Pt-VLN4.2 (Potri.018G089700) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pt-VLN4.2 | ||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.018G089700.4 pacid=42801046 polypeptide=Potri.018G089700.4.p locus=Potri.018G089700 ID=Potri.018G089700.4.v4.1 annot-version=v4.1
ATGGCTGTTTCCATGAGAGATTTAGACTCGGCCTTTCAGGGAGCTGGACAAAAGGCTGGACTTGAAATATGGCGGATTGAAAACTTTCGTCCTGTTCCTG
TGCCGAAGTCCTCTCATGGAAAATTTTTTACCGGGGACTCCTATGTAATTTTACAGACAACTGCACTAAAAAGTGGTTCCTTACGTCATGATATTCATTA
TTGGCTAGGTAAAGATACCTCTCAGGATGAAGCAGGTGCTGCTGCCATTAAAACAGTTGAACTAGATGCTGCTCTTGGAGGCCGTGCTGTGCAGTATCGT
GAAGTGCAGGGACATGAAACTGAGAAGTTCTTATCTTATTTTAAACCATGTATTATACCTCAAGAAGGTGGAGTCGCATCTGGATTCAAACAAGCTGAGG
CCATGGAACACCAGACACACTTGTTTGTTTGTAGAGGAAAACACGTTGTCCATGTGAACGAGGTTCCTTTTGCTCGATCCTCACTCAACCACGATGATAT
TTTTATTCTTGATACAAAGTCAAAAATTTTTCAATTTAATGGCTCCAACTCATCCATTCAAGAGAGAGCTAAAGCTTTAGAGGTGGTTCAGTACATCAAA
GATACTTATCATGATGGAAAATGTGAGGTGGCTGCTGTTGAGGATGGAAAATTGATGGCTGATGCTGAAACTGGAGAGTTCTGGGGTTTCTTTGGTGGCT
TTGCTCCACTTCCAAGGAAAACAACTAGTGATGAGGATAAGACTGATGTTTCTTTTCCTACTAAGCTGTTTCATGTTGAGAAGGGACAGGCAGAACCTGT
TGAAGCTGATTCTTTGACAAGGGAGTTGCTGGACACCAATAAGTGTTACATTCTAGATTGTGGAATTGAAGTCTTTGTATGGATGGGGAGAAATACTTCT
CTTGACGAAAGGAAAAGTGCCAGCGGAGCAGCGGAAGAGTTAGTCCGTGCTGCTGAACGTCCAAATTCCCGCATTGCTCGTGTAATTGAGGGGTTTGAGA
CGGTGATGTTTAGATCAAAATTTGAATCGTGGCCTCAGACAACTAACGTGACTGTGTCTGAGGATGGTAGAGGAAAAGTTGCAGCACTTCTAAGACGCCA
GGGGGTTAATGTCAATGGTCTGTTGAAAACTGCTCCTGTGAAAGAAGAACCTCAACCATACATTGATGTCACTGGGAATTTGCAGGTTTGGAGTGTGAAT
GACCAGGAAAAGATTCTCATTCCTGCTGCAAATCAGTCAAAATTTTACAGTGGAGGTTGCTATATTTTTCAGTATTCATATCCAGGAGAAGACAGGGAAG
AATATCTTATAGGAACATGGTTTGGGAAGAAGAGTGTTGAGGAGGAAAGAGCTTCTGCTATTTCATTGGCGAGCAAGATGGTTGAATCATTAAAGTTTCT
ACCTGCCCAGGCTCGCATCTTTGAAGGAAATGAGCCTATCCAGTTCTTTTCAATCTTCCAGAGCTTTATTGTATTTAAGGGTGGTCATAGCTCTGGTTAC
AAGAAATATATTGCTGAAAATGAACTCCCAGATGAGACATGCAAGGAGGATGGTGTTGCGTTATTCAGGGTTCAGGGTTCTGGACCTGATAATATGCAGG
CTATACAAGTTGAGCCAGTTGCATCATCACTGAATTCCTCCTACTGTTACATATTACATAATGATTCCTCTGTTTTTACGTGGTCTGGAAACCTTACTAC
TTCAGAGGACCAGGAACTTATTGAAAGGCAGCTGGATTTGATAAAGCCAAATATGCAGTCTAAGCCACAAAAGGAAGGTTCAGAATCTGAACAATTTTGG
GATTTGTTAGGAGGTAAATCTGAATATCCCAGTCAAAAACTTGCAAGGGAAGCTGAAAGTGATCCGCACCTATTCTCTTGCATATTTTTGAAAGGAAACC
TGAAGGTTTCTGAGATATACAACTTCACCCAGGATGATTTAATGACTGAAGATATTTTTATCTTGGATACTCACTCTGAGATCTTTGTCTGGGTGGGGCA
