External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25930 227 / 2e-69
Protein kinase family protein with leucine-rich repeat domain (.1)
AT4G28650 163 / 1e-46
Leucine-rich repeat transmembrane protein kinase family protein (.1)
AT1G08590 154 / 8e-44
Leucine-rich receptor-like protein kinase family protein (.1)
AT4G28490 152 / 3e-43
RLK5, HAESA
RECEPTOR-LIKE PROTEIN KINASE 5, HAESA, Leucine-rich receptor-like protein kinase family protein (.1)
AT5G65700 149 / 5e-42
BAM1
BARELY ANY MERISTEM 1, Leucine-rich receptor-like protein kinase family protein (.1.2)
AT3G49670 148 / 1e-41
BAM2
BARELY ANY MERISTEM 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT2G33170 148 / 2e-41
Leucine-rich repeat receptor-like protein kinase family protein (.1)
AT1G17230 148 / 2e-41
Leucine-rich receptor-like protein kinase family protein (.1)
AT5G61480 146 / 5e-41
PXY, TDR
TDIF receptor, PHLOEM INTERCALATED WITH XYLEM, Leucine-rich repeat protein kinase family protein (.1)
AT4G20270 144 / 2e-40
BAM3
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034978
229 / 5e-70
AT5G25930 927 / 0.0
Protein kinase family protein with leucine-rich repeat domain (.1)
Lus10012949
228 / 5e-70
AT5G25930 910 / 0.0
Protein kinase family protein with leucine-rich repeat domain (.1)
Lus10005491
228 / 1e-69
AT5G25930 1104 / 0.0
Protein kinase family protein with leucine-rich repeat domain (.1)
Lus10006028
228 / 1e-69
AT5G25930 1116 / 0.0
Protein kinase family protein with leucine-rich repeat domain (.1)
Lus10003503
153 / 6e-47
AT1G72180 278 / 2e-87
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10038382
160 / 9e-47
AT4G20270 734 / 0.0
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10027951
163 / 1e-46
AT1G08590 1256 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10000814
162 / 1e-46
AT1G08590 1263 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10025943
162 / 3e-46
AT4G28650 1284 / 0.0
Leucine-rich repeat transmembrane protein kinase family protein (.1)
Lus10036242
162 / 3e-46
AT4G20270 1248 / 0.0
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Potri.018G088086.1 pacid=42801438 polypeptide=Potri.018G088086.1.p locus=Potri.018G088086 ID=Potri.018G088086.1.v4.1 annot-version=v4.1
ATGCACCATGACTGCTCTCCTCCCATAGTACACCGAGATGTCAAGTCAAGCAATATACTGTTGGATTCTGAGTTCAACGCAAAAATCGCTGATTTTGGTT
TAGCAAGGATGCTTGTTAAGCAGGGAGAACTTGCTACTGTGTCTGCTGTGGCTGGTTCTTTGGGTTACATAGCTCCAGAGTATGCTCGAACAGTTAGAGT
GAATGAGAAGATTGATGTTTATAGCTTTGGAGTTGTCCTTCTTGAACTAACGACAGGGAAAGCAGCTAATTATGGTGACGAGGATACATGCCTTGCCGAA
TGGGCATGGCGTCACATGCAAGAAGGAAAACCTATAGTCGATGTGTTAGATGAGGAGATAAAGGAGCCTTGTTACGTGGATGAGATGAGAGATGTTTTCA
AACTAGGGGTGTTCTGTACGAGCATGTTGCCTTCAGAAAGGCCTAACATGAAGGACGTTGTCCAAATCTTACTTGGTAGGAACCGTCGATGGGTCTGTGG
AAGGAAGAATATGAGACATGCATGA
AA sequence
>Potri.018G088086.1 pacid=42801438 polypeptide=Potri.018G088086.1.p locus=Potri.018G088086 ID=Potri.018G088086.1.v4.1 annot-version=v4.1
MHHDCSPPIVHRDVKSSNILLDSEFNAKIADFGLARMLVKQGELATVSAVAGSLGYIAPEYARTVRVNEKIDVYSFGVVLLELTTGKAANYGDEDTCLAE
WAWRHMQEGKPIVDVLDEEIKEPCYVDEMRDVFKLGVFCTSMLPSERPNMKDVVQILLGRNRRWVCGRKNMRHA
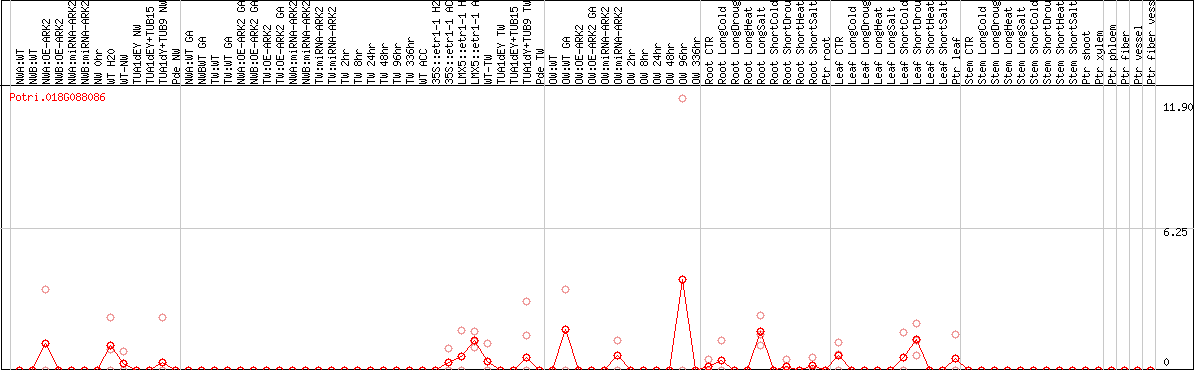
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G088086 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.