Potri.018G068100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G068100.25 pacid=42801945 polypeptide=Potri.018G068100.25.p locus=Potri.018G068100 ID=Potri.018G068100.25.v4.1 annot-version=v4.1
ATGGGGCTGGTGATGGTGCCACCCATTCTTCAGCTGCTGGGGAAACTCTCTGCCATTGGCTTTTTTTTCGCAGTTCAAGGATCTACTGAAGGAACGCCTG
GTCCCCAACTCCAGCCATTCAGAAGCAGTGTGCCAGCCTCAGAGTCTCTGTCAACTGGGTCAAACTTGCACCCTCCACTTGCATTGCCACCACTCACCTC
TGCCCCAGTGCCCCAAACAATCACAGGGCATGTACCATCATATTCTCCTAGTCCTCTGATAGTAACAAAACCACATAATAGAGCTCCTCCTCCCTTGATT
TCTCAAGGACATGTACCATCCTTGTCACCAAGTTTTTCAGTTACATCTCCAACGTATAATACAGCTCCTCCACCCACACTTAATCAAGGGCATGAGCCAC
CTAAGCCACCAAATGCTCATCGAAGAGAGGCAGCAGTTAGACAGACTCCTGTCTCTGCGTCAAGTGCGCCAGCTGCAGCACCTTCGAGACAGTCACCAGA
AAATTCACCAGTCAGCCAATCAAATGCACCTGAAGCTTCATCACTAAGTACTCATAAAAAAGATGTATCCTTCAGTGGGGCTCCTGTCCCAAATTCAGTT
TCTCCAGCTGCATCTCCATCAAGGACTTCAAAAGAAAATCAACCAGCAGCCCCCCCTATTGCACCAGCAACGATGCCATCTGCAGTTCCGGTTGTATCAC
CAATGGGAGCCATGCCACAAAATTCATCAGCAACCCACCCAGTCATGCCTGGAGAACCCCCTGCTATATTATCAGGTCCAAATGTCCCCCATCCATCTGC
TCCTACCCCAAGCATTGATGATAAGAAAGATGAAATTCCAGTTTCTGCACCGCCAAATGAAACACCCACACCTTTATCTCCCATGAGCCATTTTCCAGCT
AAAGGTTCTTTTTCTGGTATAGCTCCTTCCACGCACAATGCCATGAGGCACTCTAATGACTCTCCCATTTCTCCTTCATCATTGCCGAAAACTCCTCTGG
CTAAACAGCATCACTCTTCTGCGCCTTCACCTTCTATCCCATTTCATAAACAGAACCATGAAAGCAGCAGAATCAGTGACTCTGCTCCTGCATCCTCAAA
TCCAACATTTCCGCCATCTTCAAAACAGCAAGGTCCAGTCATGCCTCCATCATTTCTTCCAACAAGCAGGCATAAACAATATGCTTTCTCACCCCTTGGT
ACTCCAGATTCTTCAAATAATGCATCACACTACCGCTATCCTAAACCAGTGATCATTGTCTCACCCACTCCTTCACCCACTCAAACTGCAGCATCTGGAT
GGACAAAAATGCCAGCTCTTTCCCCTAAAGCTTCTCCTTCTGGGTTTTCTTCAAGAACTCCCAAGATGCCCCCCCTGCCACCACTCCACACATTGCCACC
TCCACCTCCTAATGAAGATTGTTCAACGACTGTGTGCACAGAGCCATACACAAATACTCCTCCTGGGTCACCATGTCGCTGTGTCTTGCCCATGCAAGTA
