Potri.018G047500 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G047500.6 pacid=42801363 polypeptide=Potri.018G047500.6.p locus=Potri.018G047500 ID=Potri.018G047500.6.v4.1 annot-version=v4.1
ATGGATACTGCAATGCCAGACCATGATCCTCTCAAGAGAAGCTTTGGTGACATGATGGTCAACAACAACAGCTCTTCGAGCTGTCTTGCTATGGATACTT
CCAATGGTATTACTGATCATAATGACACCACCCCACAAGGACTAGCTTCTGTTCTTAGTAATCATAAGGAGTTTTATCCAAGAGGGTATCAGTCAAAGGT
TTTTGAGGTGGCTGTTAAGAGGAATACAATTGCTGTGCTGGAAACAGGAGCTGGAAAGACAATGATTGCTGTTATGTTAATTAAACAGATTGGTCAAGCT
GTCTTTTACAGTGGTGTTAAGAGATTGATTCTTTTCTTAGCTCCAACAGTTCATCTTGTTAATCAGCAATATGAAGTCATTAAATCTCAAACAAATTTTA
GAGTGGGAGAGTACTATGGAGCTAAGGGAATAGATGAGTGGTCTCTGAAGTCCTGGGAGAAGGAAATTGATGAGCACGATGTGTTGGTTATGACACCCCA
GATCCTCTTGGATGCCTTAAGAAAGGCATTTTTGAATCTAAAAATGGTATCCTTGTTAATACTTGACGAGTGTCATCGTTCCACCGGTAACCATCCTTAT
AAGAAAATTATGAAGGATTTCTATCACAAAATGGAGAATAAGCCAAAGGTTTTTGGAATGACAGCATCTCCTGTAGTTAGAAAAGGTGTCTCGTCTGCCA
TGGACTGTGAGGATCAACTAGCAGAACTTGAGAGTGTATTGGATTCTCAGATTTATACTATTGAAGACAGGGCAGAGGTGCATGTCTATGTTCCCTCTGC
AAAAGAATTATGTAGATTTTATGACAAAGCATGGTGTTCTTATGTGGAGCTGAAAGATAAGATTGAAGCTTCATGGTCCAAGTTTGATGCTTCAATGTTA
GCTTTGCAAGGCTCAACACAAAGTTGTTACAAAGATATGGATGATAAGCTTAAAGCAACGAGAAAGCAGTTGTCCAAGGACCACGCAAAGATTTTGAATT
GCCTAGAAGATCTTGGCCTCATATGTGCTTATGAGGCCATCAAGGTTTGTCTAGAGAATGCTGGTAACCCCACTGGTGAATGCAAATTATATCAAGAAAT
TTCTTTGCAGTGTAGATATTTCCTTGAGGATGTGTTACATATAATTGGTGAATCTTTGCTGCATGGTGATAATTTCTCTTTGGATCATGGGTTTGACTGT
TCAGCAGCATTGGGTTTTGGCTACATATCCCCCAAACTACATGAACTTCTCCAACTTTTTCTATCATTTGGAGAAGCTAGGGAAGTATTGTGCCTCATTT
TTGTTGAAAGAATTATTACAGCGAAAGTGGTTGAAAGATTTATGAAGAAAGTTGAGGTTTTAGCACATTTCACTGTTTCATATTTGACTGGAACTAATGC
ATCAGCTGATGCACTGGCCCCAAAAATGCAAATGGAGACCTTGGAATCATTTCGCTCTGGGAAGGTCAATCTATTATTTGCCACTGATGTGGTGGAGGAG
GGAATTCATGTGCCAAACTGCTCCTGTGTAATACGTTTTGATCTGCCTAAGACAGTCCGCAGTTATGTCCAGTCTCGGGGACGAGCTCGACAAAATAATT
