External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G20900 122 / 9e-35
ZIM
TIFY3B, JAZ12
jasmonate-zim-domain protein 12 (.1)
AT3G43440 89 / 2e-21
ZIM
TIFY3A, JAZ11
jasmonate-zim-domain protein 11 (.1.2)
AT1G74950 74 / 6e-16
ZIM
TIFY10B, JAZ2
JASMONATE-ZIM-DOMAIN PROTEIN 2, TIFY domain/Divergent CCT motif family protein (.1)
AT1G72450 66 / 7e-13
ZIM
TIFY11B, JAZ6
TIFY DOMAIN PROTEIN 11B, jasmonate-zim-domain protein 6 (.1)
AT5G13220 64 / 2e-12
ZIM
JAS1, TIFY9, JAZ10, AT5G13220
TIFY DOMAIN PROTEIN 9, JASMONATE-ASSOCIATED 1, jasmonate-zim-domain protein 10 (.1.2.3.4)
AT1G19180 63 / 4e-12
ZIM
TIFY10A, JAZ1
jasmonate-zim-domain protein 1 (.1.2)
AT1G17380 62 / 1e-11
ZIM
TIFY11A, JAZ5
jasmonate-zim-domain protein 5 (.1)
AT4G14713 61 / 5e-11
ZIM
TIFY4A, PPD1
PEAPOD 1, TIFY domain/Divergent CCT motif family protein (.1.2)
AT1G70700 57 / 1e-09
ZIM
TIFY7, JAZ9
JASMONATE-ZIM-DOMAIN PROTEIN 9, TIFY domain/Divergent CCT motif family protein (.1.2)
AT3G17860 56 / 3e-09
ZIM
TIFY6B, JAI3, JAZ3
JASMONATE-INSENSITIVE 3, jasmonate-zim-domain protein 3 (.1.2.3)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011929
122 / 7e-34
AT5G20900 108 / 4e-29
jasmonate-zim-domain protein 12 (.1)
Lus10027639
121 / 8e-34
AT5G20900 107 / 8e-29
jasmonate-zim-domain protein 12 (.1)
Lus10039911
81 / 5e-18
AT1G19180 161 / 3e-48
jasmonate-zim-domain protein 1 (.1.2)
Lus10027648
76 / 4e-16
AT1G19180 159 / 3e-47
jasmonate-zim-domain protein 1 (.1.2)
Lus10008129
71 / 2e-14
AT1G74950 114 / 1e-30
JASMONATE-ZIM-DOMAIN PROTEIN 2, TIFY domain/Divergent CCT motif family protein (.1)
Lus10013166
71 / 2e-14
AT1G74950 118 / 2e-32
JASMONATE-ZIM-DOMAIN PROTEIN 2, TIFY domain/Divergent CCT motif family protein (.1)
Lus10035804
57 / 1e-09
AT5G20900 62 / 3e-11
jasmonate-zim-domain protein 12 (.1)
Lus10002576
56 / 1e-09
AT5G13220 112 / 2e-31
TIFY DOMAIN PROTEIN 9, JASMONATE-ASSOCIATED 1, jasmonate-zim-domain protein 10 (.1.2.3.4)
Lus10001803
54 / 9e-09
AT5G13220 100 / 1e-26
TIFY DOMAIN PROTEIN 9, JASMONATE-ASSOCIATED 1, jasmonate-zim-domain protein 10 (.1.2.3.4)
Lus10036584
54 / 1e-08
AT1G74950 64 / 9e-12
JASMONATE-ZIM-DOMAIN PROTEIN 2, TIFY domain/Divergent CCT motif family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF06200
tify
tify domain
CL0281
CCT
PF09425
Jas_motif
Jas motif
Representative CDS sequence
>Potri.018G047100.5 pacid=42801180 polypeptide=Potri.018G047100.5.p locus=Potri.018G047100 ID=Potri.018G047100.5.v4.1 annot-version=v4.1
ATGGAGGCTCAACAACCTGATTCTCGTAAGGAAATGAAGCCACGAGAAGATATGGAAGTGAAGAAGGAGCAAGAGGTTATGGGATCTTGCAAGGAAGGTG
CTGGCTTGTTGGAGAAAAATGGGGTTCATCATCTTTGGGCACCCAGTTACTGGCCCATGACAATGGCGACTTCTAGACCAAATGTCACCATTCCTACTCC
AGATCAGCTCACCATCTTCTATGGTGGAAGTGTTGTTGTGTTTGATTCAATTCCTGCAGAAAAGGTTCATGAAATTATGCTTATTGCTGCAGCTGCTGTC
AAACCTGGTGACATGAAAAAGAGTGGTTCTCCTACTGGTACACCAGTTCTTACAAGGTCTCCTTCAATGCAAAGCACTGCTGCACCCCAGGGCCAAACAT
ATAGTCGTCAGAATTCCATCTGCAGAATGCAAGCTGAGTTGCCTATTGCGAGGAGACAATCCCTGCAGCGTTTCTTCAAGAAGCGACGGGACAGGTTGGT
GAGCAAAAGTCCATATCCCACTTCGCCAGCAGGAAAAGAGGCAGACACCACAGAACCTGGTATTAGTGCCGCACCTCCACCAGATGCAGGTTGCTTTGGA
AAACCTCTAGCATCAGAAGAACTTCAACCGAAGGTTGCTGCCAACGTTGTTTGA
AA sequence
>Potri.018G047100.5 pacid=42801180 polypeptide=Potri.018G047100.5.p locus=Potri.018G047100 ID=Potri.018G047100.5.v4.1 annot-version=v4.1
MEAQQPDSRKEMKPREDMEVKKEQEVMGSCKEGAGLLEKNGVHHLWAPSYWPMTMATSRPNVTIPTPDQLTIFYGGSVVVFDSIPAEKVHEIMLIAAAAV
KPGDMKKSGSPTGTPVLTRSPSMQSTAAPQGQTYSRQNSICRMQAELPIARRQSLQRFFKKRRDRLVSKSPYPTSPAGKEADTTEPGISAAPPPDAGCFG
KPLASEELQPKVAANVV
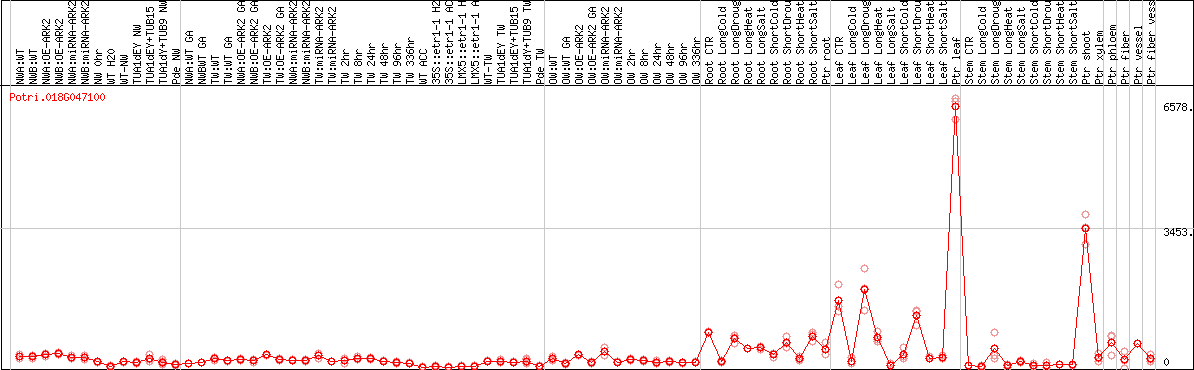
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G047100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.