GGACTGGGCGTTAGTGTTGCACTGTATACCTTTTTCCCTTTAGTTTCAGAGCTTGCTCAGGAAATTGCTGCTGGCGTTTTTATGAAACAAGGTCAAGTTC
GCATTATAGGGGCCAATGCCCCCAGCCAGCAGCTAGAGAAGACAATCGTCCTTATTGACTTGGTTCCCCTTGGAGAAAGATTTGACAATTCAACTGCCCT
TTTGACTTATCTAAGATTTTGGAAAAAACAAGTTGTTATAAACCCTTCTTTTTTTGGTGACTATGAAGTATTATATGTGCGTTATCTAGGTTTACCTCCA
CCTCCACCTATGGCACCTTTTGGAATTGCAATAATCGATGATGGGCCATATTCTGGCAATGACAACAACGCAAGGATGATAAAGCCCCTAGGGGTTGATG
TCCATAAAAGGAAGCGTAAAGATGGGCTTGCTGGTGGCATCATTGCTATTATTGCTGTGTCTGGTTTTGTAGCACTAGTTTTATTCTCTGCTGTTGCTTT
GGCATTGCTGTTTAAACATAGAGATCATGCTAGTCAACCGGCATCAGTCCTGCAGCCATTGCCACCTTCAGTTGTGAAACCATCAGGTATTGCTGGGTCA
TTGATTGGAAGTGGTCTTAGTTCTGCCTCATTGTCCTTTGCATCTAGTATTCCAGCGTACACAGGATCTGCTAAGACCTTCAGCAAAAGTGACATAGAGC
GGGCCACTAGTAGCTTTGATGCTTCAAGAATACTCGGGGAGGGTGGCTTTGGGCTTGTTTACAGTGGTGTTTTAGAAGACGGGACCAAAGTTGCTGTAAA
AGTTCTAAAGAGAAATGATCAGCAAGGTGGTCGAGAATTCTTGGCAGAAGTTGAGATGCTTAGCCGCCTTCATCACAGAAATTTAGTCAAGTTGATTGGA
ATATGCACAGAGGAATGTTCACGCAGCTTGGTCTACGAGCTCATAGCAAATGGCAGTGTCGAATCTCATCTACATGGGGTTGATAAAGAATCTGCTCTTA
ATTGGGATGCTCGGATCAAGATAGCACTTGGTGCAGCTCGAGGTCTAGCTTATCTACATGAGGATTCAAGCCCTCGTGTGATACACAGGGACTTCAAGTC
AAGCAACATTTTGTTGGAACATGATTTCACACCAAAAGTCTCAGACTTTGGTTTGGCTAGGACTGCCTTGGATGAGGAAAATAGGCACATATCAACGCAG
GTCATGGGAACTTTTGGGTATGTCGCTCCAGAGTATGCAATGACTGGCCATCTTCTTGTGAAGAGTGATGTTTACAGTTATGGTGTAGTCCTTCTTGAGC
TCCTAACAGGACGAAAACCAGTGGATATGTCCCAGCCACCTGGTCAAGAGAATCTAGTTGCATGGGCTCGTCCATTTCTAACAACTAAAGAAGGGCTTGA
AGTAATTATAGATCCGACTCTAGCAACTGATGTACCTTTTGATAGTGTGGCCAAAGTAGCAGCTATTGCATCAATGTGTGTTCAACCAGAGGTATCACAC
CGTCCTTTTATGAGTGAGGTTGTTCAGGCCTTAAAACTAGTATCTAATGAGTGTGATGAGGCAAAGGAACTGGATTCCAGGAGTTCCAGCCAAGATTTAT
CTATTTACATGGATGCTGAGGTCAGTGCAGTCTCAGGACAGCTTCGGGATTCTTTACGAAGTCAGGCCCTAGTACATAATTATGATTCCGAGCCAGATAT