CCCACTTTATCACCATGCTTGAAAGGGGAAACACCAAACAACGGGATCAGCTATTTGAAATCATTAGAAGTGAGTGGTCAATGACAGATACAGCTATAAA
TAGAGATCCTAATGTATGGAATCTGAAAGCATGTGCTTCAGAAGCAGCAAAGGCTTATGTGGTGGATGTGACAGGAGCATCAGTAACTGCAGACTCTAGT
GTTAGCCTCATACATCGATATTGTCAACACCTCCCTGGCGACAGGTACTACACACCAAAGCCAACTTTTCAGTTTGAAGTTTTTGAACAGTCTTGCCGCT
GTGCAATGAAGCTACCTCCTAATGCAGCATTTCAAACATTAGTTGGTCCAACATGTAGGAATCAACAATTAGCAAAGCAGCTTGTATGCTTGGAAGCATG
TAAGAAATTGCATCAAATGGGTGCTTTAGATGATCATCTTCTGCCATCAGTTGAAGAGCCTTCAGAAATTGCTGTTGTTAAAAGCAAGTCAACATCTGCA
GGTGCAGGAACTACAAAAAGGAAGGAATTGCATGGGACAGCTTGCATTCATGCGTTATCCGGAAGTTGGGGAGAGAAACTTGATGGAGCCACTTTTCATG
CATACAAGTTTGATTTCTCTTGCTCCATTGTCAGTCAGATCTATTCTGGATTTATTCTTCTCATTGAGTCAAAGCTCGATGATGATGTGGGAAACATTGA
GTTGGATCTTTATTTGGTTGCAAAGATAGTCAAGTCTTCTATTTCTTCATGTGGAGTAGTTCACTTGGATGCTGCACAGATGACAAAAGCAAAACGGTTT
CAAGAATTCTTTTTCAATGGCTTGTTTGGAAAGTTGTTTACTGGATCTAAATCATCTAGGGAGTTCTTACTTCAGAAAGAAACAACATTACTGTGGAGTC
CCTCAAACATGTATCTGCTTCTACCACTAGAGCCATGGAGCATTTCCAGTAATGATTGGTGTAAAATAGATTGGAAAGGAATTGAAGCTTGCTCATCTGT
GGTAGAATACTTGAAGAACTCTTTTTTGGCTGCTCGGTCTTACAGTGGTGGAGGAAATCCATTACCTGATAATGTTCAGTCATCCACCATAGAATGCAAT
GGTACAAATTTAATCCATTTTGCTAATGCTTTAGTCAACGTAGAGAACATTAAAGATATGGTGGTACTGGCAATCCACACGGGAAGAATCTACTCCATTG
TTAAAGTCGTGAACGATTCATCTGCGGAGAGTGCTTTTGAGGGAAATGCTGATAATGTGACAGAGTTTTCTACATACACAGAGTACTTCAACAAAAGGTA
CGGAATTGTGCTGATGCATCCAGGACAGCCTCTGTTGCGGTTAAAGCAAAGCCATAACCCACACAATCATCTTGTAAACTTTAATGATGAAGGTGATTCA
AAAGATGGCATGGTTGGCAGAAAACAGCAGCAGCATGTGCATATGCCACCAGAGCTTTTGATCAAGATTGATGTTCCTATCAGTGTTGTAAAATCAATTT
ATCTGATGCCATCTTTGATGCACCGTCTTGAGTGCCTGATGTTGGCCAGTCAACTTAGACAAGAGATTGATTGTCATGCTCCTAATTTTTACATCCCAAG
CTCATTGATTCTGGAAGCGATAACAACACTTAGATGCTGTGAAAGTTTTTCGATGGAGCGACTGGAGTTGCTAGGGGACTCAGTTCTAAAGTATGCTGTC
AGCTGCCACCTATTTTTAAAATATCCCAATAAACATGAAGGCCAGTTATCCTCATGGCGCTCAGGGGCTGTTTGTAATTCAACCCTACATAAATTGGGAA
