAAAGCATCCAGGGGTGCCATTTGGTAGGAAGCCCATGGCAAGGTATAATAGTGAACCAACAATACCTCCATTTTCTCCTTC
TTTTCGTCATGATGTTATGCCACCAGTCCCTGAAGTCTCATGGCCTGATAGACCTCGAAGGAGAAGGGCAGATAATTTCCACAACTCTCTTGTTGGGATG
ATGTCAGTTGCTAGTACTGGTGAGAATGACATAATTTCATCTCCAGCCCGTTTTACTGGCTTCGCAAGTTTGAACTGTCGTGCTAGAGTGACTCTCAGTT
GCCCGGATAAAGGTGAAGTTGCTGGAAAGATTGTTGCTTTACCAAATTCACTCCAAGAACTACTTGATATTGGTTCCAAAAAATTTGGGTGCAATGCCTC
CAAGATTCTAACCAAGGAAGGTGCTGAAATTGAAGATATAGAGGTACTTAGAGATGGCGATCATCTTGTTCTTGTTAGTGATGCTGGCACTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G031600.1 pacid=42801897 polypeptide=Potri.018G031600.1.p locus=Potri.018G031600 ID=Potri.018G031600.1.v4.1 annot-version=v4.1
MTKKRVLFCGQGGLENGSLYGSEQGNNGIRDKEHDELSGDDASHFSLTGEILPSLGATARSNRRVILRRYILSPFAPNYRLWDTFLVFLVFYTAWVSPFE
FGYLSKPSGGLAITDNVVNGFFAIDIVLTFFVAYLDKSSYLLVDNRKKIAWRYTKSWLVLDVISTIPSELIREILPDKLQSYGYFSMLRLWRLRRVSKFF
SRLEKDRNYNYFVVRCVKLICVTLFVVHMAGCFYYRIAVNYKDPSKTWIGSVWEDFHGESLWIRYVKSLYWSTTTLSTTGYGDLHAVNPQEMIFVMFYMM
FNLGLTSYLIGNMTNLVVHATFRTRQFRDTIQAVSSFAQRNRLPIRLQEQMLAHLCLKYRTDSEGLHQQETIGSLPKAIRSSISNYLFYSLVDKVYLFRG
VSNDLLFQLVTEMKAEYFPPREDVILQNEAPTDLYILVTGAVELIVHRNGIEQVVGEAATGDVIGEIGLLCYRPQLFTVRTKRLSQLLRLNRTAFLNNVQ
SNVGDGTVIMNNLLQHLKELNDPEMEGILHHTEHMLNQDRMDLPLTLCIAAMRGDDLLLHQLLKQGSDPNESDENGRTALHIAASNGNEHCVVLLLEYGV
DPNIKDSEGNVPLWEALQGNHKSVFKLLSDNGATITSGDVGQFAYTAAEQNNLDLLKEIVKYGGDVTLPARCGTTAIHTAISEGNTEMVKFILDQGADVD
KPDLHGWTPRALADHQGQEEIQALFENRMQTNKKTVSTIPKHPGVPFGRKPMARYNSEPTIPPFSPSFRHDVMPPVPEVSWPDRPRRRRADNFHNSLVGM
MSVASTGENDIISSPARFTGFASLNCRARVTLSCPDKGEVAGKIVALPNSLQELLDIGSKKFGCNASKILTKEGAEIEDIEVLRDGDHLVLVSDAGT
|
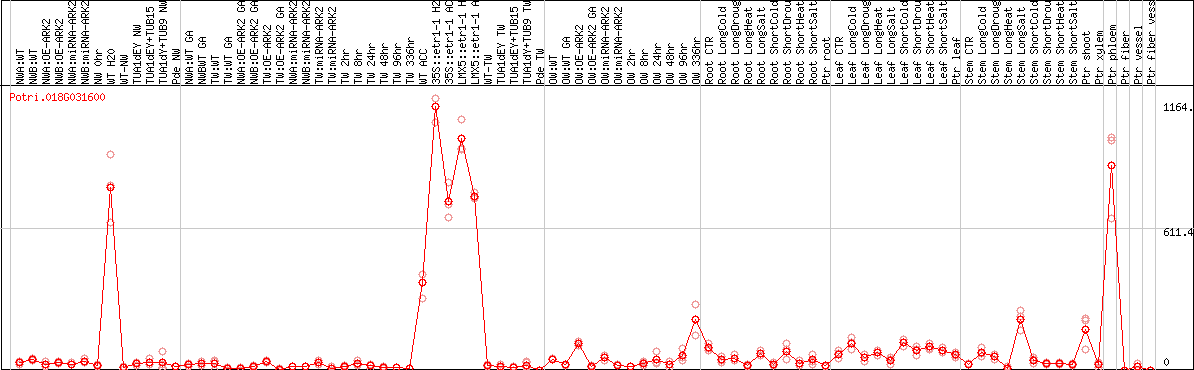
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G031600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