AGAAAGGGGACTCGTGGATTCATATTTTTTCAGTACATCAGACGGGTGTGGGAGGCAGGGCTCTGGATCATTGAGGAGATGCTCGTCAGGTCCATTGAAA
ACAGGAAGGGGCAGAGAGTTGTTTCAGAGAATGAGATTGACAGGGGAAAGTGTGAGTGAACGTGGAACCATCTTCAAAATGTGGCCAGGTTCACATTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G068100.25 pacid=42801945 polypeptide=Potri.018G068100.25.p locus=Potri.018G068100 ID=Potri.018G068100.25.v4.1 annot-version=v4.1
MGLVMVPPILQLLGKLSAIGFFFAVQGSTEGTPGPQLQPFRSSVPASESLSTGSNLHPPLALPPLTSAPVPQTITGHVPSYSPSPLIVTKPHNRAPPPLI
SQGHVPSLSPSFSVTSPTYNTAPPPTLNQGHEPPKPPNAHRREAAVRQTPVSASSAPAAAPSRQSPENSPVSQSNAPEASSLSTHKKDVSFSGAPVPNSV
SPAASPSRTSKENQPAAPPIAPATMPSAVPVVSPMGAMPQNSSATHPVMPGEPPAILSGPNVPHPSAPTPSIDDKKDEIPVSAPPNETPTPLSPMSHFPA
KGSFSGIAPSTHNAMRHSNDSPISPSSLPKTPLAKQHHSSAPSPSIPFHKQNHESSRISDSAPASSNPTFPPSSKQQGPVMPPSFLPTSRHKQYAFSPLG
TPDSSNNASHYRYPKPVIIVSPTPSPTQTAASGWTKMPALSPKASPSGFSSRTPKMPPLPPLHTLPPPPPNEDCSTTVCTEPYTNTPPGSPCRCVLPMQV
GLGVSVALYTFFPLVSELAQEIAAGVFMKQGQVRIIGANAPSQQLEKTIVLIDLVPLGERFDNSTALLTYLRFWKKQVVINPSFFGDYEVLYVRYLGLPP
PPPMAPFGIAIIDDGPYSGNDNNARMIKPLGVDVHKRKRKDGLAGGIIAIIAVSGFVALVLFSAVALALLFKHRDHASQPASVLQPLPPSVVKPSGIAGS
LIGSGLSSASLSFASSIPAYTGSAKTFSKSDIERATSSFDASRILGEGGFGLVYSGVLEDGTKVAVKVLKRNDQQGGREFLAEVEMLSRLHHRNLVKLIG
ICTEECSRSLVYELIANGSVESHLHGVDKESALNWDARIKIALGAARGLAYLHEDSSPRVIHRDFKSSNILLEHDFTPKVSDFGLARTALDEENRHISTQ
VMGTFGYVAPEYAMTGHLLVKSDVYSYGVVLLELLTGRKPVDMSQPPGQENLVAWARPFLTTKEGLEVIIDPTLATDVPFDSVAKVAAIASMCVQPEVSH
RPFMSEVVQALKLVSNECDEAKELDSRSSSQDLSIYMDAEVSAVSGQLRDSLRSQALVHNYDSEPDIERGLVDSYFFSTSDGCGRQGSGSLRRCSSGPLK
TGRGRELFQRMRLTGESVSERGTIFKMWPGSH
|
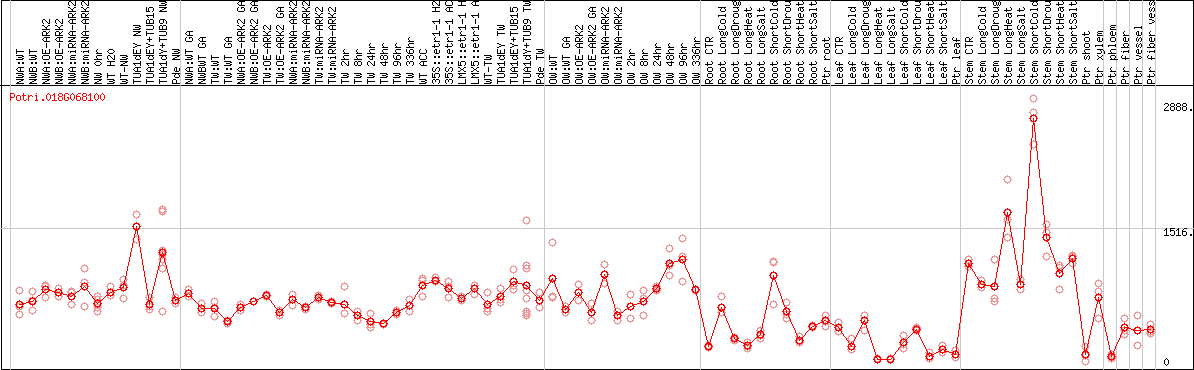
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G068100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