CAGATTGTAAAGTACAGGGATATATACTAGACAGTGCATTTGATCCCCGTCGTTGGGCTGCTCCTGGACAGAAATCTGTACGTACTCCTGCTCCTTGCAA
ATGTGGGGTTGATACTTTAGAAGTACCATTGGATCGTAAGTTCCAAACCGAAAGCGCAATTGTTAAGGTTGGAAAACCTTGTGATTCAGGCCACCGATGG
ATGGGTTCCAAAACCATATCAGATTGTGTTGAATCTGTCATAGGGGCATACTATGTCAGTGGTGGATTGATTGCTGCAATTCATGTGATGAAGTGGTTTG
GCATTAATGCTGAACTTGATCCTTCACTAATAAGCGAAGCAATTACGAGTGCATCTCTACGATCTTATATCCCTAAAGAAGATGAGATTAAGAGTCTAGA
GTCAAAGCTTGGGTATACCTTCGGTGTCAAGTTTGTCTTGCAGGAGGCCATGACTCATGCATCTATACAAGAACAGGGTGTTACATACTGTTACCAGAGG
CTTGAATTTCTTGGTGATTCTGTGTTGGACTTGCTTATAACATGGCATCTCTATCAGAGCCACACAGATGTTGATCCTGGCGAGCTGACTGACTTGCGCT
CAGCTTCTGTTAACAATGATAACTTTGCTCAAGTTGCTGTGAAACAAAACCTATATACCCATCTTCTTCATTGTTCTACACTCCTTCAAAGTCAAATAAC
AGAATATGTAAATTCTTTTCATGAATCTGATCAAGGCACAAAGGCTCCCAAGGCTCTTGGAGACCTGATTGAAAGCATTGCAGGGGCATTATTAATTGAT
ACAAAGTTCAATCTCGATGGTGTGTGGAGAATATTCAAGCCCTTGTTATCTCCAATTGTAACCCCTGAGAAACTTGAGCTGCCTCCACTGCGTGAACTTG
TTGAATTATGTGACTCTATAGGGGTTTTTGTAAAAGAAAAATGTACCAAGAAAGCTGAGATGGTTCACGCCCAGCTTTGGGTACAGCTGGACAACGAGCT
CTTGTCTGGAGAGGGGTACGAGAAGAACAGGAAAGCAGCTAAAGGAAAAGCAGCTTCTTGTTTGTTGAAGAAGCTCCAGAAAAGAGGCATCGTATACTCA
CGTGGAGGTTCAAAGAGGAGGAAACAGGACACTGACCCTGTTGTTGATTCAAGTTCCCTTGGCTTCTTAGAAAGTGAAGATTTTTCTGGAAAAACAAAGC
CAAAAAAGCAGAAAATAGAAAACCAAGTGCCCGGAGATTCAAATACAGATTGTTCCCCTGCCATCAGTCCTAGTCATGGTCCCCCAGTTATTGAGTCGAT
TAACAAGAAGAAAGGAGGACCTCGTACTAGTCTTTATGATCTCTGTAAGAAAGTTCAATGGACAATGCCTACATTTGACACAACAGAAACGAAATCCAGA
ACTGCAATTGAATTTGGTGAAGGCCCTGATAAAAGGACGGGATTTAACAGTTATGTATCAAAAATCATCATGAACATACCGTCGTATGGCGTTGTTGAAT
GTGCGGGAGAGGCTAGTGCTGATAAGAAGACCTCATATGACTCTGCGGCACTTGCAATGCTTAATGAGCTTGAAAAACGGGGACAGCTCATCATTGATGA
ATCAAAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G047500.6 pacid=42801363 polypeptide=Potri.018G047500.6.p locus=Potri.018G047500 ID=Potri.018G047500.6.v4.1 annot-version=v4.1