ACAGGTTGACTCCAAAAGCAAACTGCAAGCCTTGAGTATTGGAGAGAAATTTCTTGAGCATGATTTTCTGTTAAAGAAGTCATCCGGTGAAACTCCAATA
TATATTGTTATGGAAGGGAGTGAGCCACCTTTCTTCACACGCTTCTTCACATGGGATTCTGCAAAATCTTCAATGCATGGAAACTCATTCCAAAGGAAAC
TTGCAATTGTAAAAAATGGGGGTACTCCACTTCTAGATAAACCCAAACGTAGAACAGCCGTATCTTATGGAGGCAGATCTAGTGTACCTGACAAATCACA
GCGTTCCAGAAGCATGTCTTTCAGTCCTGACAGAGTTCGAGTTAGGGGAAGGTCTCCAGCTTTCAATGCCCTGGCTGCTAATTTTGAGAACCCTAATGCC
AGAAACCTTTCAACTCCACCACCAGTTGTAAGAAAGGTATATCCAAAGTCTGTGAGCCCTGATTCAGCAAAACTGGCTTCCAAATCTGCAGCTATAGCAG
CATTGACTGCAAGTTTTGAACAACCGCCACCAGCACGACAAGTAATTATGCCGCGGTCTGTCAAGGTGAGCCCTGAGACTCCAAAATCGACACCTGAGTC
AAACTCTAAGGAGAAGCCTATAAGCATCAGAATAGAATCCCTGACCATACAGGAGGATGTGAAAGAGGGTGAAGCTGAAGATGAGGAAGGGCTCCCAATT
TATCCATATGAAGGCCTTAAAGTTAACTCGCCAGATCCTGTAACTGAAATTGATGTGACCAAGCGAGAGACTTACCTCTCAGCAGCAGAATTTAGGGAGA
AATTTGGGATGGCAAAGGATGCTTTCTATAAGTTGCCCAAATGGAAACAGAACAAGCTCAAAATGGCCCTTCAATTGTTCTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.018G089700.4 pacid=42801046 polypeptide=Potri.018G089700.4.p locus=Potri.018G089700 ID=Potri.018G089700.4.v4.1 annot-version=v4.1
MAVSMRDLDSAFQGAGQKAGLEIWRIENFRPVPVPKSSHGKFFTGDSYVILQTTALKSGSLRHDIHYWLGKDTSQDEAGAAAIKTVELDAALGGRAVQYR
EVQGHETEKFLSYFKPCIIPQEGGVASGFKQAEAMEHQTHLFVCRGKHVVHVNEVPFARSSLNHDDIFILDTKSKIFQFNGSNSSIQERAKALEVVQYIK
DTYHDGKCEVAAVEDGKLMADAETGEFWGFFGGFAPLPRKTTSDEDKTDVSFPTKLFHVEKGQAEPVEADSLTRELLDTNKCYILDCGIEVFVWMGRNTS
LDERKSASGAAEELVRAAERPNSRIARVIEGFETVMFRSKFESWPQTTNVTVSEDGRGKVAALLRRQGVNVNGLLKTAPVKEEPQPYIDVTGNLQVWSVN
DQEKILIPAANQSKFYSGGCYIFQYSYPGEDREEYLIGTWFGKKSVEEERASAISLASKMVESLKFLPAQARIFEGNEPIQFFSIFQSFIVFKGGHSSGY
KKYIAENELPDETCKEDGVALFRVQGSGPDNMQAIQVEPVASSLNSSYCYILHNDSSVFTWSGNLTTSEDQELIERQLDLIKPNMQSKPQKEGSESEQFW
DLLGGKSEYPSQKLAREAESDPHLFSCIFLKGNLKVSEIYNFTQDDLMTEDIFILDTHSEIFVWVGQQVDSKSKLQALSIGEKFLEHDFLLKKSSGETPI
YIVMEGSEPPFFTRFFTWDSAKSSMHGNSFQRKLAIVKNGGTPLLDKPKRRTAVSYGGRSSVPDKSQRSRSMSFSPDRVRVRGRSPAFNALAANFENPNA
RNLSTPPPVVRKVYPKSVSPDSAKLASKSAAIAALTASFEQPPPARQVIMPRSVKVSPETPKSTPESNSKEKPISIRIESLTIQEDVKEGEAEDEEGLPI
YPYEGLKVNSPDPVTEIDVTKRETYLSAAEFREKFGMAKDAFYKLPKWKQNKLKMALQLF
|
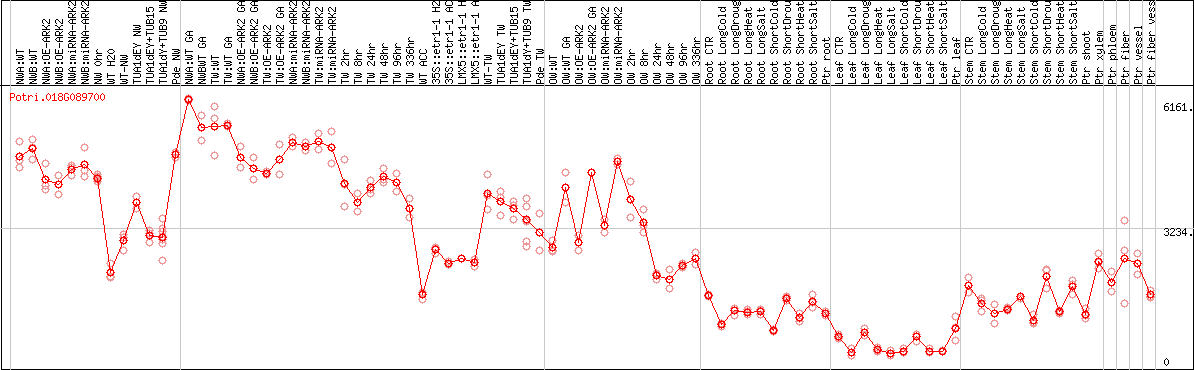
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.018G089700 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