MDTAMPDHDPLKRSFGDMMVNNNSSSSCLAMDTSNGITDHNDTTPQGLASVLSNHKEFYPRGYQSKVFEVAVKRNTIAVLETGAGKTMIAVMLIKQIGQA
VFYSGVKRLILFLAPTVHLVNQQYEVIKSQTNFRVGEYYGAKGIDEWSLKSWEKEIDEHDVLVMTPQILLDALRKAFLNLKMVSLLILDECHRSTGNHPY
KKIMKDFYHKMENKPKVFGMTASPVVRKGVSSAMDCEDQLAELESVLDSQIYTIEDRAEVHVYVPSAKELCRFYDKAWCSYVELKDKIEASWSKFDASML
ALQGSTQSCYKDMDDKLKATRKQLSKDHAKILNCLEDLGLICAYEAIKVCLENAGNPTGECKLYQEISLQCRYFLEDVLHIIGESLLHGDNFSLDHGFDC
SAALGFGYISPKLHELLQLFLSFGEAREVLCLIFVERIITAKVVERFMKKVEVLAHFTVSYLTGTNASADALAPKMQMETLESFRSGKVNLLFATDVVEE
GIHVPNCSCVIRFDLPKTVRSYVQSRGRARQNNSHFITMLERGNTKQRDQLFEIIRSEWSMTDTAINRDPNVWNLKACASEAAKAYVVDVTGASVTADSS
VSLIHRYCQHLPGDRYYTPKPTFQFEVFEQSCRCAMKLPPNAAFQTLVGPTCRNQQLAKQLVCLEACKKLHQMGALDDHLLPSVEEPSEIAVVKSKSTSA
GAGTTKRKELHGTACIHALSGSWGEKLDGATFHAYKFDFSCSIVSQIYSGFILLIESKLDDDVGNIELDLYLVAKIVKSSISSCGVVHLDAAQMTKAKRF
QEFFFNGLFGKLFTGSKSSREFLLQKETTLLWSPSNMYLLLPLEPWSISSNDWCKIDWKGIEACSSVVEYLKNSFLAARSYSGGGNPLPDNVQSSTIECN
GTNLIHFANALVNVENIKDMVVLAIHTGRIYSIVKVVNDSSAESAFEGNADNVTEFSTYTEYFNKRYGIVLMHPGQPLLRLKQSHNPHNHLVNFNDEGDS
KDGMVGRKQQQHVHMPPELLIKIDVPISVVKSIYLMPSLMHRLECLMLASQLRQEIDCHAPNFYIPSSLILEAITTLRCCESFSMERLELLGDSVLKYAV
SCHLFLKYPNKHEGQLSSWRSGAVCNSTLHKLGTDCKVQGYILDSAFDPRRWAAPGQKSVRTPAPCKCGVDTLEVPLDRKFQTESAIVKVGKPCDSGHRW
MGSKTISDCVESVIGAYYVSGGLIAAIHVMKWFGINAELDPSLISEAITSASLRSYIPKEDEIKSLESKLGYTFGVKFVLQEAMTHASIQEQGVTYCYQR
LEFLGDSVLDLLITWHLYQSHTDVDPGELTDLRSASVNNDNFAQVAVKQNLYTHLLHCSTLLQSQITEYVNSFHESDQGTKAPKALGDLIESIAGALLID
TKFNLDGVWRIFKPLLSPIVTPEKLELPPLRELVELCDSIGVFVKEKCTKKAEMVHAQLWVQLDNELLSGEGYEKNRKAAKGKAASCLLKKLQKRGIVYS
RGGSKRRKQDTDPVVDSSSLGFLESEDFSGKTKPKKQKIENQVPGDSNTDCSPAISPSHGPPVIESINKKKGGPRTSLYDLCKKVQWTMPTFDTTETKSR
TAIEFGEGPDKRTGFNSYVSKIIMNIPSYGVVECAGEASADKKTSYDSAALAMLNELEKRGQLIIDESK
|
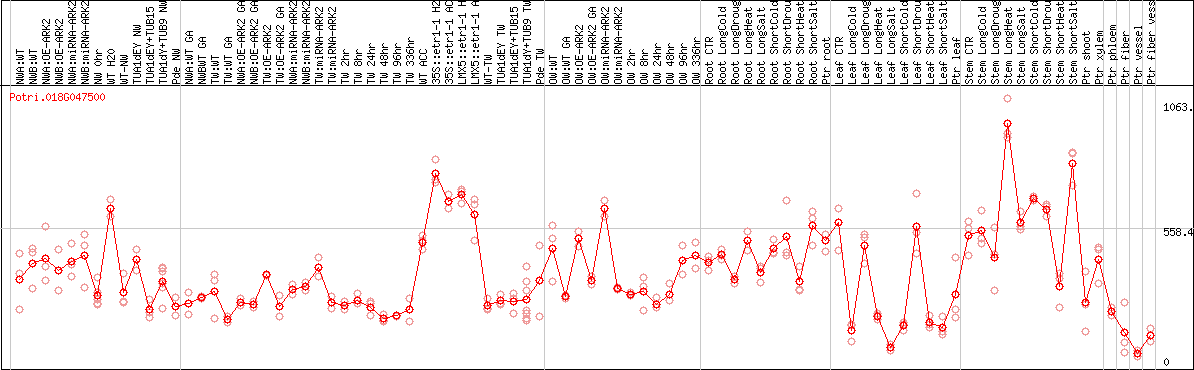
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G047500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
